External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G04130 231 / 3e-73
Tetratricopeptide repeat (TPR)-like superfamily protein (.1), Tetratricopeptide repeat (TPR)-like superfamily protein (.2)
AT3G22670 135 / 1e-36
Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT3G53700 85 / 1e-18
MEE40
maternal effect embryo arrest 40, Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT1G63150 81 / 4e-17
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT1G09900 80 / 4e-17
Pentatricopeptide repeat (PPR-like) superfamily protein (.1)
AT5G64320 81 / 5e-17
Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT5G59900 80 / 5e-17
Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT1G09820 79 / 1e-16
Pentatricopeptide repeat (PPR-like) superfamily protein (.1)
AT1G62910 78 / 3e-16
Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT4G19890 78 / 3e-16
Pentatricopeptide repeat (PPR-like) superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.010G055500
363 / 2e-124
AT3G04130 566 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1), Tetratricopeptide repeat (TPR)-like superfamily protein (.2)
Potri.010G084000
141 / 1e-38
AT3G22670 489 / 3e-168
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Potri.010G083900
134 / 3e-36
AT3G22670 439 / 1e-148
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Potri.010G083800
115 / 2e-29
AT3G22670 428 / 3e-144
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Potri.005G046000
92 / 3e-21
AT1G63130 405 / 9e-133
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.004G013300
89 / 5e-20
AT3G53700 1092 / 0.0
maternal effect embryo arrest 40, Pentatricopeptide repeat (PPR) superfamily protein (.1)
Potri.006G015900
82 / 1e-17
AT5G64320 878 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Potri.003G154800
82 / 1e-17
AT4G20090 816 / 0.0
embryo defective 1025, Pentatricopeptide repeat (PPR) superfamily protein (.1)
Potri.015G121700
82 / 2e-17
AT4G19890 923 / 0.0
Pentatricopeptide repeat (PPR-like) superfamily protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10036975
235 / 1e-75
AT3G04130 473 / 2e-165
Tetratricopeptide repeat (TPR)-like superfamily protein (.1), Tetratricopeptide repeat (TPR)-like superfamily protein (.2)
Lus10015827
236 / 7e-75
AT3G04130 535 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1), Tetratricopeptide repeat (TPR)-like superfamily protein (.2)
Lus10041193
129 / 3e-34
AT3G22670 531 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Lus10039056
87 / 2e-19
AT5G64320 832 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Lus10012037
85 / 2e-18
AT2G15630 677 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Lus10005303
84 / 4e-18
AT1G30290 1037 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10006373
83 / 8e-18
AT3G53700 1012 / 0.0
maternal effect embryo arrest 40, Pentatricopeptide repeat (PPR) superfamily protein (.1)
Lus10040633
82 / 2e-17
AT3G61520 586 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Lus10012328
81 / 4e-17
AT3G53700 1010 / 0.0
maternal effect embryo arrest 40, Pentatricopeptide repeat (PPR) superfamily protein (.1)
Lus10034419
78 / 3e-16
AT1G73400 684 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
PFAM info
Representative CDS sequence
>Potri.008G179201.1 pacid=42806034 polypeptide=Potri.008G179201.1.p locus=Potri.008G179201 ID=Potri.008G179201.1.v4.1 annot-version=v4.1
ATGAAAGGGTATGCTTGCCACCCTTGTGTAATCAGTTACTCCACCATCATCTTATTCTATTGCCGCCAATACAACTTTTATAAGGTTTATGAGCTCCTTG
ATGTGATGAAATGGAAGCACAAGGGTGCCCAACAAATGCTGTTACTTACACCACCATCACTGTTTTTTCTGGCCAAGTCACAAAATTTTGAGGAGGCTTT
ACAACTAGCTCAGAGGATGAAGTCAGCTGAATGTAAACCTGATACGGTTTTTTACAATTCCCTGATGGGAGAGCTGGTCGATTTCAGAAGGCCGTTGATT
TTTTTTGAGGAGATGCCAAATACAGGGGTCTCTCGTGATCCATCTACCTATAACTCCATGATTGCTATGCTCTGTCATGATGGTCAAGTATCAAAGGCCC
TTTCTCTTCTCAAGCAGATGGCAACGTTGGCACATTGCAAGCTTGTTGGTCAGGCGTTCTACCCATTGCTTAAGGCATGCTTTAGAATTGGAGACATGAA
TTTGTTGATTCAATTAATGGATGACATGGTGAAAAAGCATCAGCTAAGTCTTGATAGATCAGTCTATGCTCTTTTAATTCATGGCCTTTGCAGAGCAAAC
AAATGCGAGTGGGCTTGCCATCTGTTCAAGGAAATGATCGGTAAAAATATAGTACCAAAATATCAAACATGTCATTTGCTTTTGGAGGAGGTCAAACTAA
AGAATATGTATGATACTGCTGAGAAAATTGAAGATTTCATGAAAAAATTATAA
AA sequence
>Potri.008G179201.1 pacid=42806034 polypeptide=Potri.008G179201.1.p locus=Potri.008G179201 ID=Potri.008G179201.1.v4.1 annot-version=v4.1
MKGYACHPCVISYSTIILFYCRQYNFYKVYELLDVMKWKHKGAQQMLLLTPPSLFFLAKSQNFEEALQLAQRMKSAECKPDTVFYNSLMGELVDFRRPLI
FFEEMPNTGVSRDPSTYNSMIAMLCHDGQVSKALSLLKQMATLAHCKLVGQAFYPLLKACFRIGDMNLLIQLMDDMVKKHQLSLDRSVYALLIHGLCRAN
KCEWACHLFKEMIGKNIVPKYQTCHLLLEEVKLKNMYDTAEKIEDFMKKL
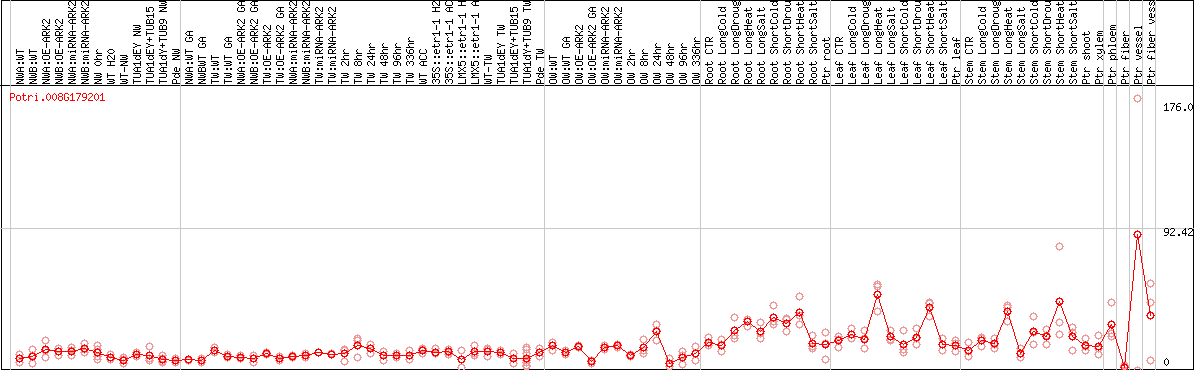
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.008G179201 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.