Potri.008G188150 [POPLAR]

| External link |
|
||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | |||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| ||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| ||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.008G188150.5 pacid=42807122 polypeptide=Potri.008G188150.5.p locus=Potri.008G188150 ID=Potri.008G188150.5.v4.1 annot-version=v4.1
ATGAGTAATAGCAACATATGGCATCATAAATTATTCCACTACCAGAGTTTGCTTCGTCCAGCTAGTTTAGAGCATCGAAGCAAGCTCGATTCTTCTGGAA
TTGTTGCAAAATATTTGTTTAAAGCTGCTCGTAGGTCGTCCAGCTTTTCTGGAAACGATTTGAAACTGAGGAGATCAGAATCAGGCATCGGAGGATCACG
CCGCTCTGTTATTTTCACTCCGCGAGCTGTCTTAGCCATGGAGCCCTCTTCTGAGCAGCTTGCAGGAAAATTCAACCTTGATGGAAATATTGAAATGCAG
GTTTTTGTCAGCAATTCCTCGGCTGCATCCATCGCACAAATAAATATTCAAATCACTTATAGTAGTGATTCATTGCTCCTACACTGGGGTGCAATACTTG
ATAGAAAGGAAAAGTGGGTGCTTCCTTCTCGTCAACCAAATGGAACAACAAATTATAAGAACAGAGCCCTCCGATCTCCTTTTATAAAGTCTGGTTCCAA
TTCTTACATCAGCATAGCGATTGATGATCCTGCAACGCAAGCTATAGAGTTTCTTGTAGTTGATGAAGCAAAGAATAAGTGGTTTAAGGATAATGGCCAG
AACTTTCATATCAAGTTACCCATGAGAGAGAAGTTGATAATTCCTAATGTCTCAGTTCCTGAAGAACTTGTACAGATTCAAGCATATCTAAGGTGGGAAA
GAAAGGGTAAACAAATGTATACCCCTGAGCAAGAGAAGGAGGAATATGAATCTGCTCGCTTTGAGCTATTGGAGGAAGTAGCCAGAGGTACTTCCATTGA
GGATCTCCGAGCAAAACTAACAAACAAAAATGATAGATGTGAAATTAAGGAGCCATCTGTCTCTAAAATAAAAAACAGAATACCGGATGATCTTGTGCAA
ATACAAGCTTATATCAGATGGGAAAAGGCAGGGAAACCCAATTATTCTCCGGAGCAGCAACTGAGGGAATTTGAAAAGGCAAGAGAGGAGTTGCAAACTG
AACTGTACAAAGGTGTCTCTATCGATGAGATTCAGAAGAAGATTACTAAAGGAGAAATCAAGACTGAGGAGTCAAAGCAACTTCAAAACAAAAGGTATTT
TAGCAATGAAAGAATTCAGCGCAAAAAGTGGGACATAATGCAGCTTGTCAACAAACATGCTGCTAAATCTGTAGAGGACAAGGTTTCTAAATCTGCAGAG
GGAAAAGCTTCAGTGGAATCAAAAGTCTTGAAGGCTGTTGAGCTTTTTGCTAAGAAGAAGGAAGAACATGATGGTGGTGCTGTGCTGAACAAGAAAATCT
TTAAGCTTGCTGATAAGGAACTTTTGGTACTTGTAACCAAGCCTGGTGGTAAGGTGAAGGTTCATTTGGCCACTGATTTGGAACAGCCAGTTACTCTTCA
CTGGGCTTTATCTAAAAAGGCTGGAGAATGGTTGGAGCCACCTCCAAATGTGTTGCCTCCTGGTTCGATTTCTCTTCAGGTGGCTGCTGAAACACAGCTT
AAAAATGAGTCCTCTACTGAATTTTCATACCAGGTGCAATCTTTTGAAATGGAGATTGAAAAAGATACTTTCATAGGGATGCCATTTGTCCTTTTGTCTA
ATGGAAGATGGATAAAGAATAATGGCTCAGACTTTTATATTGAATTTGGCAGAGGATCTAAGCATGTCCAAAAGGATGCTGGTGATGGCATAGGTACTGC
CAGGGCGTTGTTGGATAAAATTGCAAAAATGGAGAGTGAGGCACAGAAGTCTTTCATGCACAGATTCAATATTGCGGCAGACTTGATGGATCAAGCCAAG
GATGCTGGGGAATTGGGTCTTGCTGGGATCTTTGTCTGGATGCGCTTTATGGCCACAAGGCAGCTCATATGGAATAAAAACTACAACGTCAAACCACGTG
AAATCAGTAAAGCTCAGGATAGGCTTACAAACTTGGTTCAGGATATTTATGCAAACAATCCACAGCATCGGGAGCTCTTGAGAATGATTATGTCCACTGT
TGGTCGAGGAGGTGAAGGGGATGTGGGACAACGTATTCGGGATGAAATCCTAGTGATACAGAGAAACAATGATTGCAAAGGTGGAATGATGGAGGAATGG
CATCAGAAGTTGCATAATAACACCAGTCCTGATGATGTCATAATCTGCCAGGCATTAATTGATTGCATTAAAAGCGACTTTGACATCAGTGTGTACTGGA
AAACTTTGAATGATAATGGAATAACAAAAGAGAGACTTTTAAGTTATGATCGTGCCATCCATTCCGAACCAAATTTCAGGAGAGATCAAAGGGATGGTCT
CTTGCGTGATCTTGGAAACTATATGAGGACTTTGAAGGCAGTTCATTCAGGTGCAGATCTTGACTCTGCTATTGGAAATTGTATGGGCTACAGATCTGAG
GGTCAAGGCTTCATGGTTGGAGTGCAAACAAATCCCGTTCCAGGCTTGCCATCTGGTTTCCCAGAATTGCTTAAATTTCTACTAGAACACGTTGAAGATA
AGAATGTAGAAGCACTTCTTGAGGGTTTGCTAGAGGCTCGTCAGGAGCTTAGAACTCTTCTATTTAAGTCCAAGGATCGTCTTAAGGATCTTTTATTTTT
GGATATTGCACTTGACTCCACTGTTAGGACAGTTATTGAACGGGGTTATGAGGAATTAAACAATGCTAGACCAGAGAAAATTATGTACTTCATCACTTTG
GTTCTTGAAAATCTCGCACTTTCATCAGATGATAATGAAGATCTTATTTACTGCTTGAAGGGATGGAAACTTGCACTAAGTATGTCAAATAGCAAAAGTG
ACCACTGGGCGTTATTTTCAAAATCAGTCCTTGACAGAACTCGCCTTGCCCTTGCTTGCAAAGCAGAATGGTACCAGAATGTTTTGCAACCATCAGCGGA
GTATCTTGGATCATTGCTTGGAGTGGACCAGTGGGCTCTGAACATATTCACTGAAGAAATCATCCGTGCTGGATCAGCTGCAGCATTATCTACACTTCTT
AATCGACTTGATCCAATTCTTCGGCAGACTGCTCATCTAGGAAGCTGGCAGGTTATTAGCCCAGTTGAAGCTGTTGGATATGTTGTTGCTGTGGATGAGT
TGCTCACTGTGCAGAATAAAACTTACAAGTTCCCAACAATCTTAGTTGCAAAACGAGTAAAAGGAGAAGAAGAAATTCCGGATGGTACAGTTGCTGTGCT
GACACCCGATATGCCTGATGTGCTATCTCATGTTTCTGTGCGAGCAAGAAACAGCAAGGTTTGCTTTGCTACATGCTTTGATCCTGATATTCTGGCCGAC
CTTAAAGCATATGAAGGGAAATTGTTGCATCTGAAACCAACTTCGGCAGATATAGCATATAGTGAACTAAAGGAGGGCAAGCTAGCAGATTCAAGTTCAA
CTAACTTGAAAGAAGTCAGTCCTTCACCTATAAAGTTGGTCAGAAAGCAGTTCAGAGGCAGATATGCCATATCTTCTGAGGAGTTTAGCAGTGAAATGGT
TGGTGCCAAATCACGCAACATCTCATATTTGAAAGGAAAAGTGCCGTCTTGGATCGGGATTCCTACATCAGTTGCTTTACCATTTGGAGTTTTTGAGAAG
GTTCTAAACGAGGATTTGAATTGGGAAGTGGCCAATAAGTTGCAACTGTTGAAGAAAAACTTGGCAGAAGAGTCTAGTGCTCTTGGACAGATTCGCCATA
CAGTTTTACAGCTGGCTGCACCACCACAGTTGGTCCAAGAGCTGAAGACCAAGATGCAAAGTTCTGGGATGCCATGGCCTGGTGACGAAGGTGAACGAAG
ATGGGAGCAAGCATGGATGGCTATAAAGAAGGTCTGGGCTTCAAAATGGAATGAAAGGGCATACTTCAGCACAAGAAAAGTAAAGTTAGACCATGATTAT
CTCTGCATGGCTGTCCTAGTTCAGGAAGTAATAAATGCTGATTATGCATTTGTTATTCATACAACCAATCCATCTTCGGGGGATTCATCTGAGATATATG
CTGAGGTGGTGAAGGGTCTTGGAGAAACTCTAGTTGGAGCTTATCCTGGCCGTGCATTGAGTTTTATCTGCAAGAAAAATGATTTAAACACTCCTCTGGT
GTTGGGTTACCCGAGCAAACCCATTGGGCTTTTTATAAAACGTTCTATTATCTTCCGATCTGACTCCAATGGTGAAGATCTAGAAGGTTATGCCGGTGCA
GGCCTTTATGATAGTGTGCCAATGGATGAAGAAGAGAAGGTTGTGCTCGACTATTCATCTGATCCATTGATCACTGATGATAACTTTCGACGAACAATCC
TCTCTAGCATCGCTCGTGCAGGAAGTGCAATTGAAGAGCTGTATGGATCTCCTCAAGACATTGAAGGTGTAATACGTGATGGTAAGGTCTATGTAGTTCA
GACAAGACCCCAGATGTGA
|
||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.008G188150.5 pacid=42807122 polypeptide=Potri.008G188150.5.p locus=Potri.008G188150 ID=Potri.008G188150.5.v4.1 annot-version=v4.1
MSNSNIWHHKLFHYQSLLRPASLEHRSKLDSSGIVAKYLFKAARRSSSFSGNDLKLRRSESGIGGSRRSVIFTPRAVLAMEPSSEQLAGKFNLDGNIEMQ
VFVSNSSAASIAQINIQITYSSDSLLLHWGAILDRKEKWVLPSRQPNGTTNYKNRALRSPFIKSGSNSYISIAIDDPATQAIEFLVVDEAKNKWFKDNGQ
NFHIKLPMREKLIIPNVSVPEELVQIQAYLRWERKGKQMYTPEQEKEEYESARFELLEEVARGTSIEDLRAKLTNKNDRCEIKEPSVSKIKNRIPDDLVQ
IQAYIRWEKAGKPNYSPEQQLREFEKAREELQTELYKGVSIDEIQKKITKGEIKTEESKQLQNKRYFSNERIQRKKWDIMQLVNKHAAKSVEDKVSKSAE
GKASVESKVLKAVELFAKKKEEHDGGAVLNKKIFKLADKELLVLVTKPGGKVKVHLATDLEQPVTLHWALSKKAGEWLEPPPNVLPPGSISLQVAAETQL
KNESSTEFSYQVQSFEMEIEKDTFIGMPFVLLSNGRWIKNNGSDFYIEFGRGSKHVQKDAGDGIGTARALLDKIAKMESEAQKSFMHRFNIAADLMDQAK
DAGELGLAGIFVWMRFMATRQLIWNKNYNVKPREISKAQDRLTNLVQDIYANNPQHRELLRMIMSTVGRGGEGDVGQRIRDEILVIQRNNDCKGGMMEEW
HQKLHNNTSPDDVIICQALIDCIKSDFDISVYWKTLNDNGITKERLLSYDRAIHSEPNFRRDQRDGLLRDLGNYMRTLKAVHSGADLDSAIGNCMGYRSE
GQGFMVGVQTNPVPGLPSGFPELLKFLLEHVEDKNVEALLEGLLEARQELRTLLFKSKDRLKDLLFLDIALDSTVRTVIERGYEELNNARPEKIMYFITL
VLENLALSSDDNEDLIYCLKGWKLALSMSNSKSDHWALFSKSVLDRTRLALACKAEWYQNVLQPSAEYLGSLLGVDQWALNIFTEEIIRAGSAAALSTLL
NRLDPILRQTAHLGSWQVISPVEAVGYVVAVDELLTVQNKTYKFPTILVAKRVKGEEEIPDGTVAVLTPDMPDVLSHVSVRARNSKVCFATCFDPDILAD
LKAYEGKLLHLKPTSADIAYSELKEGKLADSSSTNLKEVSPSPIKLVRKQFRGRYAISSEEFSSEMVGAKSRNISYLKGKVPSWIGIPTSVALPFGVFEK
VLNEDLNWEVANKLQLLKKNLAEESSALGQIRHTVLQLAAPPQLVQELKTKMQSSGMPWPGDEGERRWEQAWMAIKKVWASKWNERAYFSTRKVKLDHDY
LCMAVLVQEVINADYAFVIHTTNPSSGDSSEIYAEVVKGLGETLVGAYPGRALSFICKKNDLNTPLVLGYPSKPIGLFIKRSIIFRSDSNGEDLEGYAGA
GLYDSVPMDEEEKVVLDYSSDPLITDDNFRRTILSSIARAGSAIEELYGSPQDIEGVIRDGKVYVVQTRPQM
|
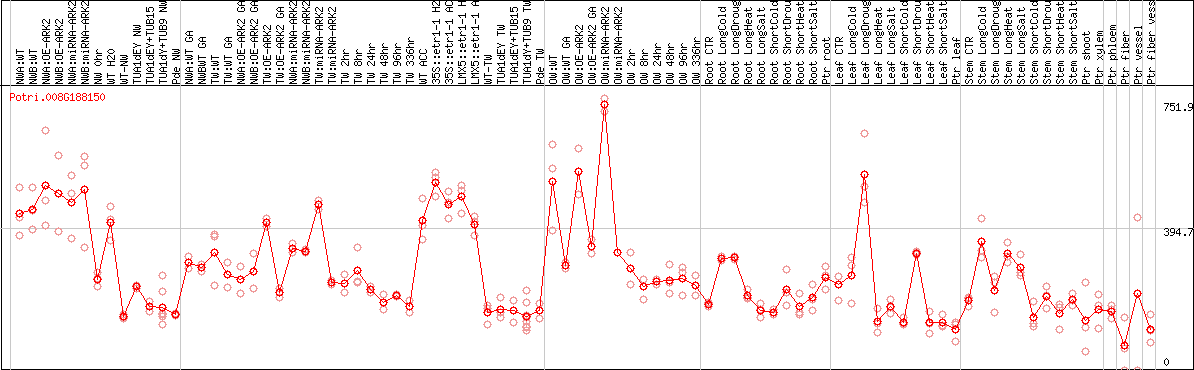
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.008G188150 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
