Potri.008G201600 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||
| PFAM info | ||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.008G201600.3 pacid=42808097 polypeptide=Potri.008G201600.3.p locus=Potri.008G201600 ID=Potri.008G201600.3.v4.1 annot-version=v4.1
ATGGCCATATCTATGCAAAACACCATATGGGATAGTGTTTTAGAGATAACTAAGGTGGCCCAAGAAAAAGGCAGTGATCCTCTAATATGGGCTCTTCAGG
TATCGTCTAGTTTGAGCTCCAGTGGAGTCGGTTTGCCTTCTCCAGAGCTTGCTAATGTCTTGGTTTCTTACATTTTCTGGGATAATAATATGCCCATTTT
GTGGAAGCTTCTTGAGAAGGCCTTGGCTCTCAGGATTGTGCCTCCTTTGATGGTTCTTGCACTTCTTTCTGACAGGGTTGTTCCATGCAGACGGTCTCGG
CCAGTGGCATACAGGCTTTATATGGAACTCCTTAAAACATTTGCTTTTGCATTGAAGGGCCAAATTAATGTGCCAAACTATGAGATGGTCATGAAATCAA
TAGATGGTGTTCTTCATCTTTCCCATAATTTTGGTCTCGAAGCTACCAGTCCTGGGATTCTGGTGGTTGAGTTTCTCTATTCAATCGTGTCACAGTTGTT
AGATGCCTCATTGGATGATGAAGGCTTGTTGGAACTTATACCAGAGATGAAGTCCAGATGGGCAACCAAACCTCAAGAAATGGAAATTGATGCAAATGAT
AATTACAACCAGATGCAAACTGAGTATCACGAGAAATTGTATAAGATGAATACTATAATGGCTATTGAGATGATTGGAAAATTTCTCCAAGATAAATCAA
CTTCAAGGATTCTTGACTTGGTGCGCCAGAACTTTCCCACACATTGGATAAGGTTCTTCCAGAGATTGCAGCTGCTTGGAACAAATTCATCAGCATTAAG
AAATTCAAAAATTCTTACTGCTGAGGATCTTCTGCAGTTGACTACAGGTTCAGGTAGTAACATAGTATTGAGTCGAGAATCGAAGACAAGTTCACTGCAA
AAGTTCCATTCAGTTATGGCTTTTGGTTCTCTAGTGTCTTCCTCTGGTTTGTGTCAAGGAGCTAGTCATTCTGCCCTATGGCTTCCTCTTGATCTTGCAT
TAGAAGATGCCATGGATGGGTATCAAGTTAATGCAACAAGTGCTATTGAAATCATAACCGGTTCAGTTAAAGCCCTTCAAGCAATAAATGGAACCACATG
GCATGAAACCTTTCTAGGCCTTTGGGTTGCAGCTCTCCGTCTTGTTCAAAGGGAAAGGGAACCCATTGAGGGTCCTATCCCTCGATTGGATGCTCGCTTA
TGCATTTTACTATCCATCACAACTCTAGTGGTTGCTGATCTTATTGCTGAAGATGAGAACACACCTATTGATGAATCAGAATGTGGCTCAACAAATCACT
GGAAAGAAAAAAAACTTTCAGGAAAACGCCGCATTGATTTAGTGTCCAGCTTACAGTTATTGGGTGATTATCAGACATTGCTGAGTCCACCTCAATCTGT
TGTTTCTTCAGCCAATCAGGCTGTTGCGAAAGCCATGTTATTTGTTTCAGGCATCAATGTTGGGAGTACATACTCTGAGTGCATCAGCATGAAAGACCTG
CCGATAAACTGCTGTTTCCTAATTGTAGGTCTTGTGCTCTTACTCTCAGCTGGAAACATGAGACATCTGATTGTTGAGGCTTGCATTGCAAGAGGTCTAT
TGGACACGTCAGCATATTTCTGGCCTGGGTATGTCAATGGATGCATCAACCAAATACCTCACAGTATGCCAGCTCAGGTGCCTGGTTGGTCATCATTCAT
GAAGGGGGTCCCACTTTCTCTGTCGATGGTCAATGCTCTGGTTTCAAGTCCCGCTTCAAGCTTAGCAGAGCTCGAGAAAATATTTGAGCTTGCGGTCAAG
GGATCAGATGATGAGAAGATATCTGCTGCTACAGTTCTTTGCGGGGCTTCTCTTCTCCGGGGTTGGAATATACAGGAGCACACTGCTCATTTCATTACTA
GATTATTGTCTCCTCCGGTTCCTGCAGAATATTCTGGGAGTGAGAGCCATTTGATACGTTACGCTCCAATCCTGAATGTACTAATTGTGGGAATAGCAAC
TGTGGATTGTGTTCAGATTTTCTCCCTACATGGCTTGGTTCCACAACTTGCATGCTCGTTGATGCCCATCTGTGAGGTTTTTGGTTCATGCGTGCCTGAT
GTCTCATGGACTCTGCCCACAGGAGAAGACATCTCTGCCCATGCTGTGTTTTCGAATGCATTTGCTCTTCTTTTGAAGCTATGGAGGTTTAATCATCCTC
CTCTTGAACGTGGAGTGGGAGATGTACCCACAGTTGGATCCCAGCTAACTCCTGAATACCTCCTATCAGTACGAAACTCCCACCTAGTCTCTTCTGGAAA
TGTCCTCAAGGATCAGAACAAGAGGAGACTTTCAGCAGTTGCGACTTCGTCATCTGCACAGCCCATATTTTTGGACTCATTTCCCAAATTGAAAGTCTGG
TATCGACAGCATCAAAAATGTCTTGCTGCAACTCTCTCTGATCTTGTTCATGGAACTCCTGTTCATCAGATTGTCAACGTGCTTCTTAACATGATGTTCA
GAAAGATTAATAGAGGAAGCCAGTCATTAACTACCGTTACTTCTGTAAGTAGTGGTTCATCTGGACCTGGAACTGATGATTCTACTCCAAGACCTAAATT
ACCAGCTTGGGATATCTTGGAAGCTGTTCCTTTTGTGGTTGATGCTGCTTTGACTGCCTGTGCTCATGGGAGACTTTCTCCTCGTGAACTGGCTACAGGC
CTTAAAGATTTAGCTGATTTTCTTCCCGCATCTTTGGCAACCATTGTGAGCTACTTCTCAGCTGAAGTGAGCCGTGGAGTGTGGAAACCAGTATTTATGA
ATGGTACAGACTGGCCCAGTCCTGCAGCAAATTTGTCCATTGTCGAGGAGAAGATCAAGAAAATATTAGCTGCTACTGGTGTTGATGTCCCAAGCCTTGC
TGCAGGTGTGAGCTCTCTGGCAACAATTCCACTGCCCCTGGCTGCTTTTGTGAGCCTCACCATAACCTATAAAATTGACAAAGCCTCAGAACGATTTCTC
AATTTGGCTGGTCCAGCATTGGAATCCCTTGCAGCTGGTTGTCCTTGGCCTTGCATGCCAATTGTGGCGTCTCTGTGGACCCAAAAGGCAAAGCGATGGT
TTGACTTCCTTGTGTTTTCTGCATCCCGTACCGTCTTCCTTCACAATAATGATGCAGTGTTTCAACTTCTTAAGAGCTGCTTCTCTGCGACACTTGGTCC
GAATGCTGCAGCTATCTCAAGCAATGGTGGTGTTGGAGCTCTTCTTGGCCATGGGTTTGGTTCCCATTTTAGTGGGGGGATTTCCCCAGTTGCTCCAGGA
ATTCTCTATCTGCGTGTTTACCGAAGCATCAGAGATATTGTATCGCTAATGGAAGATATCATTTCACTCATGATGCTTTCTGTGAGAGAAATAGCATGCA
CTGGCCTTCCAAGGGAGAGGTTAGAGAAGTTGAAGAGATCAAAAAATGGACTGAGGTGTGGACAGTTTTCGCTCACCGCAGCAATGACCCGAGTGAAACT
TGCAGCTTCACTCGGGGCCTCTTTAATATGGTTATCTGGTGGCTTGGGTTTGGTTCAGGCATTGTTTAAAGAAACTTTACCATCTTGGTTTATAGCTGTT
CACAGGTCTGAGCAGGAAGAAGGGTCCAAGGGGATGGTCGCTATGCTTGGGGGTTATGCCCTGGCATTCTTTTCAGTCCATTGCGGGGCACTGGCATGGG
GAGTTGACTCATCATCAAAACGCCGTCCCAAAGTACTTGGTGTGCACATGGAATTTCTTGCGAGTGCACTTGATGGGAAAATATCACTTGGCTGTGATTG
CACTACTTGGCGGGCCTACGTGTCTGGTTTTGTGAGCCTAATGGTGGGCTGCACGCCTTCCTGGGTGCTAGAGGTGGATGCTGATGTTTTGAAGAGACTT
AGTAAAGGATTGAGGCAGTGGAATGAGAAGGATCTTGCCCTGGCTTTGCTGGAAACTGGTGGGGTGGAGACTATGGGTGAGGCTGCTGAACTGATAATCG
AAGATCAATGA
|
|||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.008G201600.3 pacid=42808097 polypeptide=Potri.008G201600.3.p locus=Potri.008G201600 ID=Potri.008G201600.3.v4.1 annot-version=v4.1
MAISMQNTIWDSVLEITKVAQEKGSDPLIWALQVSSSLSSSGVGLPSPELANVLVSYIFWDNNMPILWKLLEKALALRIVPPLMVLALLSDRVVPCRRSR
PVAYRLYMELLKTFAFALKGQINVPNYEMVMKSIDGVLHLSHNFGLEATSPGILVVEFLYSIVSQLLDASLDDEGLLELIPEMKSRWATKPQEMEIDAND
NYNQMQTEYHEKLYKMNTIMAIEMIGKFLQDKSTSRILDLVRQNFPTHWIRFFQRLQLLGTNSSALRNSKILTAEDLLQLTTGSGSNIVLSRESKTSSLQ
KFHSVMAFGSLVSSSGLCQGASHSALWLPLDLALEDAMDGYQVNATSAIEIITGSVKALQAINGTTWHETFLGLWVAALRLVQREREPIEGPIPRLDARL
CILLSITTLVVADLIAEDENTPIDESECGSTNHWKEKKLSGKRRIDLVSSLQLLGDYQTLLSPPQSVVSSANQAVAKAMLFVSGINVGSTYSECISMKDL
PINCCFLIVGLVLLLSAGNMRHLIVEACIARGLLDTSAYFWPGYVNGCINQIPHSMPAQVPGWSSFMKGVPLSLSMVNALVSSPASSLAELEKIFELAVK
GSDDEKISAATVLCGASLLRGWNIQEHTAHFITRLLSPPVPAEYSGSESHLIRYAPILNVLIVGIATVDCVQIFSLHGLVPQLACSLMPICEVFGSCVPD
VSWTLPTGEDISAHAVFSNAFALLLKLWRFNHPPLERGVGDVPTVGSQLTPEYLLSVRNSHLVSSGNVLKDQNKRRLSAVATSSSAQPIFLDSFPKLKVW
YRQHQKCLAATLSDLVHGTPVHQIVNVLLNMMFRKINRGSQSLTTVTSVSSGSSGPGTDDSTPRPKLPAWDILEAVPFVVDAALTACAHGRLSPRELATG
LKDLADFLPASLATIVSYFSAEVSRGVWKPVFMNGTDWPSPAANLSIVEEKIKKILAATGVDVPSLAAGVSSLATIPLPLAAFVSLTITYKIDKASERFL
NLAGPALESLAAGCPWPCMPIVASLWTQKAKRWFDFLVFSASRTVFLHNNDAVFQLLKSCFSATLGPNAAAISSNGGVGALLGHGFGSHFSGGISPVAPG
ILYLRVYRSIRDIVSLMEDIISLMMLSVREIACTGLPRERLEKLKRSKNGLRCGQFSLTAAMTRVKLAASLGASLIWLSGGLGLVQALFKETLPSWFIAV
HRSEQEEGSKGMVAMLGGYALAFFSVHCGALAWGVDSSSKRRPKVLGVHMEFLASALDGKISLGCDCTTWRAYVSGFVSLMVGCTPSWVLEVDADVLKRL
SKGLRQWNEKDLALALLETGGVETMGEAAELIIEDQ
|
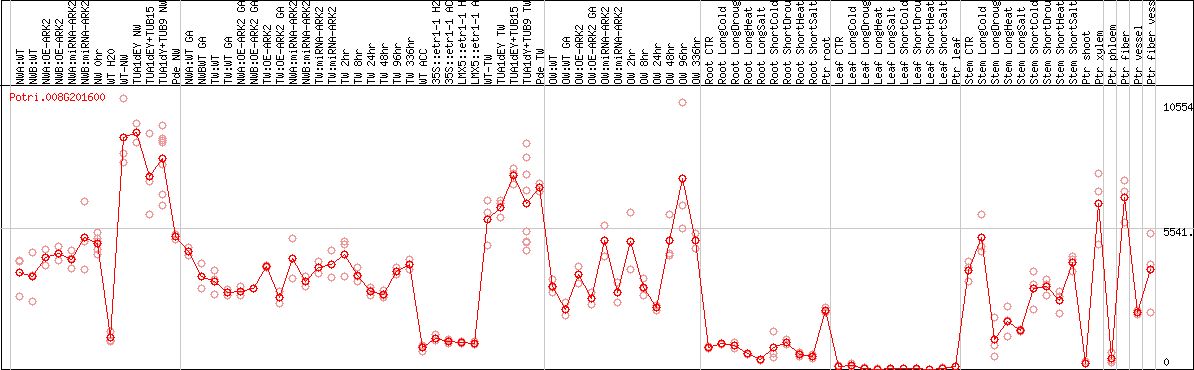
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.008G201600 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
