External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G28160 236 / 8e-76
bHLH
ATFIT1, ATBHLH29, bHLH029, FIT1, ATBHLH029, FRU
ARABIDOPSIS FE-DEFICIENCY INDUCED TRANSCRIPTION FACTOR 1, BASIC HELIX-LOOP-HELIX PROTEIN 29, FER-like regulator of iron uptake (.1)
AT5G57150 76 / 7e-16
bHLH
bHLH035
basic helix-loop-helix (bHLH) DNA-binding superfamily protein
AT4G29930 74 / 3e-15
bHLH
bHLH027
basic helix-loop-helix (bHLH) DNA-binding superfamily protein
AT2G16910 76 / 6e-15
bHLH
AMS, bHLH021
ABORTED MICROSPORES, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
AT1G01260 58 / 5e-09
bHLH
bHLH013, INU4
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.2), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.3)
AT2G46510 56 / 2e-08
bHLH
ATAIB, bHLH107, bHLH017, INU1, JAM1, MYL1
ABA-inducible BHLH-type transcription factor (.1)
AT5G46760 56 / 2e-08
bHLH
bHLH005, ATR2, MYC3
Basic helix-loop-helix (bHLH) DNA-binding family protein (.1)
AT1G12860 56 / 2e-08
bHLH
SCRM2, ICE2, bHLH033
SCREAM 2, INDUCER OF CBF EXPRESSION 2, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
AT5G10570 55 / 2e-08
bHLH
bHLH061
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
AT1G10610 54 / 7e-08
bHLH
bHLH090
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.018G141800
71 / 7e-14
AT5G57150 256 / 6e-86
basic helix-loop-helix (bHLH) DNA-binding superfamily protein
Potri.018G141700
70 / 1e-13
AT5G57150 205 / 4e-66
basic helix-loop-helix (bHLH) DNA-binding superfamily protein
Potri.018G141600
68 / 5e-13
AT5G57150 191 / 2e-60
basic helix-loop-helix (bHLH) DNA-binding superfamily protein
Potri.006G074700
68 / 5e-13
AT5G57150 175 / 2e-54
basic helix-loop-helix (bHLH) DNA-binding superfamily protein
Potri.009G136300
69 / 7e-13
AT2G16910 461 / 1e-156
ABORTED MICROSPORES, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Potri.006G074900
68 / 8e-13
AT5G57150 200 / 3e-64
basic helix-loop-helix (bHLH) DNA-binding superfamily protein
Potri.001G142200
64 / 4e-11
AT1G32640 604 / 0.0
JASMONATE INSENSITIVE 1, Basic helix-loop-helix (bHLH) DNA-binding family protein (.1)
Potri.018G141500
60 / 4e-10
AT5G57150 214 / 1e-69
basic helix-loop-helix (bHLH) DNA-binding superfamily protein
Potri.008G189600
61 / 5e-10
AT1G10610 223 / 3e-67
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10021416
199 / 1e-63
AT2G28160 191 / 7e-61
ARABIDOPSIS FE-DEFICIENCY INDUCED TRANSCRIPTION FACTOR 1, BASIC HELIX-LOOP-HELIX PROTEIN 29, FER-like regulator of iron uptake (.1)
Lus10016151
183 / 5e-55
AT2G28160 235 / 1e-75
ARABIDOPSIS FE-DEFICIENCY INDUCED TRANSCRIPTION FACTOR 1, BASIC HELIX-LOOP-HELIX PROTEIN 29, FER-like regulator of iron uptake (.1)
Lus10006951
66 / 3e-12
AT5G57150 243 / 2e-81
basic helix-loop-helix (bHLH) DNA-binding superfamily protein
Lus10030880
66 / 1e-11
AT2G16910 224 / 7e-69
ABORTED MICROSPORES, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Lus10030609
64 / 6e-11
AT2G16910 239 / 1e-74
ABORTED MICROSPORES, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Lus10000037
58 / 1e-09
AT4G17880 300 / 3e-99
Basic helix-loop-helix (bHLH) DNA-binding family protein (.1)
Lus10026278
57 / 1e-08
AT2G16910 233 / 2e-69
ABORTED MICROSPORES, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Lus10030970
56 / 2e-08
AT1G32640 630 / 0.0
JASMONATE INSENSITIVE 1, Basic helix-loop-helix (bHLH) DNA-binding family protein (.1)
Lus10030243
56 / 2e-08
AT1G01260 505 / 2e-173
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.2), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.3)
Lus10042395
56 / 3e-08
AT2G16910 323 / 1e-103
ABORTED MICROSPORES, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF00010
HLH
Helix-loop-helix DNA-binding domain
Representative CDS sequence
>Potri.009G005600.1 pacid=42771436 polypeptide=Potri.009G005600.1.p locus=Potri.009G005600 ID=Potri.009G005600.1.v4.1 annot-version=v4.1
ATGGCCCTCCCAATTCCCATCAATTTTTACTTGAAAGAGACCTCTCTTTTTCAAGGAATGGATAGGATGGATGATCCAACTGGAAACCCTTTAGCAGCAC
AGACTAATTTTCAGTTTCAGTTGCAAGATTTCATTGATGAAGCCAACTTCGATCGATACATTGATCTAATTCGAGGAGAGAATGAAATCACTGCTTTCGA
TTGTGATCTTATCAATGGCTTTTTGGTCGATAACCAATTTGGTCTGTCCACAGGAGATAAATTTGATTGTGATCTCATCAACCATGTTCCTACACACACT
TCTAGTGCAATGGAACAGGATCCCAATTACGTCCCCATTGCATTGCCGAGTTTTGACGGGGACATGGGATTAGAGGCTGAAGAAGATACTGATGAGGAAG
ACTCTTCTGGAACCGCCACAACGACTAAGAAGACGAAGAAGGATCGATCAAGGACATTGATTTCGGAGCGAAGGAGGAGGGGTAGGATGAAGGAGAAGCT
CTATGCTTTGCGGTCCTTGGTTCCTAATATAACTAAGATGGACAAGGCCTCCATAATTGGGGATGCCGTATTATTTGTGCAAGAGCTGCAAATGCAAGCC
AATAAGCTAAAAGCTGATATTGCCAGCCTTGAATCATCCTTAATAGGATCAGATAGATATCAAGGGTCAAATAGAAATCCAAAGAATCTTCAAAATACAA
GTAACAACCATCCAATTCGCAAGAAGATCATCAAGATGGATGTGTTTCAAGTGGAAGAAAGAGGGTTTTACGTGAGATTGGTGTGCAACAAGGGAGAAGG
CGTTGCACCCTCGCTCTATAGGGCTCTCGAGTCCCTTACAAGCTTCAGTGTTCAAAACTCAAACTTGGCTACAACTTCTGAGGGGTTTGTATTAACATTT
ACTTTAAATGTTAAAGAAAGTGAGCAGGACATGAACTTGCCAAATTTGAAGCTATGGGTTACTGGGGCTCTTTTGAATCAAGGATTTGAACTGTTAACGG
CTTAA
AA sequence
>Potri.009G005600.1 pacid=42771436 polypeptide=Potri.009G005600.1.p locus=Potri.009G005600 ID=Potri.009G005600.1.v4.1 annot-version=v4.1
MALPIPINFYLKETSLFQGMDRMDDPTGNPLAAQTNFQFQLQDFIDEANFDRYIDLIRGENEITAFDCDLINGFLVDNQFGLSTGDKFDCDLINHVPTHT
SSAMEQDPNYVPIALPSFDGDMGLEAEEDTDEEDSSGTATTTKKTKKDRSRTLISERRRRGRMKEKLYALRSLVPNITKMDKASIIGDAVLFVQELQMQA
NKLKADIASLESSLIGSDRYQGSNRNPKNLQNTSNNHPIRKKIIKMDVFQVEERGFYVRLVCNKGEGVAPSLYRALESLTSFSVQNSNLATTSEGFVLTF
TLNVKESEQDMNLPNLKLWVTGALLNQGFELLTA
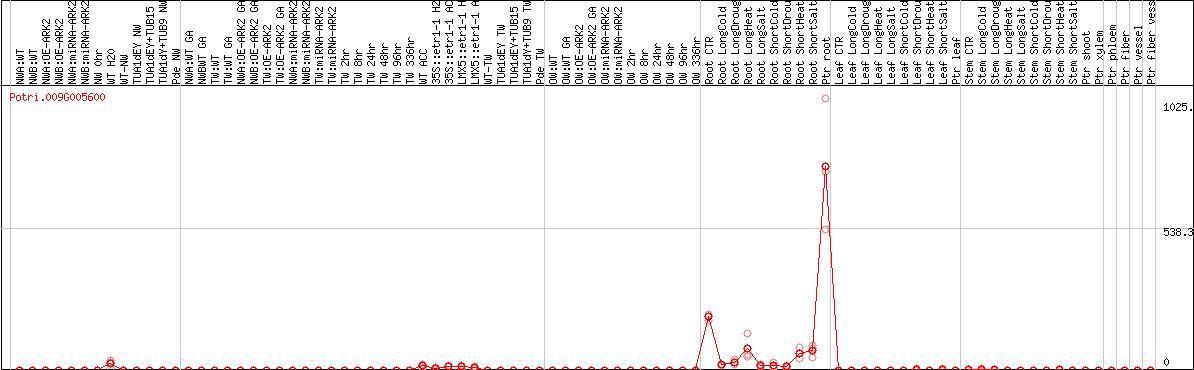
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.009G005600 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.