External link
Symbol
FLA14.12
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G60490 168 / 3e-51
FLA12
FASCICLIN-like arabinogalactan-protein 12 (.1)
AT5G03170 159 / 1e-47
ATFLA11, FLA11, IRX13
ARABIDOPSIS FASCICLIN-LIKE ARABINOGALACTAN-PROTEIN 11, FASCICLIN-like arabinogalactan-protein 11 (.1)
AT5G44130 113 / 6e-30
FLA13
FASCICLIN-like arabinogalactan protein 13 precursor (.1)
AT1G03870 105 / 4e-27
FLA9
FASCICLIN-like arabinoogalactan 9 (.1)
AT2G20520 96 / 2e-23
FLA6
FASCICLIN-like arabinogalactan 6 (.1)
AT2G04780 74 / 1e-15
FLA7
FASCICLIN-like arabinoogalactan 7 (.1.2)
AT3G46550 67 / 2e-12
FLA4, SOS5
salt overly sensitive 5, fasciclin-like arabinogalactan-protein 4, Fasciclin-like arabinogalactan family protein (.1)
AT3G60900 56 / 9e-09
FLA10
FASCICLIN-like arabinogalactan-protein 10 (.1)
AT2G45470 54 / 3e-08
AGP8, FLA8
FASCICLIN-like arabinogalactan protein 8 (.1)
AT5G55730 49 / 1e-06
FLA1
FASCICLIN-like arabinogalactan 1 (.1.2)
Paralogs
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10036113
201 / 1e-63
AT5G03170 173 / 6e-53
ARABIDOPSIS FASCICLIN-LIKE ARABINOGALACTAN-PROTEIN 11, FASCICLIN-like arabinogalactan-protein 11 (.1)
Lus10036112
194 / 4e-61
AT5G03170 197 / 1e-62
ARABIDOPSIS FASCICLIN-LIKE ARABINOGALACTAN-PROTEIN 11, FASCICLIN-like arabinogalactan-protein 11 (.1)
Lus10002985
192 / 1e-60
AT5G60490 180 / 4e-56
FASCICLIN-like arabinogalactan-protein 12 (.1)
Lus10002978
188 / 5e-59
AT5G03170 209 / 1e-67
ARABIDOPSIS FASCICLIN-LIKE ARABINOGALACTAN-PROTEIN 11, FASCICLIN-like arabinogalactan-protein 11 (.1)
Lus10036111
185 / 6e-58
AT5G03170 209 / 1e-67
ARABIDOPSIS FASCICLIN-LIKE ARABINOGALACTAN-PROTEIN 11, FASCICLIN-like arabinogalactan-protein 11 (.1)
Lus10002984
185 / 1e-57
AT5G03170 192 / 1e-60
ARABIDOPSIS FASCICLIN-LIKE ARABINOGALACTAN-PROTEIN 11, FASCICLIN-like arabinogalactan-protein 11 (.1)
Lus10036114
172 / 2e-52
AT5G03170 191 / 3e-60
ARABIDOPSIS FASCICLIN-LIKE ARABINOGALACTAN-PROTEIN 11, FASCICLIN-like arabinogalactan-protein 11 (.1)
Lus10026499
162 / 6e-49
AT5G03170 291 / 5e-100
ARABIDOPSIS FASCICLIN-LIKE ARABINOGALACTAN-PROTEIN 11, FASCICLIN-like arabinogalactan-protein 11 (.1)
Lus10019929
162 / 9e-49
AT5G03170 295 / 2e-101
ARABIDOPSIS FASCICLIN-LIKE ARABINOGALACTAN-PROTEIN 11, FASCICLIN-like arabinogalactan-protein 11 (.1)
Lus10002979
140 / 6e-41
AT5G60490 143 / 1e-42
FASCICLIN-like arabinogalactan-protein 12 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF02469
Fasciclin
Fasciclin domain
Representative CDS sequence
>Potri.009G012200.1 pacid=42772757 polypeptide=Potri.009G012200.1.p locus=Potri.009G012200 ID=Potri.009G012200.1.v4.1 annot-version=v4.1
ATGAGGCCACAATCTTTCATCTTAGCATTGTCTCTTATCTTTTTCTTCCTCCATTGTACAAAAACGTTGTGCCAGTCACCTGCCGCGGCCCCGGCTATGG
CGCCCCCTAAGACACCAGTGAAGGCACCGCCTGCAGACTCCTCACAGGCACCATCTGCACAGGTTGCTACATCACCTGGCCCCGTTGACGTCAATAAAAT
CTTGCAGAAGGCTGGCCACTTCACAGTCTTCGCCCGCCTGATGCAAGCTACAACAGAAGACACCGAGTTAAACAAGGAGCTGAACACCACAAACAATGGA
ATAACAATCCTGGCACCAACTGACAACGCATTTTCGAGCCTCAAAGCAGGCTTTCTCAACTCTCTAAGCGATGAAGACAAGACTGAGCTGGTGAAGTTTC
ACGTGCTGCCTGCATTTATATCAACATCCCAATTTCAGACTGTGAGTAACCCTGTAAGGACACAGGCGGGGACAGGTCCCAGGGTAACACTAAATGTCAC
CACCACCGGGAATTTCGTGAATATAAGTTCAGGGCTGACAAATACAAGCATATCAGGCACTGTATACACTGACAGCCAGCTTGCTATTTATCAACTTGAT
AAGGTACTATTTCCTTTGGACATATTTACTCCTAAGCCTCCAGCCCCTGCTCCCGAGCCAGCACTGGGAAAGCCTAGGAAAGCAGCACCCGATGCAGAGA
GCCCTACTGCTCCTAAGGATATTTCTGGTGCTCCAGCTCTGCTTTTTCTGCATAACAACGCCTTGCTTCTTGCAGTCAGCTGTGCGTTTGGTGCAATAAT
ACATTCATGA
AA sequence
>Potri.009G012200.1 pacid=42772757 polypeptide=Potri.009G012200.1.p locus=Potri.009G012200 ID=Potri.009G012200.1.v4.1 annot-version=v4.1
MRPQSFILALSLIFFFLHCTKTLCQSPAAAPAMAPPKTPVKAPPADSSQAPSAQVATSPGPVDVNKILQKAGHFTVFARLMQATTEDTELNKELNTTNNG
ITILAPTDNAFSSLKAGFLNSLSDEDKTELVKFHVLPAFISTSQFQTVSNPVRTQAGTGPRVTLNVTTTGNFVNISSGLTNTSISGTVYTDSQLAIYQLD
KVLFPLDIFTPKPPAPAPEPALGKPRKAAPDAESPTAPKDISGAPALLFLHNNALLLAVSCAFGAIIHS
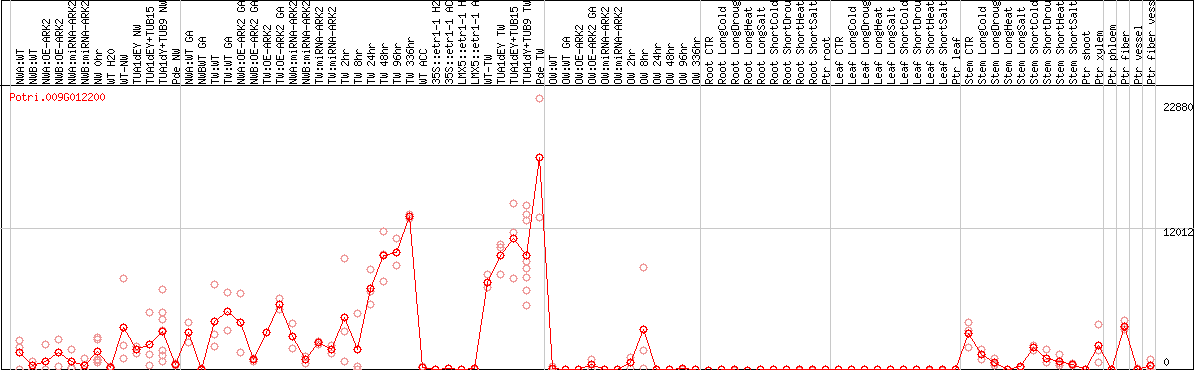
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.009G012200 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.