External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G44350 258 / 9e-87
NAC
ANAC061
NAC domain containing protein 61 (.1.2)
AT5G22380 256 / 3e-86
NAC
ANAC090
NAC domain containing protein 90 (.1)
AT2G17040 139 / 4e-40
NAC
ANAC036
NAC domain containing protein 36 (.1)
AT4G27410 137 / 7e-39
NAC
RD26, ANAC072
NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.1), NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.2), NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.3)
AT3G15500 136 / 2e-38
NAC
ATNAC3, ANAC055
NAC domain containing protein 55, NAC domain containing protein 3 (.1)
AT2G02450 137 / 3e-38
NAC
LOV1, ANAC034, ANAC035
LONG VEGETATIVE PHASE 1, Arabidopsis NAC domain containing protein 34, NAC domain containing protein 35 (.1.2)
AT4G17980 133 / 7e-38
NAC
ANAC071
NAC domain containing protein 71 (.1)
AT5G08790 134 / 8e-38
NAC
ATAF2, ANAC081
Arabidopsis NAC domain containing protein 81, NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.1)
AT1G52890 134 / 9e-38
NAC
ANAC019
NAC domain containing protein 19 (.1)
AT1G33060 139 / 1e-37
NAC
ANAC014
NAC 014 (.1.2)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.001G218800
401 / 3e-143
AT5G22380 272 / 3e-92
NAC domain containing protein 90 (.1)
Potri.016G076100
233 / 7e-77
AT5G22380 214 / 2e-69
NAC domain containing protein 90 (.1)
Potri.016G076000
232 / 1e-76
AT5G22380 226 / 4e-74
NAC domain containing protein 90 (.1)
Potri.006G209200
228 / 9e-75
AT5G22380 241 / 3e-80
NAC domain containing protein 90 (.1)
Potri.003G046700
140 / 2e-39
AT2G02450 318 / 2e-106
LONG VEGETATIVE PHASE 1, Arabidopsis NAC domain containing protein 34, NAC domain containing protein 35 (.1.2)
Potri.009G141600
137 / 4e-39
AT2G17040 314 / 6e-108
NAC domain containing protein 36 (.1)
Potri.008G081500
137 / 2e-38
AT1G54330 316 / 1e-107
NAC domain containing protein 20 (.1)
Potri.001G404100
136 / 3e-38
AT4G27410 370 / 8e-129
NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.1), NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.2), NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.3)
Potri.011G123300
135 / 5e-38
AT4G27410 354 / 2e-122
NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.1), NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.2), NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.3)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10020643
288 / 1e-98
AT5G22380 283 / 1e-96
NAC domain containing protein 90 (.1)
Lus10004846
284 / 6e-97
AT5G22380 275 / 1e-93
NAC domain containing protein 90 (.1)
Lus10029410
176 / 3e-54
AT5G22380 171 / 1e-52
NAC domain containing protein 90 (.1)
Lus10022636
139 / 1e-39
AT2G17040 287 / 6e-97
NAC domain containing protein 36 (.1)
Lus10008419
140 / 1e-38
AT2G02450 327 / 7e-109
LONG VEGETATIVE PHASE 1, Arabidopsis NAC domain containing protein 34, NAC domain containing protein 35 (.1.2)
Lus10003333
136 / 2e-38
AT2G17040 293 / 2e-99
NAC domain containing protein 36 (.1)
Lus10030446
135 / 2e-38
AT1G69490 302 / 2e-103
Arabidopsis NAC domain containing protein 29, NAC-like, activated by AP3/PI (.1)
Lus10003367
140 / 3e-38
AT2G02450 333 / 3e-110
LONG VEGETATIVE PHASE 1, Arabidopsis NAC domain containing protein 34, NAC domain containing protein 35 (.1.2)
Lus10008420
137 / 6e-38
AT2G02450 320 / 2e-106
LONG VEGETATIVE PHASE 1, Arabidopsis NAC domain containing protein 34, NAC domain containing protein 35 (.1.2)
Lus10042731
134 / 6e-38
AT1G01720 414 / 7e-147
Arabidopsis NAC domain containing protein 2, NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF02365
NAM
No apical meristem (NAM) protein
Representative CDS sequence
>Potri.009G019200.2 pacid=42772317 polypeptide=Potri.009G019200.2.p locus=Potri.009G019200 ID=Potri.009G019200.2.v4.1 annot-version=v4.1
ATGGAAGAACTCCCTCTAGGTTACCGCTTCTACCCAACGGAAGAAGAGCTAGTCTCCTTCTACTTGCATAATAAGCTGGAAGGGACGAGACAAGAACTGC
AACGTGTGATTCCAGAGATCAGCATATACGACATCGAGCCGGGGGATCTTCCACAGCTTTCAGGAGAATTATGTCGAGGCAACACGGAGCAGTGGTTCTT
CTTTACACCCAGGCAAGCGAGAGAAGCTAGGGGAGGGCGACCTAATCGGACTACAGCAACCGGGTACTGGAAGGCCACAGGCTCTCCTGGTTATGTTTAC
TCATCGGATAATCGAGTGATTGGATTGAAGAAAACCATGGTGTTCTATATAGGAAAAGCTCCGACTGGAAGAAAAACCAAATGGAAGATGAATGAGTATA
GAGCCATTGAAGTTCATGAATCGTCTAGAAATGCTACTTCAAAGTTGAGGCATGAATTCAGTTTGTGCCGTGTCTATGTGGTATCAGGAAGCTTTCGGGC
ATTTGATAGACGGCCTCTAGAAGCTGTAACAAGAGGGACACAACATCTTCTTAGGGGGACACATGTTGGTGATGCAGCAACTACACCTGCTCAGGACCCT
ATAACTGTGGAGATGACAAGCTCACCTGAAACTTCATACTCGGGAGGAGACCATGTTGATTACCCAGGGACTGCAGCAAGTGCCAACTGGGGGCTGATCC
TGCATGGGAATGACCGGAACAACTGGATTGGCCCAGCACAGATAGATTTGTAA
AA sequence
>Potri.009G019200.2 pacid=42772317 polypeptide=Potri.009G019200.2.p locus=Potri.009G019200 ID=Potri.009G019200.2.v4.1 annot-version=v4.1
MEELPLGYRFYPTEEELVSFYLHNKLEGTRQELQRVIPEISIYDIEPGDLPQLSGELCRGNTEQWFFFTPRQAREARGGRPNRTTATGYWKATGSPGYVY
SSDNRVIGLKKTMVFYIGKAPTGRKTKWKMNEYRAIEVHESSRNATSKLRHEFSLCRVYVVSGSFRAFDRRPLEAVTRGTQHLLRGTHVGDAATTPAQDP
ITVEMTSSPETSYSGGDHVDYPGTAASANWGLILHGNDRNNWIGPAQIDL
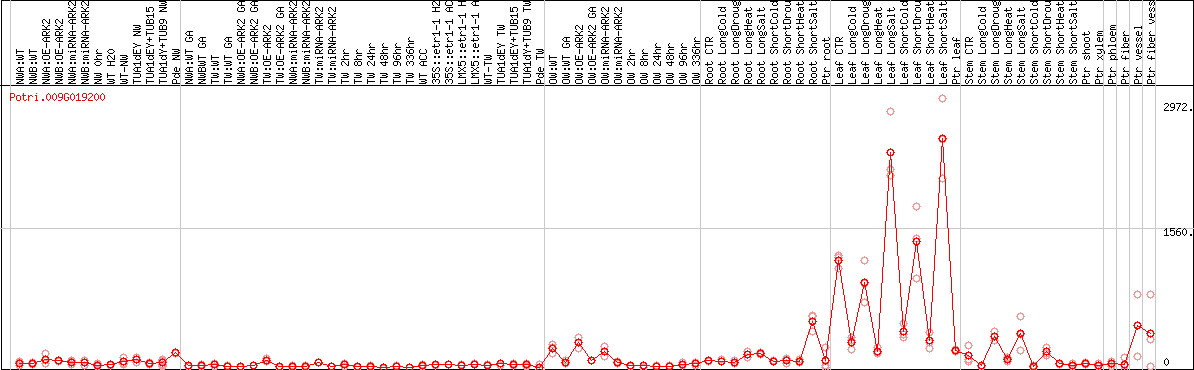
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.009G019200 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.