Potri.009G065300 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.009G065300.4 pacid=42771790 polypeptide=Potri.009G065300.4.p locus=Potri.009G065300 ID=Potri.009G065300.4.v4.1 annot-version=v4.1
ATGGCTGGGAAATCTAATAAAGGGAGGAACCGGAGAGGATCCAATAGCACAACAAATTCTTCAGAGCCAACAGCGTCATCTGATGCTCCTGTGAAAGATG
ACATAACCGCTTCTGAAGCTGCTTCAGAAGTTGGTGTAGCTATTGTTAATGGAGCACCAGCTGGGGGTGAACCTGCTAATGGCACCTCAGAGATAAAGGA
GTCTGAAACAGCAAATTCTACGAGTGAAGCAAAGCAAGGTGATCTTCATCTTTACCCCGTCTCTGTCAAAACACAAAGTAGCGAGAAGCTTGAGTTACAA
TTGAACCCAGGAGATTCCGTCATGGATGTAAGACAATTCCTTCTCGATGCTCCCGAAACATGTTTCTTTACATGCTATGACTTGTTGCTGCACACAAAGG
ATGGTTCAACTCTTCAAATAGAAGATTATAATGAAATATCTGAAGTAGCTGACATTACTTCTGGTGGTTGCTCCTTGGAGATGGTTGCTGCACCTTATGA
TGATAGATCGATCAGGGCTCATGTTCACCGCACAAGAGAATTGCTTTCACTTTCTACACTTCATGCTTCATTATCAACGTCCCTTGCTTTAGAGTACGAA
AAAGCACAAAACAAAGCTCTTGGTTCAGATACTGGAAAAACTGAGGTCCCTGAGCTTGATGGTATGGGATTTATGGAGGATGTTGCAGGTTCGGTTGGTA
AATTGTTGTCATTCCCAGCAAAGGAGATCATGTGTGTGGACAGTATTATATTTTCATCCTTCAATCCTCCCCCAAGCCATAGAAGGCTCGTTGGGGATTT
GATTTATTTGGATGTAATAACATTGGAGGGTAACAAATATTGTATTACTGGAACCACAAAGATGTTCTATGTCAATTCAAGCACAGGGAATGTCCTTGAC
CCAAGGCCAAGTAAAGCTACCTCTGAAGCAACTACCCTTGTTGGACTCCTGCAGAGAATCAGTCCCAAGTTCAAAAAAGCTCTTCGTGAAATTTTGGAGC
ATAAGGGCTCTGCACACCCTTTTGAAAATGTTCAATCTCTCCTGCCACCTTGTTCATGGCTAGGATTATACCCAGTTCCTGACCACAGACGTGATGCGGC
CAGAGCAGAGGATGCACTGACTCTTTCATATGGTAGTGAGCTGATTGGCATGCAAAGAGATTGGAATGAGGAGTTGCAATCCTGTCGGGAGTTTCCCCAT
TCAGCTCCTCAGGAAAGGATTTTGCGTGATAGGGCTCTCTACAAAGTAACTTCGGACTTTGTTGATGCAGCAGTTAAAGGTGCTATTGGAGTCATCAACA
GATGTATTCCCCCCATAAATCCAACTGATCCAGAGTGCTTTCATATGTATGTTCACAACAATATATTCTTCAGTTTTGCTGTGGACGTGGACCTTGAACA
GCTCTCCAAGAAATGTAATTCAGATACTAGTTCAAAGACAGAGAACACCAGTTCTTCAATTAATTCATCTGAAAAGGTCACCAGCGATTTGCATGGGGAT
GGTGGAATTGCCAATGGGGTGAAATCTGATGGATCAACCGTTGAAGTGGTGGAATTGCATCCCGAGTCATCTTCTGAGCCGCAGTTGGCCGAGAGTGAAC
AAGCTACATATGCGTCGGCAAATAATGATCTGAAAGGCACCAAAGCCTACCAGGAAGCTGATGTCCCTGGTTTATATAATCTTGCCATGGCCATAATTGA
TTACAGGGGTCACAGAGTGGTAGCCCAGAGTGTCTTACCTGGCATTCTTCAAGGGGATAAATCAGACTCTCTTCTATATGGTTCTGTTGATAATGGAAAG
AAAATATGCTGGAATGAAGATTTTCATTCTAAGGTAGTGGAAGCTGCTAAACGCCTTCATCTCAAGGAACACACTGTCCTAGATGGATCCGGGGATGCTT
TCAAATTAGCTGCTCCAGTAGAATGCAAGGGCATTGTAGGTAGTGATGACAGGCATTATCTACTAGACTTGATGAGAGCAACTCCTCGTGATGCAAACTA
TACTGGACCTGGCTCGCGGTTTTGCATATTGAGACCAGAATTAATTTCTGCCTTCTGTCAGGCTGAAGCTGTGGCAAGATTGAAGAGTAGACCAAAATCC
GAGGGAGGGGCTCACGTGGCTGCAGATTCTACAGAAGTTACTACTGGTGATGAGCAAGTTAAACCTGAGGAAGCTGCAGCTTCCATCAACAATCAAGAGA
TTGCTAAAGAAGGGAAAGCTGACACTGTAGAAGAAAGTGCTCCTGCTCTTGCTGGAAGCTCTGAATCTTGTGAAGAAATACTTTTTAACCCTAATGTTTT
TACCGAGTTCAAGTTGTCTGGTGATCCGGAGGAGATTGCAGCAGATGAAGAAAATGTGAAGAAAGTGGGTTCATATCTTGCAAATACTGTGCTTCCAAAG
TTCATCCAAGATCTTTGCACACTTGAAGTCTCGCCCATGGATGGACAGACTCTTACCGAAGCACTACATGCTCATGGAATTAATGTTCGCTACATGGGAA
AGGTAGCTGAAGGGATAAAACACTTACCTCATCTATGGGATCTTTGTTCCAATGAGATTGTTGTCCGGTCTGCAAAGCACATCCTCAAGGATCTTCTCAG
GGATACAGATGATAATGATCTAGGGCCAGCAATTTCTCATTTTTTCAACTGTTTCTTTGGAACTTGTCAAGCTGTTGGCATAAAAGTTTCAGCTAATGGT
CCGCATTCAAGAGCAGCAAAGAAAGAACAATCTGGCAATCAGTCTTCAGGAAAATCCTCTAGGGGACAAACAAGGTGGAAAGGAGCATCTGCAAGAAAGA
ACCAATCCTCATATATGAATGTTAGCTCAGAAACTCTATGGTCTGACATTCAGGAACTTGCAGAGCTCAAATATCAGTTTGAGTTACCAGAGGATGCGAG
GTCACAAGTGAAGAAAGTCTCGGTTATTCGGAATCTGTGCCAAAAGATGGGCATTACCATCGCAGCTCGCAAATATGATCTCAATGCTGCAATGCCATTC
CAACTGTCAGATATCTTGAATCTCCAACCCGTGGTAAAGCATTCGGTTCCTCTATGTTCAGAAGCTAAGGATATTGTGGAAACCGGAAAGGTTCAACTAG
CTGAGGGGATGCTTAGTGAAGCCTACACGTCATTTTCCGATGCATTTTCCATTCTTCAACAGGTGACTGGTCCAATGCATCGGGAGGTTGCCAACTGTTG
TCGGTACCTGGCCATGGTTTTGTATCATGCTGGTGATATGGCTGGAGCAATCATCCAACAGCACAAGGAGTTGATCATAAATGAACGTTGCCTGGGATTA
GACCACCCCGACACTGCCCACAGTTATGGAAACATGGCTCTCTTTTACCATGGACTAAACCAAACAGAACTTGCATTGCGACATATGTCCCGTGCGCTGC
TCTTACTGAGTTTGTCTTCTGGGCCAGACCACCCTGATGTGGCTGCAACCTTCATTAATGTTGCAATGATGTACCAGGACATTGGAAAGATGAATACAGC
TCTTCGCTATCTGCAAGAGGCCTTGAAAAAGAACGAGAGGCTTTTAGGAGAGGAACATATTCAAACTGCTGTATGCTACCATGCTCTTGCAATTGCATTT
AACTGTATGGGCGCGTTCAAGCTATCACACCAGCATGAGAAGAAAACCTATGACATACTTGTCAAACAGCTTGGTGAGGAGGACTCAAGGACTCGAGACT
CCCAAAACTGGATGAAGACATTTAAGATGCGAGAACTGCAGAAGCAGAAAGGTCAGGCATTGAATGCAGCTTCTGCTCAAAAGGCGATCGATATATTGAA
GGCTAATCCTGACCTGTTGCATGCATTCCAAGCTGCTGCGGTTGCTGGAGGATCTGGATCTGGGAGTACCAGTGGTTCCATGAACAAATCCCTTAATGCA
GCTATTGTTGGTGAGACACTTCCACGGGGAAGGGGAGTAGATGAACGAGCTGCTCGTGCAGCAGCAGAAGTTAGGAAGAAAGCAGCTGCTAGGGGGCTCT
TGACTCGTCCACATGGTGTTCCTGTTCAAGCTTTGCCACCTCTCACTCAGCTCCTCAACATCATTAATTCAGGGGCAACTCCAGATACAGTCAACAACGA
GGAAGCAGCAGGTGGAGTGGAGGAAACTAATGGACAATCTTCTAATGACCCAGTAGATACTCAAAAAGACCAAACATCAGGAGGGCAAGATCAGGCTCCT
GTTGGATTGGGCAAAGGCTTAGCATCATTAGATGCCAAGAAACAGAAAGCCAAAGCCAAAGTTGCAGCTTAA
|
|||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.009G065300.4 pacid=42771790 polypeptide=Potri.009G065300.4.p locus=Potri.009G065300 ID=Potri.009G065300.4.v4.1 annot-version=v4.1
MAGKSNKGRNRRGSNSTTNSSEPTASSDAPVKDDITASEAASEVGVAIVNGAPAGGEPANGTSEIKESETANSTSEAKQGDLHLYPVSVKTQSSEKLELQ
LNPGDSVMDVRQFLLDAPETCFFTCYDLLLHTKDGSTLQIEDYNEISEVADITSGGCSLEMVAAPYDDRSIRAHVHRTRELLSLSTLHASLSTSLALEYE
KAQNKALGSDTGKTEVPELDGMGFMEDVAGSVGKLLSFPAKEIMCVDSIIFSSFNPPPSHRRLVGDLIYLDVITLEGNKYCITGTTKMFYVNSSTGNVLD
PRPSKATSEATTLVGLLQRISPKFKKALREILEHKGSAHPFENVQSLLPPCSWLGLYPVPDHRRDAARAEDALTLSYGSELIGMQRDWNEELQSCREFPH
SAPQERILRDRALYKVTSDFVDAAVKGAIGVINRCIPPINPTDPECFHMYVHNNIFFSFAVDVDLEQLSKKCNSDTSSKTENTSSSINSSEKVTSDLHGD
GGIANGVKSDGSTVEVVELHPESSSEPQLAESEQATYASANNDLKGTKAYQEADVPGLYNLAMAIIDYRGHRVVAQSVLPGILQGDKSDSLLYGSVDNGK
KICWNEDFHSKVVEAAKRLHLKEHTVLDGSGDAFKLAAPVECKGIVGSDDRHYLLDLMRATPRDANYTGPGSRFCILRPELISAFCQAEAVARLKSRPKS
EGGAHVAADSTEVTTGDEQVKPEEAAASINNQEIAKEGKADTVEESAPALAGSSESCEEILFNPNVFTEFKLSGDPEEIAADEENVKKVGSYLANTVLPK
FIQDLCTLEVSPMDGQTLTEALHAHGINVRYMGKVAEGIKHLPHLWDLCSNEIVVRSAKHILKDLLRDTDDNDLGPAISHFFNCFFGTCQAVGIKVSANG
PHSRAAKKEQSGNQSSGKSSRGQTRWKGASARKNQSSYMNVSSETLWSDIQELAELKYQFELPEDARSQVKKVSVIRNLCQKMGITIAARKYDLNAAMPF
QLSDILNLQPVVKHSVPLCSEAKDIVETGKVQLAEGMLSEAYTSFSDAFSILQQVTGPMHREVANCCRYLAMVLYHAGDMAGAIIQQHKELIINERCLGL
DHPDTAHSYGNMALFYHGLNQTELALRHMSRALLLLSLSSGPDHPDVAATFINVAMMYQDIGKMNTALRYLQEALKKNERLLGEEHIQTAVCYHALAIAF
NCMGAFKLSHQHEKKTYDILVKQLGEEDSRTRDSQNWMKTFKMRELQKQKGQALNAASAQKAIDILKANPDLLHAFQAAAVAGGSGSGSTSGSMNKSLNA
AIVGETLPRGRGVDERAARAAAEVRKKAAARGLLTRPHGVPVQALPPLTQLLNIINSGATPDTVNNEEAAGGVEETNGQSSNDPVDTQKDQTSGGQDQAP
VGLGKGLASLDAKKQKAKAKVAA
|
DESeq2's median of ratios [POPLAR]
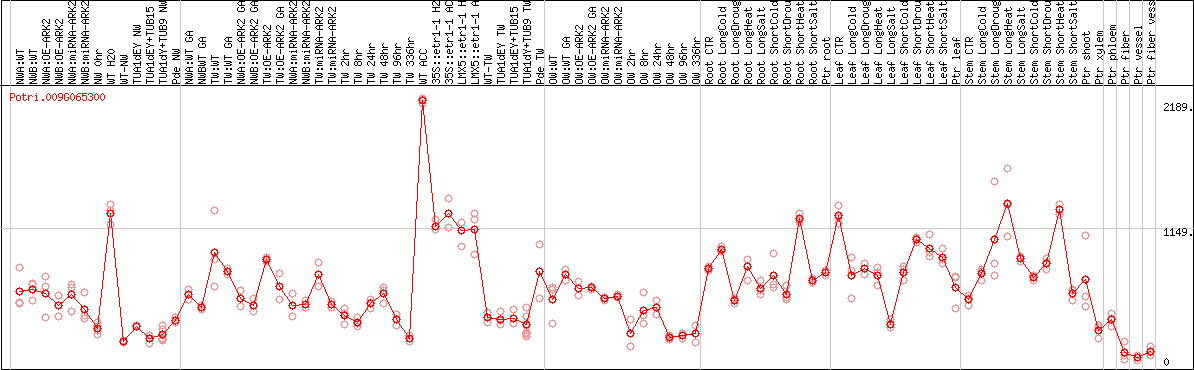
Coexpressed genes
Potri.009G065300 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
