External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G12320 197 / 6e-66
ankyrin repeat family protein (.1)
AT4G35450 81 / 1e-18
AKR2A, AFT, AKR2
ankyrin repeat-containing protein 2 (.1.2.3.4.5)
AT2G17390 73 / 7e-16
AKR2B
ankyrin repeat-containing 2B (.1)
AT4G27780 67 / 9e-14
ACBP2
acyl-CoA binding protein 2 (.1)
AT2G03430 66 / 9e-14
Ankyrin repeat family protein (.1)
AT4G19150 66 / 1e-13
Ankyrin repeat family protein (.1.2)
AT3G02850 64 / 2e-12
SKOR
STELAR K+ outward rectifier, STELAR K+ outward rectifier (.1)
AT5G53470 64 / 2e-12
ACBP1
acyl-CoA binding protein 1 (.1)
AT5G37500 61 / 3e-11
GORK
gated outwardly-rectifying K+ channel, gated outwardly-rectifying K+ channel (.1)
AT2G14255 59 / 7e-11
Ankyrin repeat family protein with DHHC zinc finger domain (.1)
Paralogs
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10036033
236 / 4e-81
AT5G12320 186 / 2e-61
ankyrin repeat family protein (.1)
Lus10009696
232 / 2e-79
AT5G12320 184 / 7e-61
ankyrin repeat family protein (.1)
Lus10001414
72 / 1e-15
AT4G19150 245 / 9e-82
Ankyrin repeat family protein (.1.2)
Lus10001605
67 / 3e-13
AT4G38130 833 / 0.0
ARABIDOPSIS HISTONE DEACETYLASE 19, ARABIDOPSIS HISTONE DEACETYLASE 1, histone deacetylase 1 (.1.2)
Lus10035498
65 / 3e-13
AT3G02850 206 / 2e-61
STELAR K+ outward rectifier, STELAR K+ outward rectifier (.1)
Lus10037694
66 / 4e-13
AT3G02850 659 / 0.0
STELAR K+ outward rectifier, STELAR K+ outward rectifier (.1)
Lus10015687
66 / 5e-13
AT3G02850 1231 / 0.0
STELAR K+ outward rectifier, STELAR K+ outward rectifier (.1)
Lus10013859
63 / 3e-12
AT2G17390 427 / 1e-150
ankyrin repeat-containing 2B (.1)
Lus10009035
63 / 4e-12
AT3G23280 524 / 0.0
XB3 ortholog 5 in Arabidopsis thaliana (.1.2)
Lus10001999
62 / 7e-12
AT3G02850 970 / 0.0
STELAR K+ outward rectifier, STELAR K+ outward rectifier (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0465
Ank
PF12796
Ank_2
Ankyrin repeats (3 copies)
Representative CDS sequence
>Potri.009G070700.1 pacid=42771108 polypeptide=Potri.009G070700.1.p locus=Potri.009G070700 ID=Potri.009G070700.1.v4.1 annot-version=v4.1
ATGGGGACGGCGACAAACCAAGCAGAGCGGCAAACCCAAGTCGAAACAACGCCGGAGATTGTTGATGCTTTACTTGAGGCTGCTAGATATGATGATTTTG
AGGATATCACAAGTTTAGCATCTTCTGGGGTTTCTCTTGACTCCAAAGATTCACAGGGCAGAACAGCCCTCCATATGGCTGCAGCTAATGGGCATCTTGA
CATTGTGGAGTATCTTATCAATCAAGGAGTGGACCTCAATGCTTCTAATGAGGAGAAGAATACGCCTCTTCATTGGGCCTGTCTCAATGGTCATATAGAG
GTTGTTAAGAAATTGATTCTGGCAGGAGCGAGTTTAGGCATTCTTAATAGCCATGAACGGACTCCAATGGATGAAGCTGTTACTAGGGGAAAATTGGATG
TTATTGATGCAATTAATGCAGCTGTGGCACAACAGGAACTTTCAGGTGTTACAGTTTCCTAG
AA sequence
>Potri.009G070700.1 pacid=42771108 polypeptide=Potri.009G070700.1.p locus=Potri.009G070700 ID=Potri.009G070700.1.v4.1 annot-version=v4.1
MGTATNQAERQTQVETTPEIVDALLEAARYDDFEDITSLASSGVSLDSKDSQGRTALHMAAANGHLDIVEYLINQGVDLNASNEEKNTPLHWACLNGHIE
VVKKLILAGASLGILNSHERTPMDEAVTRGKLDVIDAINAAVAQQELSGVTVS
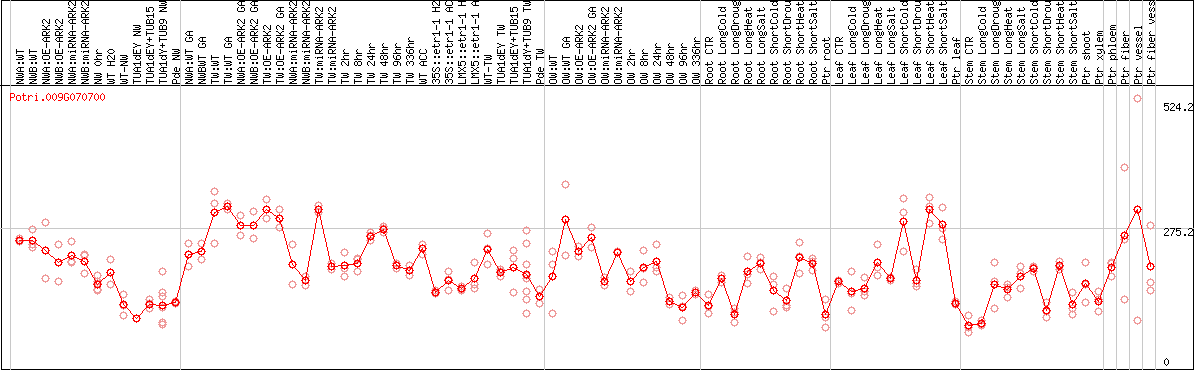
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.009G070700 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.