External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G07030 426 / 2e-150
Mitochondrial substrate carrier family protein (.1)
AT2G30160 417 / 6e-147
Mitochondrial substrate carrier family protein (.1)
AT1G34065 125 / 4e-33
SAMC2
S-adenosylmethionine carrier 2 (.1)
AT4G39460 116 / 5e-30
SAMC1, SAMT1
SAM TRANSPORTER1, S-adenosylmethionine carrier 1 (.1.2)
AT1G07025 111 / 1e-29
Mitochondrial substrate carrier family protein (.1)
AT4G11440 114 / 5e-28
Mitochondrial substrate carrier family protein (.1)
AT1G74240 109 / 5e-27
Mitochondrial substrate carrier family protein (.1)
AT4G32400 102 / 2e-24
EMB42, EMB104, ATBT1, SHS1
SODIUM HYPERSENSITIVE 1, EMBRYO DEFECTIVE 42, EMBRYO DEFECTIVE 104, ARABIDOPSIS THALIANA BRITTLE 1, Mitochondrial substrate carrier family protein (.1)
AT3G53940 97 / 2e-22
Mitochondrial substrate carrier family protein (.1)
AT5G15640 93 / 2e-21
Mitochondrial substrate carrier family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.001G282400
573 / 0
AT1G07030 426 / 1e-150
Mitochondrial substrate carrier family protein (.1)
Potri.016G125200
422 / 6e-149
AT1G07030 436 / 1e-154
Mitochondrial substrate carrier family protein (.1)
Potri.003G127600
130 / 2e-33
AT4G11440 437 / 3e-146
Mitochondrial substrate carrier family protein (.1)
Potri.005G085900
124 / 9e-33
AT4G39460 487 / 9e-175
SAM TRANSPORTER1, S-adenosylmethionine carrier 1 (.1.2)
Potri.005G197200
116 / 6e-30
AT4G39460 417 / 3e-147
SAM TRANSPORTER1, S-adenosylmethionine carrier 1 (.1.2)
Potri.006G252100
108 / 2e-26
AT4G32400 493 / 4e-175
SODIUM HYPERSENSITIVE 1, EMBRYO DEFECTIVE 42, EMBRYO DEFECTIVE 104, ARABIDOPSIS THALIANA BRITTLE 1, Mitochondrial substrate carrier family protein (.1)
Potri.018G029200
105 / 2e-25
AT4G32400 483 / 2e-171
SODIUM HYPERSENSITIVE 1, EMBRYO DEFECTIVE 42, EMBRYO DEFECTIVE 104, ARABIDOPSIS THALIANA BRITTLE 1, Mitochondrial substrate carrier family protein (.1)
Potri.012G066000
101 / 2e-24
AT1G74240 446 / 6e-158
Mitochondrial substrate carrier family protein (.1)
Potri.017G098500
99 / 9e-24
AT5G15640 460 / 3e-164
Mitochondrial substrate carrier family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10042258
480 / 1e-171
AT2G30160 484 / 2e-173
Mitochondrial substrate carrier family protein (.1)
Lus10026395
473 / 7e-169
AT2G30160 483 / 5e-173
Mitochondrial substrate carrier family protein (.1)
Lus10006812
114 / 7e-29
AT4G39460 478 / 6e-171
SAM TRANSPORTER1, S-adenosylmethionine carrier 1 (.1.2)
Lus10005787
113 / 1e-28
AT4G39460 494 / 2e-177
SAM TRANSPORTER1, S-adenosylmethionine carrier 1 (.1.2)
Lus10002026
108 / 1e-26
AT4G32400 525 / 0.0
SODIUM HYPERSENSITIVE 1, EMBRYO DEFECTIVE 42, EMBRYO DEFECTIVE 104, ARABIDOPSIS THALIANA BRITTLE 1, Mitochondrial substrate carrier family protein (.1)
Lus10002902
106 / 9e-26
AT4G32400 522 / 0.0
SODIUM HYPERSENSITIVE 1, EMBRYO DEFECTIVE 42, EMBRYO DEFECTIVE 104, ARABIDOPSIS THALIANA BRITTLE 1, Mitochondrial substrate carrier family protein (.1)
Lus10017849
101 / 3e-24
AT1G74240 430 / 6e-151
Mitochondrial substrate carrier family protein (.1)
Lus10012597
100 / 2e-23
AT5G15640 488 / 9e-173
Mitochondrial substrate carrier family protein (.1)
Lus10041498
98 / 7e-23
AT5G15640 494 / 2e-177
Mitochondrial substrate carrier family protein (.1)
Lus10029770
97 / 2e-22
AT5G42130 478 / 5e-169
MitoFerrinLike1, Mitochondrial substrate carrier family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF00153
Mito_carr
Mitochondrial carrier protein
Representative CDS sequence
>Potri.009G077600.7 pacid=42771711 polypeptide=Potri.009G077600.7.p locus=Potri.009G077600 ID=Potri.009G077600.7.v4.1 annot-version=v4.1
ATGGCCACCGATGCCACCACCACCAAATTCCAAAACCCCACCGATTTCCGCCCCGATTTCCATTCGGAGAAAATCTCCTCCGCAACCTCTTACGACGGTC
TCCATTTTTGGCAATACATGATTTCCGGTTCAATAGCCGGTCTTGTAGAACACATGGCCATGTTCCCTGTTGACACAGTCAAGACCCATATGCAAGCAAT
CGGGTCTTGCCCCATTAAATCTGTTAGTGTAACCCACGTCCTCAATTCCCTTCTCGAATCCGGAGGCCCTTCTTCCCTTTACCGTGGCATTGCCGCCATG
GCTCTTGGCGCTGGTCCTGCCCACGCTGTCCATTTTTCAGTTTATGAAGTCTGCAAGAAACATTTGTCGCGAGACAACCCAAATAGTTCAATTGCTCACG
CCATTTCTGGTGTTTGTGCAACGGTTGCTAGCGATGCGGTTTTTACACCAATGGATATGGTAAAGCAGAGATTGCAATTGGGGAGTGATAGTGTTTATAA
GGGTGTTCGGGATTGTGTTAAGAGGGTGGTGAGAGAAGAGGGGTTTGGTGCGTTTTATGCTTCGTATAGGACGACTGTTTTGATGAATGCTCCATTTACT
GCGGTTTATTTTGCAACATATGAGGCTGCGAAGAAGGGCTTGATGGAGATTTCTCCTGAGAGTGCTAATGATGAGAACTGGGTTCTTCATGCTACTGCTG
GTGCTGCAGCTGGAGCATTGGCGGCTGCTATTACCACACCACTCGATGTTGTTAAAACTCAGTTGCAGTGTCAGGGTGTTTGTGGGTGCGACAGATTTAA
AAGTGGTTCAATTGGAGATGTGATTAAAACAATAGTTAAAAAGGATGGTTACAGAGGCCTCATAAGAGGTTGGATCCCAAGGATGCTCTTCCATGCACCA
GCCGCAGCAATCTCCTGGTCTACATACGAGGCCTCAAAATCTTTCTTTCAGGAACTCAATGATAATAGTAACAGTGACAATGTCACTTGA
AA sequence
>Potri.009G077600.7 pacid=42771711 polypeptide=Potri.009G077600.7.p locus=Potri.009G077600 ID=Potri.009G077600.7.v4.1 annot-version=v4.1
MATDATTTKFQNPTDFRPDFHSEKISSATSYDGLHFWQYMISGSIAGLVEHMAMFPVDTVKTHMQAIGSCPIKSVSVTHVLNSLLESGGPSSLYRGIAAM
ALGAGPAHAVHFSVYEVCKKHLSRDNPNSSIAHAISGVCATVASDAVFTPMDMVKQRLQLGSDSVYKGVRDCVKRVVREEGFGAFYASYRTTVLMNAPFT
AVYFATYEAAKKGLMEISPESANDENWVLHATAGAAAGALAAAITTPLDVVKTQLQCQGVCGCDRFKSGSIGDVIKTIVKKDGYRGLIRGWIPRMLFHAP
AAAISWSTYEASKSFFQELNDNSNSDNVT
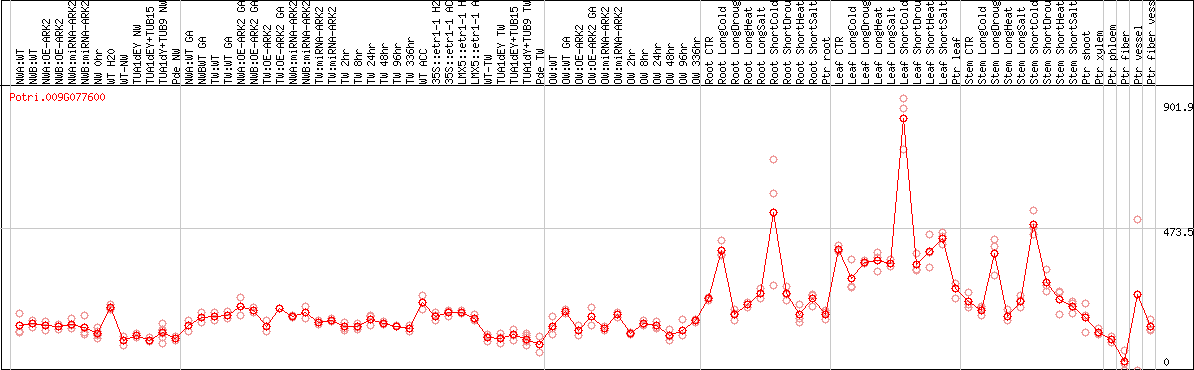
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.009G077600 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.