External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G37850 209 / 2e-65
bHLH
bHLH025
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
AT2G22750 173 / 1e-51
bHLH
bHLH018
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.2)
AT2G22760 169 / 3e-50
bHLH
bHLH019
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
AT2G22770 158 / 9e-46
bHLH
NAI1, bHLH020
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
AT2G46510 83 / 2e-17
bHLH
ATAIB, bHLH107, bHLH017, INU1, JAM1, MYL1
ABA-inducible BHLH-type transcription factor (.1)
AT1G32640 82 / 4e-17
bHLH
JIN1, JAI1, ZBF1, RD22BP1, ATMYC2
JASMONATE INSENSITIVE 1, Basic helix-loop-helix (bHLH) DNA-binding family protein (.1)
AT4G17880 82 / 8e-17
bHLH
bHLH004, MYC4
Basic helix-loop-helix (bHLH) DNA-binding family protein (.1)
AT4G29930 79 / 8e-17
bHLH
bHLH027
basic helix-loop-helix (bHLH) DNA-binding superfamily protein
AT5G57150 76 / 4e-16
bHLH
bHLH035
basic helix-loop-helix (bHLH) DNA-binding superfamily protein
AT1G01260 79 / 7e-16
bHLH
bHLH013, INU4
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.2), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.3)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.001G287200
576 / 0
AT4G37850 215 / 7e-68
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Potri.005G095250
226 / 3e-72
AT4G37850 213 / 5e-67
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Potri.007G009400
221 / 3e-69
AT4G37850 130 / 1e-34
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Potri.002G172101
90 / 1e-19
AT1G01260 526 / 0.0
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.2), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.3)
Potri.014G099700
89 / 2e-19
AT1G01260 556 / 0.0
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.2), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.3)
Potri.005G208600
85 / 8e-18
AT4G09820 296 / 2e-92
TRANSPARENT TESTA 8, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Potri.002G054100
79 / 8e-16
AT4G09820 317 / 6e-100
TRANSPARENT TESTA 8, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Potri.002G042000
78 / 1e-15
AT4G16430 475 / 4e-165
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Potri.001G142200
77 / 3e-15
AT1G32640 604 / 0.0
JASMONATE INSENSITIVE 1, Basic helix-loop-helix (bHLH) DNA-binding family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10020503
269 / 1e-89
AT4G37850 187 / 4e-58
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Lus10012457
265 / 6e-88
AT4G37850 188 / 2e-58
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Lus10011578
230 / 1e-73
AT4G37850 209 / 1e-65
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Lus10003488
230 / 8e-73
AT4G37850 212 / 6e-66
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Lus10019247
223 / 2e-70
AT4G37850 217 / 3e-68
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Lus10039639
202 / 2e-62
AT4G37850 191 / 3e-58
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Lus10039640
171 / 4e-50
AT4G37850 156 / 2e-44
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Lus10000037
91 / 3e-21
AT4G17880 300 / 3e-99
Basic helix-loop-helix (bHLH) DNA-binding family protein (.1)
Lus10030970
90 / 2e-19
AT1G32640 630 / 0.0
JASMONATE INSENSITIVE 1, Basic helix-loop-helix (bHLH) DNA-binding family protein (.1)
Lus10042572
84 / 5e-19
AT4G09820 156 / 1e-45
TRANSPARENT TESTA 8, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF00010
HLH
Helix-loop-helix DNA-binding domain
Representative CDS sequence
>Potri.009G081400.1 pacid=42772538 polypeptide=Potri.009G081400.1.p locus=Potri.009G081400 ID=Potri.009G081400.1.v4.1 annot-version=v4.1
ATGGAGACTTCATCAATTACAGCTTTATATGAACTAGGAATGGAGGATCCAGGCTTCACCAACCAGTGGTACATGAACTCCCTTGATGATATCAGCTTAC
TACCATTAGCTGCTGCTGCATTTGGAGAGAACGTACACCATCCTTTCTCTAACCAGAATTTCAACCTCAAAACCTCCATGGATAGTACTCCTACAAGCAT
TAATGTCAGACCAACAAAACAAATGAAAACTTTTCATTTGTCAGACCCCCAATCTGCCTTTTCTCCTAACTTTCTTTCATTTGTCAATCCCAACCACGCA
AATCAAATGGGATTGGTGAAGCCTAAAGAGGAGGCAGTATGTTCCAAAAGCATCAATAACTTCCCTTCTGACATGGTAGTTTCCCAAGATATCTTTGGGA
GCCAAAACTATGTAATCAAGGGTTGCCAAGGACCTGAGAGGATTAGCACAAACACTCCTAGACTTTCTCAAAGTCAAGATCACATTATTGCAGAAAGGAA
GCGGCGAGAGAAGCTTAGTCAACGATTCATAGCTCTATCTGCTGTAGTTCCTGGCCTGAAGAAGATGGACAAAGCTTCTGTTTTAGGAGATGCTATCAAG
TACTTGAAACAACTGCAAGAGAAAGTTAAGACACTTGAGGAACAAACAAAAAGGAAAACCATGGAATCAGTGGTCATTGTGAAGAAATCTCATATCTATG
TCGATGAGGGTGATGTCAACGCTTCCTCAGACGAGTCCAAAGGCCCTATTCATGAAACTTTGCCAGAAATTGAAGCAAGATTCTGTGACAAACATGTCCT
CATAAGAATCCATTGTGAGAAAAGGAAAGGAGTTTTGGAGAAAACAGTGGCTGAAATTGAGAAACTCCACCTATCAGTAATCAATAGCAGTGTTCTCGCA
TTTGGGACATCTGCTCTTCATGTTACTTTTATTGCACAGATGGATATCGACTTCAACATGTCATTGAAGGATCTAGTGAAGACTTTACGCTCAGCTTTCG
AGTTCTTCATGTGA
AA sequence
>Potri.009G081400.1 pacid=42772538 polypeptide=Potri.009G081400.1.p locus=Potri.009G081400 ID=Potri.009G081400.1.v4.1 annot-version=v4.1
METSSITALYELGMEDPGFTNQWYMNSLDDISLLPLAAAAFGENVHHPFSNQNFNLKTSMDSTPTSINVRPTKQMKTFHLSDPQSAFSPNFLSFVNPNHA
NQMGLVKPKEEAVCSKSINNFPSDMVVSQDIFGSQNYVIKGCQGPERISTNTPRLSQSQDHIIAERKRREKLSQRFIALSAVVPGLKKMDKASVLGDAIK
YLKQLQEKVKTLEEQTKRKTMESVVIVKKSHIYVDEGDVNASSDESKGPIHETLPEIEARFCDKHVLIRIHCEKRKGVLEKTVAEIEKLHLSVINSSVLA
FGTSALHVTFIAQMDIDFNMSLKDLVKTLRSAFEFFM
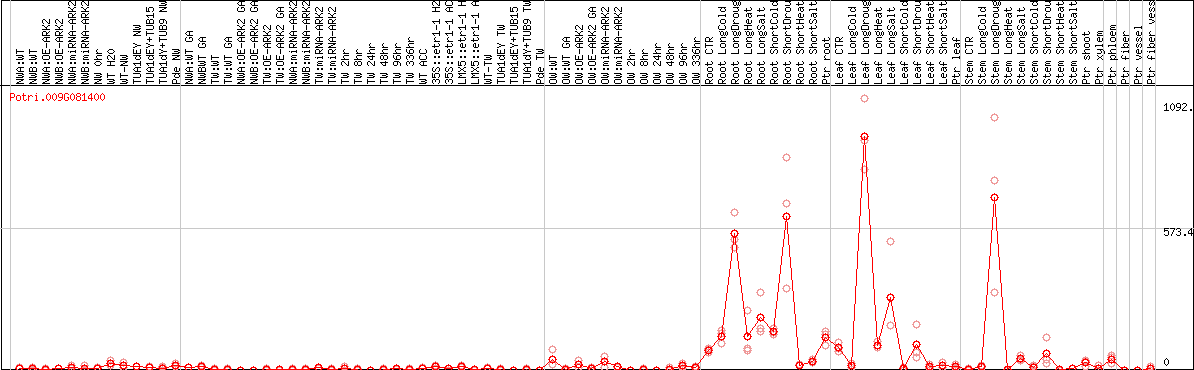
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.009G081400 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.