External link
Symbol
ATHM3.1,PtrTrxm5
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G15570 193 / 2e-63
TRX-M3, GAT1, ATHM3, ATM3
THIOREDOXIN-M3, GFP ARRESTED TRAFFICKING 1, Arabidopsis thioredoxin M-type 3, Thioredoxin superfamily protein (.1.2)
AT3G15360 120 / 1e-34
ATHM4, ATM4, TRX-M4
ARABIDOPSIS THIOREDOXIN M-TYPE 4, thioredoxin M-type 4 (.1)
AT1G03680 100 / 3e-27
ATHM1, ATM1, TRX-M1
ARABIDOPSIS THIOREDOXIN M-TYPE 1, thioredoxin M-type 1 (.1)
AT4G03520 97 / 9e-26
ATHM2
Thioredoxin superfamily protein (.1.2)
AT1G76760 79 / 1e-18
ATY1, TRX-Y1
thioredoxin Y1 (.1)
AT1G43560 75 / 3e-17
ATY2
thioredoxin Y2 (.1)
AT4G12170 70 / 8e-16
Thioredoxin superfamily protein (.1)
AT1G50320 68 / 1e-14
ATHX, ATX
thioredoxin X (.1)
AT1G52990 69 / 3e-14
thioredoxin family protein (.1)
AT3G06730 59 / 2e-11
TRXz, TRXP ,TRX z
thioredoxin putative plastidic, Thioredoxin z (.1)
Paralogs
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10019847
213 / 1e-71
AT2G15570 184 / 8e-60
THIOREDOXIN-M3, GFP ARRESTED TRAFFICKING 1, Arabidopsis thioredoxin M-type 3, Thioredoxin superfamily protein (.1.2)
Lus10014069
211 / 1e-70
AT2G15570 182 / 3e-59
THIOREDOXIN-M3, GFP ARRESTED TRAFFICKING 1, Arabidopsis thioredoxin M-type 3, Thioredoxin superfamily protein (.1.2)
Lus10014798
130 / 7e-39
AT3G15360 181 / 2e-58
ARABIDOPSIS THIOREDOXIN M-TYPE 4, thioredoxin M-type 4 (.1)
Lus10040887
130 / 1e-38
AT3G15360 182 / 4e-59
ARABIDOPSIS THIOREDOXIN M-TYPE 4, thioredoxin M-type 4 (.1)
Lus10029755
120 / 8e-35
AT3G15360 162 / 3e-51
ARABIDOPSIS THIOREDOXIN M-TYPE 4, thioredoxin M-type 4 (.1)
Lus10029752
120 / 9e-35
AT3G15360 165 / 3e-52
ARABIDOPSIS THIOREDOXIN M-TYPE 4, thioredoxin M-type 4 (.1)
Lus10042784
119 / 1e-34
AT3G15360 165 / 4e-52
ARABIDOPSIS THIOREDOXIN M-TYPE 4, thioredoxin M-type 4 (.1)
Lus10018875
74 / 9e-17
AT1G76760 176 / 6e-57
thioredoxin Y1 (.1)
Lus10028569
73 / 1e-16
AT1G76760 174 / 3e-56
thioredoxin Y1 (.1)
Lus10014186
60 / 1e-11
AT3G08710 192 / 3e-64
THIOREDOXIN TYPE H 9, thioredoxin H-type 9 (.1.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0172
Thioredoxin
PF00085
Thioredoxin
Thioredoxin
Representative CDS sequence
>Potri.009G100700.1 pacid=42771484 polypeptide=Potri.009G100700.1.p locus=Potri.009G100700 ID=Potri.009G100700.1.v4.1 annot-version=v4.1
ATGGCTTCAAGTGCTACTTCACTTTACTCCCCACCACTCACGAGCTCACGCGCCGCCGTTCTCCACCAATGTCAGCAGCTGAACCCTAACAGACTCTCTT
TTCCAAGCGACTTCAACACCGCTAAAAGAGCAACGAATCTCACTGTTCAACACGTGCCTCTCCCTCTCAAGGTTCTCTGCGGTCGCGGAAACCGAGCTAC
AGCTGTTACTCAGGACTCTTGGGAGAATTCAATTTTGAAAAGTGATATTCCTGTTCTTGTTGAATTCTATGCAAGTTGGTGTGGGCCTTGCAGGATGGTT
CATCGAGTAATTGATGAGATTGCAGCAGAGTATGATGGGAAACTCAAATGCTTTGTGCTTAACACGGACAATGACCTTCAAATTGCTGAGGACTATGAAA
TTAAGGCGGTACCAGTTGTTCTACTTTTCAAGAATGGAGAGAAGCGAGAGTCGGTGGTCGGTACCATGCCGAAGGAATTCTATATTGCTGCAGTTGAGAG
AGTTCTGCAGTCGTAA
AA sequence
>Potri.009G100700.1 pacid=42771484 polypeptide=Potri.009G100700.1.p locus=Potri.009G100700 ID=Potri.009G100700.1.v4.1 annot-version=v4.1
MASSATSLYSPPLTSSRAAVLHQCQQLNPNRLSFPSDFNTAKRATNLTVQHVPLPLKVLCGRGNRATAVTQDSWENSILKSDIPVLVEFYASWCGPCRMV
HRVIDEIAAEYDGKLKCFVLNTDNDLQIAEDYEIKAVPVVLLFKNGEKRESVVGTMPKEFYIAAVERVLQS
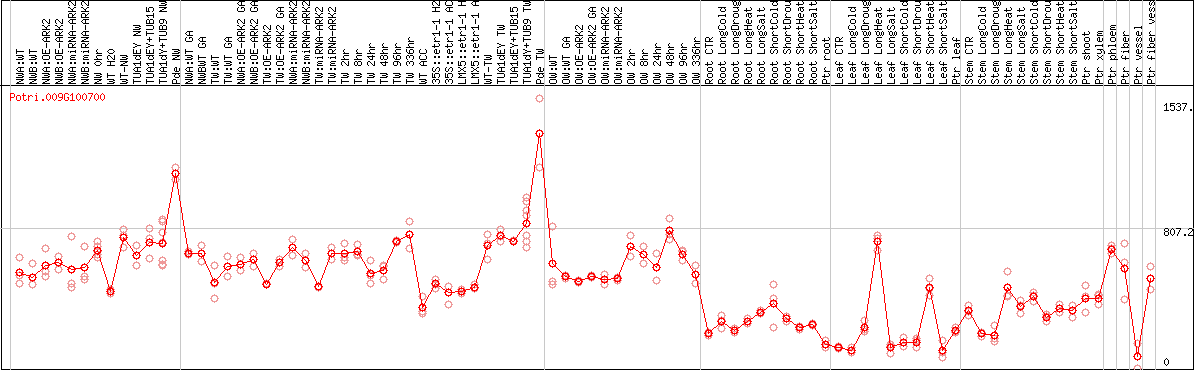
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.009G100700 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.