External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G33985 131 / 1e-39
Protein of unknown function (DUF1685) (.1)
AT2G15590 99 / 4e-27
Protein of unknown function (DUF1685) (.1), Protein of unknown function (DUF1685) (.2)
AT3G50350 90 / 2e-23
Protein of unknown function (DUF1685) (.1), Protein of unknown function (DUF1685) (.2)
AT2G15610 67 / 7e-15
Protein of unknown function (DUF1685) (.1)
AT1G08790 57 / 2e-10
Protein of unknown function (DUF1685) (.1)
AT3G04700 55 / 6e-10
Protein of unknown function (DUF1685) (.1)
AT2G31560 54 / 3e-09
Protein of unknown function (DUF1685) (.1), Protein of unknown function (DUF1685) (.2)
AT3G04710 54 / 5e-09
TPR10
tetratricopeptide repeat 10, ankyrin repeat family protein (.1.2.3)
AT5G28690 52 / 7e-09
Protein of unknown function (DUF1685) (.1)
AT1G05870 49 / 8e-08
Protein of unknown function (DUF1685) (.1), Protein of unknown function (DUF1685) (.2), Protein of unknown function (DUF1685) (.3), Protein of unknown function (DUF1685) (.4)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.005G138100
98 / 2e-26
AT4G33985 139 / 2e-42
Protein of unknown function (DUF1685) (.1)
Potri.007G043900
89 / 1e-22
AT4G33985 134 / 3e-40
Protein of unknown function (DUF1685) (.1)
Potri.013G042400
62 / 2e-12
AT1G08790 151 / 2e-46
Protein of unknown function (DUF1685) (.1)
Potri.005G055100
59 / 3e-11
AT1G08790 126 / 1e-36
Protein of unknown function (DUF1685) (.1)
Potri.017G032800
53 / 4e-09
AT1G05870 190 / 1e-61
Protein of unknown function (DUF1685) (.1), Protein of unknown function (DUF1685) (.2), Protein of unknown function (DUF1685) (.3), Protein of unknown function (DUF1685) (.4)
Potri.007G126700
52 / 1e-08
AT1G05870 188 / 7e-61
Protein of unknown function (DUF1685) (.1), Protein of unknown function (DUF1685) (.2), Protein of unknown function (DUF1685) (.3), Protein of unknown function (DUF1685) (.4)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10027898
106 / 1e-29
AT4G33985 141 / 2e-43
Protein of unknown function (DUF1685) (.1)
Lus10019852
105 / 2e-29
AT4G33985 149 / 9e-47
Protein of unknown function (DUF1685) (.1)
Lus10002400
104 / 7e-29
AT4G33985 150 / 7e-47
Protein of unknown function (DUF1685) (.1)
Lus10014064
101 / 7e-28
AT4G33985 145 / 2e-45
Protein of unknown function (DUF1685) (.1)
Lus10006560
68 / 7e-15
AT1G08790 166 / 2e-52
Protein of unknown function (DUF1685) (.1)
Lus10021997
63 / 2e-13
AT1G08790 129 / 4e-39
Protein of unknown function (DUF1685) (.1)
Lus10042536
62 / 5e-13
AT1G08790 141 / 2e-43
Protein of unknown function (DUF1685) (.1)
Lus10025562
45 / 3e-06
AT1G05870 183 / 1e-58
Protein of unknown function (DUF1685) (.1), Protein of unknown function (DUF1685) (.2), Protein of unknown function (DUF1685) (.3), Protein of unknown function (DUF1685) (.4)
Lus10027022
45 / 3e-06
AT2G31560 183 / 1e-58
Protein of unknown function (DUF1685) (.1), Protein of unknown function (DUF1685) (.2)
Lus10005536
44 / 4e-06
AT5G28690 119 / 3e-35
Protein of unknown function (DUF1685) (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF07939
DUF1685
Protein of unknown function (DUF1685)
Representative CDS sequence
>Potri.009G101300.3 pacid=42771033 polypeptide=Potri.009G101300.3.p locus=Potri.009G101300 ID=Potri.009G101300.3.v4.1 annot-version=v4.1
ATGTCAATTCAACCACGACCCCTCCCGCCACCACGGAATCCCCGGCCTACTATGCTCAACAAGCAGCTATCGTGGTCTCCAGACATGACCCGCGAGGAAG
TATGGCTCAGAAGGAAAGGAAATTCTGCAACCCGACGCCGGTGCAGCAAGAGTGTGACTGACGATGATTTAGAGGAGTTAAAAGCATGTATTGAATTAGG
ATTCGGGTTCGGACCCGACTCGTCTGATTTGGATCCGAAGTTGTCGGATACCTTGCCCGCTTTAGGTTTTTACTGTGCTCTTAATAAACAGTATAGTAGT
TGTTTGTCTAGATCAGCTTCGACGTCGTCGCTTTTGTCTGTTTCAGGTGATGATCCGGAGATGGTGAAGAAGAGGTTGCGACAATGGGCCCAGATTGTCG
CATGCTCGGTCAAGCAATTTTCAGGGGAACCAAATTGA
AA sequence
>Potri.009G101300.3 pacid=42771033 polypeptide=Potri.009G101300.3.p locus=Potri.009G101300 ID=Potri.009G101300.3.v4.1 annot-version=v4.1
MSIQPRPLPPPRNPRPTMLNKQLSWSPDMTREEVWLRRKGNSATRRRCSKSVTDDDLEELKACIELGFGFGPDSSDLDPKLSDTLPALGFYCALNKQYSS
CLSRSASTSSLLSVSGDDPEMVKKRLRQWAQIVACSVKQFSGEPN
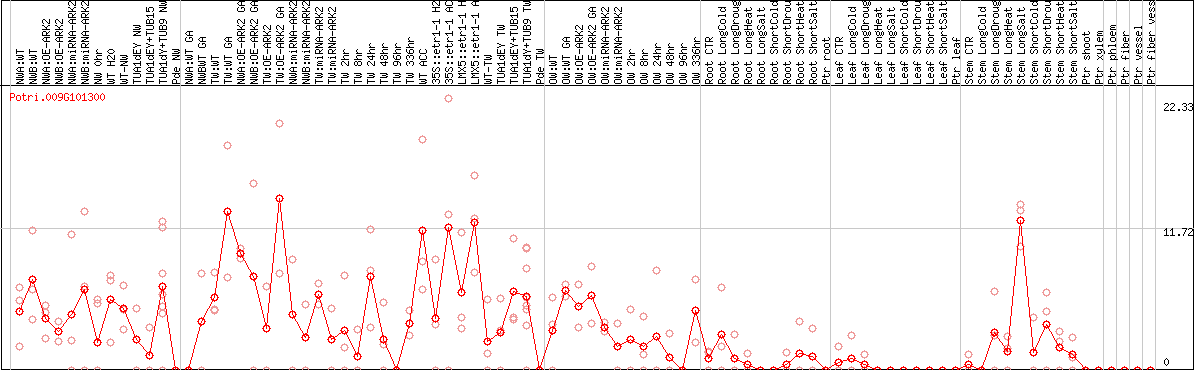
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.009G101300 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.