External link
Symbol
Pt-ICK5.1,7
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G49620 92 / 6e-23
ICN6, ICK5, KRP7
KIP-RELATED PROTEIN 7, Cyclin-dependent kinase inhibitor family protein (.1)
AT3G19150 75 / 1e-16
ACK1, ICK4, KRP6
ARABIDOPSIS CDK INHIBITOR 1, KIP-related protein 6 (.1.2)
AT5G48820 71 / 6e-15
ICK6, KRP3
KIP-RELATED PROTEIN 3, inhibitor/interactor with cyclin-dependent kinase (.1.2)
AT2G32710 62 / 1e-11
ICK7, ACK2, KRP4
KIP-RELATED PROTEIN 4, INTERACTORS OF CDC2 KINASE 7, CYCLIN-DEPENDENT KINASE INHIBITOR 2, Cyclin-dependent kinase inhibitor family protein (.1.2)
AT3G50630 61 / 1e-11
ICK2, KRP2
KIP-related protein 2 (.1)
AT3G24810 60 / 5e-11
ICK3, KRP5
kip-related protein 5, Cyclin-dependent kinase inhibitor family protein (.1)
AT2G23430 47 / 1e-06
ICK1, KRP1
KIP-RELATED PROTEIN 1, Cyclin-dependent kinase inhibitor family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.004G142100
307 / 1e-107
AT1G49620 91 / 7e-23
KIP-RELATED PROTEIN 7, Cyclin-dependent kinase inhibitor family protein (.1)
Potri.002G242100
71 / 4e-15
AT5G48820 177 / 6e-56
KIP-RELATED PROTEIN 3, inhibitor/interactor with cyclin-dependent kinase (.1.2)
Potri.007G042300
71 / 4e-15
AT2G23430 /
KIP-RELATED PROTEIN 1, Cyclin-dependent kinase inhibitor family protein (.1)
Potri.005G137500
70 / 1e-14
AT3G19150 50 / 2e-07
ARABIDOPSIS CDK INHIBITOR 1, KIP-related protein 6 (.1.2)
Potri.017G054000
67 / 2e-13
AT5G48820 127 / 3e-36
KIP-RELATED PROTEIN 3, inhibitor/interactor with cyclin-dependent kinase (.1.2)
Potri.001G314000
63 / 4e-12
AT5G48820 131 / 2e-37
KIP-RELATED PROTEIN 3, inhibitor/interactor with cyclin-dependent kinase (.1.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10011862
65 / 1e-12
AT5G48820 146 / 2e-43
KIP-RELATED PROTEIN 3, inhibitor/interactor with cyclin-dependent kinase (.1.2)
Lus10043139
56 / 2e-09
AT2G32710 143 / 2e-41
KIP-RELATED PROTEIN 4, INTERACTORS OF CDC2 KINASE 7, CYCLIN-DEPENDENT KINASE INHIBITOR 2, Cyclin-dependent kinase inhibitor family protein (.1.2)
Lus10032611
52 / 4e-08
AT2G32710 139 / 1e-39
KIP-RELATED PROTEIN 4, INTERACTORS OF CDC2 KINASE 7, CYCLIN-DEPENDENT KINASE INHIBITOR 2, Cyclin-dependent kinase inhibitor family protein (.1.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF02234
CDI
Cyclin-dependent kinase inhibitor
Representative CDS sequence
>Potri.009G103600.1 pacid=42772015 polypeptide=Potri.009G103600.1.p locus=Potri.009G103600 ID=Potri.009G103600.1.v4.1 annot-version=v4.1
ATGGGAGAGAGCGCGAGGAATAGTAAGCGAATAGCTGAAACTGGAATTCTAGAATCTTCAGTCACAAGTTTCTCTAAGAAAATGAAATTTGATTTTGACG
AGTTTAGCTTACCTTCGTTTAATTTCAAGCTCCAAGCTCATCTTTGCACCACTATGTCACCGGAAAATTTGATTTCTTCGGCGGCTTCATCGAATTCTGA
TCGCTTTATAACCGGCAATTCCAGCTGTGGCGACTCTCCGGTTTCCTGCTGCTCCAGCAATGAATCGATCAAGGTTGTGAAGGACAGCTTGAGGTTTATA
GATCTGGAGGCGAAGAGTTCCGAAACGGAAAGCTCGACGTGCAATGACAGGAAATTCAGTAGAGACACCACTCCTTCAAGCGAGTTTCACGGGATGTACT
CGCCGGCAGCCGTGGAGAAGAAAGAGAATTCTCACAGGAGAAAGTCACCGGCTGTGAAAATGCCGAGTCAGGCTGAGATCGATGCGTTTTTCGCGGGGGC
AGAGAGAGAGGAGCAGAAAAGATTTGCAGAGAAGTACAACTACGATGTTGTGAAGGATTTGCCAGTGGAGGGTCGCTACCAGTGGATTTGTTTGAAGCCA
CAGAGGAAAATAGAGAATTGA
AA sequence
>Potri.009G103600.1 pacid=42772015 polypeptide=Potri.009G103600.1.p locus=Potri.009G103600 ID=Potri.009G103600.1.v4.1 annot-version=v4.1
MGESARNSKRIAETGILESSVTSFSKKMKFDFDEFSLPSFNFKLQAHLCTTMSPENLISSAASSNSDRFITGNSSCGDSPVSCCSSNESIKVVKDSLRFI
DLEAKSSETESSTCNDRKFSRDTTPSSEFHGMYSPAAVEKKENSHRRKSPAVKMPSQAEIDAFFAGAEREEQKRFAEKYNYDVVKDLPVEGRYQWICLKP
QRKIEN
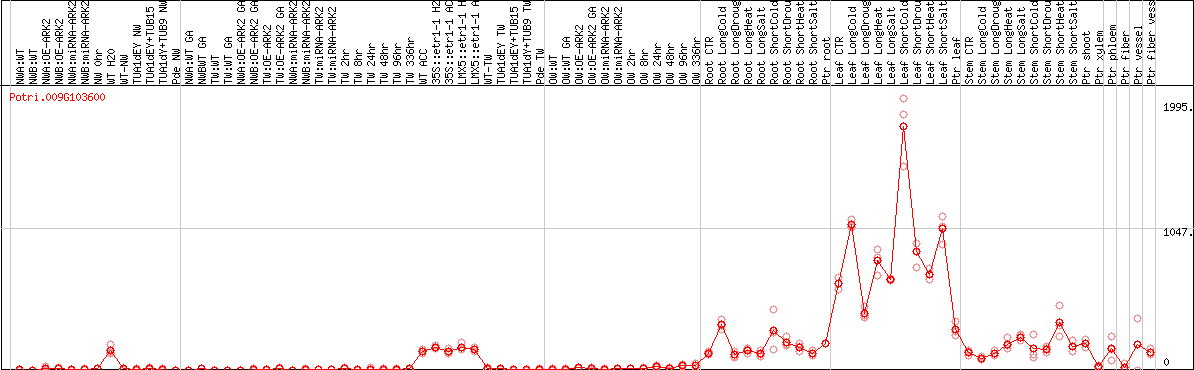
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.009G103600 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.