External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G04900 70 / 3e-14
Heavy metal transport/detoxification superfamily protein (.1)
AT4G23882 65 / 5e-12
Heavy metal transport/detoxification superfamily protein (.1)
AT3G06130 60 / 4e-10
Heavy metal transport/detoxification superfamily protein (.1.2)
AT5G37860 58 / 1e-09
Heavy metal transport/detoxification superfamily protein (.1)
AT1G30473 55 / 9e-09
Heavy metal transport/detoxification superfamily protein (.1)
AT5G19090 56 / 1e-08
Heavy metal transport/detoxification superfamily protein (.1.2.3)
AT1G23000 55 / 2e-08
Heavy metal transport/detoxification superfamily protein (.1)
AT3G05220 55 / 2e-08
Heavy metal transport/detoxification superfamily protein (.1.2)
AT1G71050 49 / 4e-07
HIPP20
heavy metal associated isoprenylated plant protein 20, Heavy metal transport/detoxification superfamily protein (.1)
AT5G27690 50 / 9e-07
Heavy metal transport/detoxification superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.001G326100
103 / 3e-26
AT3G04900 66 / 8e-13
Heavy metal transport/detoxification superfamily protein (.1)
Potri.004G144200
79 / 7e-17
AT3G04900 58 / 7e-10
Heavy metal transport/detoxification superfamily protein (.1)
Potri.004G091700
65 / 4e-12
AT1G23000 99 / 1e-23
Heavy metal transport/detoxification superfamily protein (.1)
Potri.008G202800
59 / 7e-10
AT3G06130 150 / 6e-41
Heavy metal transport/detoxification superfamily protein (.1.2)
Potri.010G175800
58 / 1e-09
AT3G06130 81 / 3e-17
Heavy metal transport/detoxification superfamily protein (.1.2)
Potri.005G036300
58 / 2e-09
AT3G06130 114 / 7e-28
Heavy metal transport/detoxification superfamily protein (.1.2)
Potri.010G114300
56 / 7e-09
AT1G23000 128 / 2e-33
Heavy metal transport/detoxification superfamily protein (.1)
Potri.019G107500
50 / 2e-07
AT1G06330 213 / 5e-72
Heavy metal transport/detoxification superfamily protein (.1)
Potri.010G114600
50 / 2e-07
AT1G22990 194 / 9e-65
heavy metal associated isoprenylated plant protein 22, Heavy metal transport/detoxification superfamily protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10041228
57 / 4e-09
AT3G06130 149 / 3e-41
Heavy metal transport/detoxification superfamily protein (.1.2)
Lus10032445
55 / 2e-08
AT1G23000 129 / 2e-34
Heavy metal transport/detoxification superfamily protein (.1)
Lus10013911
52 / 4e-08
AT1G06330 103 / 2e-28
Heavy metal transport/detoxification superfamily protein (.1)
Lus10032446
50 / 3e-07
AT1G71050 175 / 2e-56
heavy metal associated isoprenylated plant protein 20, Heavy metal transport/detoxification superfamily protein (.1)
Lus10001892
49 / 4e-07
AT1G06330 101 / 1e-27
Heavy metal transport/detoxification superfamily protein (.1)
Lus10042946
47 / 2e-06
AT1G71050 196 / 3e-65
heavy metal associated isoprenylated plant protein 20, Heavy metal transport/detoxification superfamily protein (.1)
Lus10031514
49 / 3e-06
AT5G19090 118 / 1e-28
Heavy metal transport/detoxification superfamily protein (.1.2.3)
Lus10020704
47 / 3e-06
AT1G06330 173 / 3e-56
Heavy metal transport/detoxification superfamily protein (.1)
Lus10009115
46 / 6e-06
AT5G17450 206 / 5e-69
heavy metal associated isoprenylated plant protein 21, Heavy metal transport/detoxification superfamily protein (.1.2)
Lus10028531
46 / 6e-06
AT5G17450 207 / 1e-69
heavy metal associated isoprenylated plant protein 21, Heavy metal transport/detoxification superfamily protein (.1.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF00403
HMA
Heavy-metal-associated domain
Representative CDS sequence
>Potri.009G126600.1 pacid=42771803 polypeptide=Potri.009G126600.1.p locus=Potri.009G126600 ID=Potri.009G126600.1.v4.1 annot-version=v4.1
ATGGCTGTCCCTATTTTCACCGTGAAAGTGCATATCAGTTGCTGTTCAAGATGTACGCAGAGAGCCAAGGAAAAGTTGCAGAAAATCAAAGGGGTGAATT
CCATCACCATCGACACAGCAAAAGATTTGGTGATCGTTTCCGGCAGTGTTGAGCCTGCGGTAATTCTAGAGAAGTTCGCTGAGTGGGGAAAAAAGGCAGA
ACTTTTTTCCTTTCAAAAAGAACCCATGGAGAGTGGCGGTGGCCACGACAAGGATAAAACTAATTTGAATAAAAATTATTATAGGAAGAGTGATTCGGTT
CCCGGCGCAGTACCCGCCAACAGGTTTTTGCATGATAACCCTAACATGTACGTCGCGAGAGACAGCAAGAAGGGTAAATGGTGTTGGCTTGGTAAAATAA
TATTCGGAAACAAACCGAGGGCTAGATCTTTGGTTGCAAAAGCTCATGCTGACAATAAGTGGCAGGCTCCAAGGATACCTATGAATGAATTTGGATCATC
AAAGCCAATATATGGAAACCAATTAGCTATGCATGAATTTAGATCCTCACAGCCATTTTATGGTAGCCAATTACCTATGGATCAATTTGGATCCTCAAGA
CCATTTAATGGAAGCCATTTCAATAGACCATCGATGTATGGAAGACCAAGTCGACAACCGGTGCCTCCACAACCAATGTATATGCCAAGGCCACCATTCT
TACCCCCATATGGGGGGATGCAGCCAGGGGCGGCAGCTGCACCGTCGCACTCGGTCAATCCCATGATCCACTACACAAGTTATGACGATAATTATCGTCA
TTGGTGA
AA sequence
>Potri.009G126600.1 pacid=42771803 polypeptide=Potri.009G126600.1.p locus=Potri.009G126600 ID=Potri.009G126600.1.v4.1 annot-version=v4.1
MAVPIFTVKVHISCCSRCTQRAKEKLQKIKGVNSITIDTAKDLVIVSGSVEPAVILEKFAEWGKKAELFSFQKEPMESGGGHDKDKTNLNKNYYRKSDSV
PGAVPANRFLHDNPNMYVARDSKKGKWCWLGKIIFGNKPRARSLVAKAHADNKWQAPRIPMNEFGSSKPIYGNQLAMHEFRSSQPFYGSQLPMDQFGSSR
PFNGSHFNRPSMYGRPSRQPVPPQPMYMPRPPFLPPYGGMQPGAAAAPSHSVNPMIHYTSYDDNYRHW
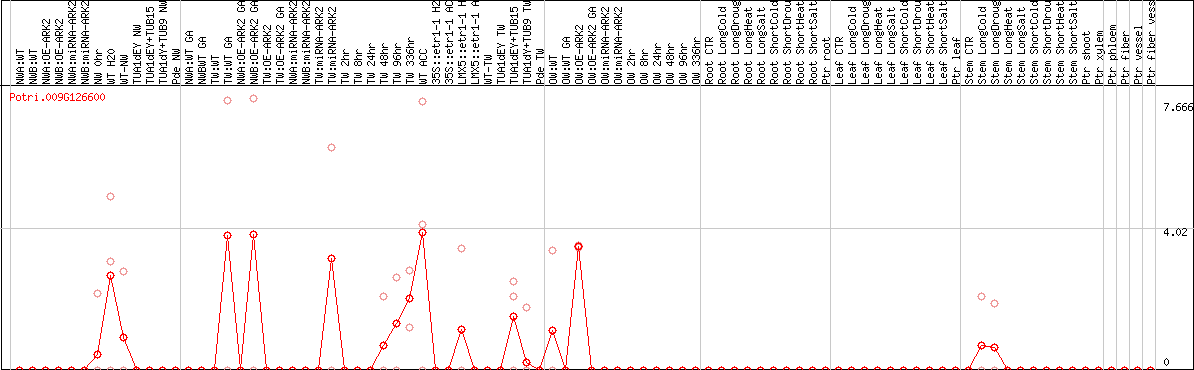
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.009G126600 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.