Potri.009G134300 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.009G134300.9 pacid=42772000 polypeptide=Potri.009G134300.9.p locus=Potri.009G134300 ID=Potri.009G134300.9.v4.1 annot-version=v4.1
ATGGAAACTCGTAGCCGGAAACGGGCGGAGGCCTCCTCATCTGCCGCCACCAACACCACCGGTACCACTACTCGCTCCAACAAACGATCCCGCACCAACG
CTGCCACCGCAACAGCAACGACAACAACCGCCACCGCTACCCGCTCTCGCTCCACACGTGCTCACCCTCTCCCCATGGACTCTACACCTGTTGAATCGTC
TTCTTCTTCTCGCTCGCGTCGGAATAGGAACAATAATAGCAATAGTGAATCGGAGAAAGGCAAAGAGAAAGAACATGAAGTTAGGGTTTCGCGTGAAAAT
AGGGAAATCACTAATAATTTGGATTCTGGAAATGACAACAATAACCCTAATGTCGATGACGACGATGATGATGACAGCGAAGGCGGTGGAATTGCTGCTT
TCCATCAGAATTTGACTTCAGCTAGTAGTGCTTTACAGGGTTTGTTGAGAAAGTTGGGTGCCGGTCTTGATGATTTACTGCCTTCTCCAGTGATGGGTTC
AGGTTCAAGCTCGCATCAATCAGGGAGATTGAAGAAGATTTTGTCTGGGTTAAGGGCGGATGGAGAAGAAGGGAAGCAAGTGGAAGCTTTAACGCAGCTT
TGTGAAATGCTTTCTATCGGGACTGAAGAGAGTCTGAGTACTTTTTCTGTGGATTCCTTTGTTCCGGTTCTTGTTGGGTTGTTGAATAATGAGAGTAATC
CTGATATTATGTTGCTTGCTGCAAGGGCGATTACGCATTTGTGTGATGTCTTGCCTTCGTCTTGTGCTGCTGTTGTTCATTATGGCGCTGTTTCGTGTTT
TGTTGCAAGGTTGATTACTATTGAGTACATGGACCTTGCTGAACAGTCTCTGCAAGCTCTAAAGAAGATATCTCAAGAGCACCCAACTGCATGTCTGCGT
GCCGGTGCTCTTATGGCAGTGCTTTCTTATCTTGATTTTTTTTCTACTGGAGTTCAGCGAGTAGCATTATCTACTGCTGCAAATATGTGCAAGAAACTTC
CTTCAGATGCAGCTGACTTTGTGATGGAAGCAGTTCCTCTGTTGACCAACCTTCTTCAGTACCATGATGCCAAGGTTTTAGAGCACGCTTCAGTTTGTTT
GACAAGGATTGCTGAAGCCTTTGCCTCATCTCCAGACAAATTAGATGAACTGTGTAATCATGGACTTGTCACACAAGCTGCCTCTCTCATTTCCACTAGC
AGTTCCGGCGGTGGACAGGCTTCTCTCAGCACTCCAACATATACGGGTTTAATTCGACTGCTATCCACTTGTGCAAGTGGGTCTCCTTTAGGAGCCAAAA
CATTACTTCTTCTTGGTGTCAGTGGCATTCTTAAAGAGATACTATCAGGTTCCGGGGTTTCTGCTAACTCCCCTGTTCCTCCTGCTCTAAGTCGACCTGC
AGATCAGATTTTTGAGATTGTCAATCTGGCAAATGAGCTTCTTCCTCCGCTTCCACAAGGAACTATCTCTCTCCCAACCAGCTCTAGTATGTTGGTCAAA
GGATCTGTTGTGAAGAAGTGTCCTTCCAGTAGTTCTGGGAAACAAGATGACATCAATGGAAATGTTCCTGAGGTCTCAGCTCGTGAGAAATTATTAAATG
ATCAACCAGAACTTCTGCAGCAATTTGGAATGGACCTTCTTCCTGTTCTGATACAGATTTATGGTTCCAGTGTCAACAGTCCTGTTCGCCACAAGTGTCT
CTCAGTTATTGGGAAGTTGATGCACTTCAGCAATGCAGAGATGATTCAGTCTCTACTTAGCATGACAAACATATCAAGTTTCCTAGCTGGGGTGTTAGCA
TGGAAAGATCCACATGTCTTGGTTCCTGCCCTTCAAGTAGCCGAGATTCTTATGGAAAAACTTCCTGGGACTTTCTCCAAGATTTTTGTCAGAGAAGGTG
TGGTTTATGCTGTAGATCAACTTATCTTAGCTGGAAACCCAAATACTGCTCCTACCCATGGTTCTTCTGCTGAAAAGGATAATGAGTCTGTTCCTGGAAC
TTCGTCACGTTCCAGGCGTTACAAACGTCGCAGTGGGAGCTCAAATCCTGAAGCAAATTCATCTGAAGAATCCAAAAATCCTATTTCTGCAAATGCTGGC
TCACCTCCAAGTTCTATAGAAATCCCAATGGTCAATTCAAATTTGCGAATGGCAGTCAGTGCATGTGCTAAAGCTTTCAGAGATAAGTACTTCCCCTCAG
ATCCCGGGGCTGCTGAAGACGGGGTTACTGATGACCTTTTACACTTGAAGAATCTCTGCACAAAGTTGAATGCTGGTGTTGATGACCAAAAGACTAAAGC
AAAGGGAAAATCAAAGGCTTCTGCATCCCGTCTTATTGACAGCTCCACAAATAAGGAAGAGTACTTGATTGGGGTGATATCCGAGATGCTAGCAGAGCTT
GGCAAAGGGGATGGTGTATCCACTTTTGAGTTTATTGGAAGTGGTGTTGTGGCAACCTTGCTGAATTTTTTTTCTTGTGGCTATTCTACCAAGGAGAAAA
TTTCAGAAGCTAACCTGCCAAAGCTTCGGCAACAAGCACTTAGAAGATTTAAGTCATTTGCTATTCTGGCCCTTCCTTCCAGCATTGATGAAGGGGGTGC
GGCACCCATGGCAGTCTTAGTTCAGAAGCTTCAAAATGCTTTGTCATCCTTGGAGCGTTTTCCTGTTGTTCTGAGCCATTCTTCTAGGTCATCTAGTGGA
GGTGCACGCCTGTCTTCTGGATTAAGTGCACTATCTCAACCTTTTAAGTTGCGTCTCTGTCGGGCCCAAGGGGAAAAGGCTCTCCGTGATTACTCTTCCA
ATGTTGTACTTATTGATCCATTAGCAAGTTTAGCTGCTGTAGAAGAATTTCTTTGGCCGCGAGTGCAGCGAAGTGAAACTGGTCACAAGGCTTCAGCATC
TGCAGGAAACTCTGAATCTGGGAATGCACAGCCTGGAGCTGGTGCATCATCTCCATCTACCTCTATCCCTGCTTCTGCTACTCGTCGTCATTCTTCAAGA
TCCAGATCATCTGTTAATATAGGAGATTCAGCTAGAAAGGAACCGATACCAGAGAAAAGCACAAGCACAAGCACATCAAAGGGAAAGGGCAAAGCTGTTT
TGAAACCACCCCTGGAGGAGACGAAAGGACCTCAAACTAGAAATGCTGCTCGTAGAAGAGCAGCCATTGATAAAGATGCACAAATGAAACCAGTGCATGG
GGATTCTAGTTCCGAGGATGAGGAATTGGATATATCTCCTGTTGAGATCGATGATGCATTGGTGATTGAAGATGATGACATCTCTGATGATGATGATGAT
GATGATGATGACCATGAAGATGTGTTAAGAGATGATTCTCTTCCTGTTTGCATGCCTGAAAAGGTACATGATGTGAAACTGGGTGCTGCATCAGAGGATA
GCAATGTTGCACCACCAGCAAGTGACAGCCAAAGTAATCCTGCTTCTGGTTCTAGTAGCAGGGCTGTTGCTGTTAGGGGCTCGGATTCCACTGATTTTAG
GAGTGGTAGTTCTTATGGATCTAGGGGAGCTATGTCATTTGCCGCCGCTGCCATGGCTGGGCTTGGATCTGCTAATGGCAGAGGCATCAGGGGAGGAAGA
GATCGACAAGGACGTCCCTTGTTTGGTAGCTCTAGTGACCCTCCAAAGTTAATCTTTACTGCTGCTGGGAAGCAACTAAATAGGCATTTGACAATCTATC
AGGCCATCCAGCGACAACTTGTTCTGGAAGAGGATGATGAAGACAGGTACGGTGGCAGGGATTTTATCTCCAGCGATGGGAGTAGGCTTTGGAGTGATAT
ATATACCCTTACATATCAAAGGGCAGATGGCCAAGCTGATAGGGCTTCTGTTGGGGGCCCAAGTTCTAGTGCATCAAAGTCTATCAAAGGTGGCTCTTCC
AATTCAAACTCAGATACCCAAGTGCACCGCATGTCACTTTTAGATAGCATCTTGCAAGCGGACCTTCCTTGTGATTTGGAAAAATCTAATCCTACTTATA
ATATATTGGCTCTACTACGTATACTTGAGGGTTTGAACCAGCTTGCTCCTCGTCTGAGAGTTCAGCTAGTTTCTGACAACTTTTCTGAGGGGAAAATCTC
AAGCTTGGATGAGTTGATGACTGCTACTGGTGTCAGGGTTCCTGCTGAAGAATTCATAAATAGCAAGCTCACACCTAAATTAGCTCGACAAATTCAAGAT
GCCCTTGCTCTGTGCAGTGGGAGCCTTCCGTCATGGTGTTATCAATTAACAAAAGCATGCCCATTCCTGTTCCCTTTTGAGACCCGGCGACAATACTTCT
ATTCAACTGCTTTTGGATTATCCCGTGCTTTGTATCGTCTTCAGCAGCAGCAGGGGGCTGATGGTCATGGATCAGCGAATGAAAGAGAGGTGAGGGTTGG
AAGATTACAACGCCAGAAAGTTCGTGTGTCCCGAAACCGTATTTTGGATTCTGCTGCAAAAGTAATGGAGATGTATTCTAGCCAGAAGGCTGTGCTTGAA
GTAGAATATTTTGGTGAGGTTGGCACAGGGTTGGGTCCTACCTTAGAGTTCTACACGCTTTTGAGTCACGATCTTCAGAAAGTCACTCTTGGAATGTGGA
GGTCAAATTCAGCAGCAGAGAAACCTTCAATGGAAATTGATGGAGATGATGACAAAAATGGAAAATCCAATAATGAATCTGGTACTGCTGTTGCTGCAGA
TCTTGTCCAGACTCCACTTGGTTTATTCCCTCGTCCTTGGCCACCAACTGCTAGTGCTTCTGAGGGGAGCCAAATATATAAGACCATTGAGTATTTCCGG
CTGGTTGGACGGGTGATGGCCAAAGCTCTTCAAGATGGACGGCTTTTGGACCTACCACTTTCGATGGCATTTTATAAGCTTGTGCTTGGTCAAGAACTTG
ATCTGTATGATATTCTTTCGTTTGATGCTGAATTTGGGAAGACTTTACAAGAATTGCATGCCCTTGTTTGTCGGAAACATTATCTAGAATCAATAGGGAG
TGATCATGAAGCCATTGCTGATTTACATTTTCATGGGACCCCGATAGAAGATCTCTGCTTGGATTTTACTCTTCCGGGTTATCCAGACTACATTTTGAAG
CCAGGAGATGAAACTGTCGATATCAATAACTTGGAGGAATTCATATCCTTGGTAGTTGACGCAACTGTCAAGACTGGGATTACGCGACAAATGGAAGCAT
TTAGAGAGGGGTTCAATCAGGTTTTTGACATCTCCTCATTGCAAATATTCACTCCACAAGAATTGGATTATTTGCTTTGCGGTCGCAGAGAGTTGTGGGA
GCCTGATACACTTGTTGATCATATAAAATTTGATCATGGATACACTGCCAAGAGTCCAGCCATTGTTAACCTGCTTGAGATTATGGGAGAGTTCACACCG
GATCAGCAGCGTGCATTCTGCCAGTTTGTTACTGGGGCACCCAGGCTACCACCTGGAGGTCTGGCTGTCCTAAACCCAAAATTAACCATTGTGAGAAAGC
ATTCTTCATCTGCAGGGAATGCAATGCCAAATGGAACTGGACCTTCAGAATCAGCTGATGATGACTTGCCTAGTGTCATGACTTGTGCTAATTACCTAAA
GCTTCCTCCTTACTCTACCAAGGAAGTTATGTACAAGAAATTGCTGTACGCAATCAGTGAAGGGCAGGGATCTTTTGATCTGTCATGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.009G134300.9 pacid=42772000 polypeptide=Potri.009G134300.9.p locus=Potri.009G134300 ID=Potri.009G134300.9.v4.1 annot-version=v4.1
METRSRKRAEASSSAATNTTGTTTRSNKRSRTNAATATATTTTATATRSRSTRAHPLPMDSTPVESSSSSRSRRNRNNNSNSESEKGKEKEHEVRVSREN
REITNNLDSGNDNNNPNVDDDDDDDSEGGGIAAFHQNLTSASSALQGLLRKLGAGLDDLLPSPVMGSGSSSHQSGRLKKILSGLRADGEEGKQVEALTQL
CEMLSIGTEESLSTFSVDSFVPVLVGLLNNESNPDIMLLAARAITHLCDVLPSSCAAVVHYGAVSCFVARLITIEYMDLAEQSLQALKKISQEHPTACLR
AGALMAVLSYLDFFSTGVQRVALSTAANMCKKLPSDAADFVMEAVPLLTNLLQYHDAKVLEHASVCLTRIAEAFASSPDKLDELCNHGLVTQAASLISTS
SSGGGQASLSTPTYTGLIRLLSTCASGSPLGAKTLLLLGVSGILKEILSGSGVSANSPVPPALSRPADQIFEIVNLANELLPPLPQGTISLPTSSSMLVK
GSVVKKCPSSSSGKQDDINGNVPEVSAREKLLNDQPELLQQFGMDLLPVLIQIYGSSVNSPVRHKCLSVIGKLMHFSNAEMIQSLLSMTNISSFLAGVLA
WKDPHVLVPALQVAEILMEKLPGTFSKIFVREGVVYAVDQLILAGNPNTAPTHGSSAEKDNESVPGTSSRSRRYKRRSGSSNPEANSSEESKNPISANAG
SPPSSIEIPMVNSNLRMAVSACAKAFRDKYFPSDPGAAEDGVTDDLLHLKNLCTKLNAGVDDQKTKAKGKSKASASRLIDSSTNKEEYLIGVISEMLAEL
GKGDGVSTFEFIGSGVVATLLNFFSCGYSTKEKISEANLPKLRQQALRRFKSFAILALPSSIDEGGAAPMAVLVQKLQNALSSLERFPVVLSHSSRSSSG
GARLSSGLSALSQPFKLRLCRAQGEKALRDYSSNVVLIDPLASLAAVEEFLWPRVQRSETGHKASASAGNSESGNAQPGAGASSPSTSIPASATRRHSSR
SRSSVNIGDSARKEPIPEKSTSTSTSKGKGKAVLKPPLEETKGPQTRNAARRRAAIDKDAQMKPVHGDSSSEDEELDISPVEIDDALVIEDDDISDDDDD
DDDDHEDVLRDDSLPVCMPEKVHDVKLGAASEDSNVAPPASDSQSNPASGSSSRAVAVRGSDSTDFRSGSSYGSRGAMSFAAAAMAGLGSANGRGIRGGR
DRQGRPLFGSSSDPPKLIFTAAGKQLNRHLTIYQAIQRQLVLEEDDEDRYGGRDFISSDGSRLWSDIYTLTYQRADGQADRASVGGPSSSASKSIKGGSS
NSNSDTQVHRMSLLDSILQADLPCDLEKSNPTYNILALLRILEGLNQLAPRLRVQLVSDNFSEGKISSLDELMTATGVRVPAEEFINSKLTPKLARQIQD
ALALCSGSLPSWCYQLTKACPFLFPFETRRQYFYSTAFGLSRALYRLQQQQGADGHGSANEREVRVGRLQRQKVRVSRNRILDSAAKVMEMYSSQKAVLE
VEYFGEVGTGLGPTLEFYTLLSHDLQKVTLGMWRSNSAAEKPSMEIDGDDDKNGKSNNESGTAVAADLVQTPLGLFPRPWPPTASASEGSQIYKTIEYFR
LVGRVMAKALQDGRLLDLPLSMAFYKLVLGQELDLYDILSFDAEFGKTLQELHALVCRKHYLESIGSDHEAIADLHFHGTPIEDLCLDFTLPGYPDYILK
PGDETVDINNLEEFISLVVDATVKTGITRQMEAFREGFNQVFDISSLQIFTPQELDYLLCGRRELWEPDTLVDHIKFDHGYTAKSPAIVNLLEIMGEFTP
DQQRAFCQFVTGAPRLPPGGLAVLNPKLTIVRKHSSSAGNAMPNGTGPSESADDDLPSVMTCANYLKLPPYSTKEVMYKKLLYAISEGQGSFDLS
|
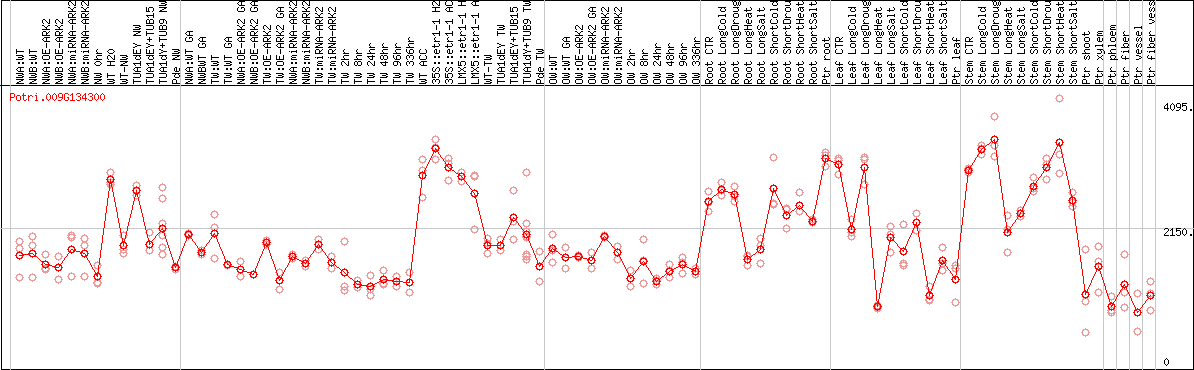
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.009G134300 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
