Potri.009G147100 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.009G147100.2 pacid=42772503 polypeptide=Potri.009G147100.2.p locus=Potri.009G147100 ID=Potri.009G147100.2.v4.1 annot-version=v4.1
ATGATGGAAGAGTCATTAAAGCAGGTAATGAGAAATGATCCTAGGATGGTGCTTGATGAGGGTGCCAAACTTGGGACTGAAGATAAGAAGCAATTCATGG
AGAGTCCACGCAAAATTGTCGAGGAAGACTATGATTACCTGAGGAGGCTAAGGAAAAGAGTTGACAGGGTAGGAATGGAGCTTCCTAGGATTGAGATTCG
GTTTCAGAATTTGTCAGTTGAGGGAGAAGCATATGTTGGGACCCGAGCACTTCCAACTTTGCTTAATACCACCTTGAATGCAGTTGAGGGTGTTGCTCAG
ATGGTTGGGCTTTCTCCATCAAAGAAAAGAGCTGTCAAGATACTTCAAGATGTGAAGGGAATTGTAAAACCTTCAAGGATGTCACTGCTTCTTGGGCCTC
CTGGCTCTGGGAAAACAACTCTGCTAAAAGCACTTGCAGGGAAACTTGACAATGACATAAAGGTAACAGGGAAAGTCACCTACTGCGGCCATGAGTTTTC
AGAATTTGTTCCCCAGAAAACATGTGCTTATATTAGCCAGCATGAACTCCACTATGGGCAGATGACAGTTCGTGAGACATTAGATTTTTCAGGACGATGC
ATGGGAGCTGGGACTAGGCATCAAATACTTTCGGAGTTGTTGAGACGGGAGAAAGAAGCAGGTATCAAACCGAATCCTCGTATTCGTAAGGAAGCTGCAG
CCATGACTTGTCAGGATACCAGTTTGATTACAGAAAACATTCTGAAGATACTTAAATTGGATAGTTGTGCAGATACTAAGGTAGGAGATGACATGATAAG
GGGCATCTCTGGTGGAGAAAAGAAGCGCGTAACTACTGGTGAACTATTGGTTGGACCGGCAAGGGCATTTGTCATGGATGAAATCTCAACAGGGTTGGAC
AGTTCTACCGCTTACCAGATTGTCAAATTCATGAGGAAGATGGTCCATCTCCTGGATATGACCATGGTCACGTCTCTCCTGCAGCCTACACCAGAGACAT
TTGAACTTTTTGATGACATTATCCTTCTTTCAGAGGGTCAGATTGTTTACCAGGGTCCACGAGATAATGTCCTTGAGTTCTTCGAACATATGGGTTTCAA
ATGCCCTGAGAGGAAGGGGGTTGCAGATTTCCTGCAGGAAGTAACTTCCAAGAAGGATCAAGAGCGATACTGGTTCAGAAAGAATCAACCTTACGAGTAT
GTTTCAGTACCGAAGTTCGTGCGCGCTTTCAATTCTTTTCACATTGGCCTACAGCTTTCAGAACATCTAAAAGTTCCTTTCAACAAATTTAGAGTTCATC
CTGATGCTTTAGTGAGTGAAAAGTATGGCGTCTCCAACTGGGAACTCTTTAAGGCGTGCTTTTCAAGGGAATGGCTTCTGATGAAGCGCAACTCTATAGT
TTCCATCTTCAAAATTATCCAAATAACAATCATTGCCATAATTGCATTTACTGCTTTCTCAAAAACCGGAAGGAAAGCTGGACAAAAGAACGGTGCAGCA
AACTTTTGGGGAGCACTATTCTTCGGTCTCACAAATTTCATAATCAATGCGATGATAGAACTTACAATGACAGTTTTCAGACTTCCCGTGTTCTTTAAGC
AGAGGAGTTCAATGCTATATCCTGCATGGGCTTTTGGATTACCTATTTGTCTCTTCAGCATTCCTGTCTCACTGATTGAATCAGGAATTTGGGTCACTCT
TACATATTACTCCATTGGCTTTGCTCCAGCTGCCAGCAGGTTCTTCAAACAGCTCTTGGCTTTCTTCAGTACATATCAGATGACTCTATCTCTATACCGT
TTCATAGCAGTGGTTGGAAGAAAATTGCTTGTCGCGAACATACTTGGTTTCTTAACTATGGTGACTGTTATTGTTCTTGGAGGTTTCATCATCACCAAAG
ATGATATAGAGTTGTGGATGAGATGGGGCTACTATCTGTCTCCTATAATGTATGGCCAAAATGCCATTTCTATCAATGAATTTCTCGACAACAGATGGGG
CAATCTCACTGGCAGCCCACATGAGTCTACTGTTGGAAAGTCCCTTCTTAAGGAAAGGGGATTCTTTACGGATGAATACTGGTATTGGATTTGCATTGGG
GTGCTTCTTGGGTTCTCTCTTATTTTCAATTTTCTCTTCATTGCAGCACTCGAATTTTTTAATGCTCCTGCCGATTCGAGAGCTGTAATTGCGGATGATG
ACACTGAAAATGTTAGTACGCGGAAACTGGCACCTATTTCTGAAGTGAAAATCTCCCGGGGGGAATACAAACATTCCAAGAATCCAAACAAGCAATATAA
AAAAGGAACGGTTCTGCCTTTTCAGCCACTTTCACTTGCCTTCAACAATGTGAATTATTATGTCGATATGCCTGTGGAAACGAGGAAACAAGGGACTGAG
AAAAACCGATTGCAGCTGCTAAAAGATGTGAGTGGAGCTTTTAGGCCAGGTACTCTGACAGCATTAGTTGGTGTCAGCGGTGCTGGTAAGACCACCTTGA
TGGATGTATTGGCAGGAAGGAAGATTATGGGGTACATTGAAGGAAGTATAAGCATTTCTGGATACCCAAAGAACCAAGTCACATTTGCTCGGGTCAGTGG
TTATTGTGAACAGATCGACATGCATTCACCATGTGTTACTGTCTATGAATCTCTCCTCTATTCTGCCTCGATGCGTCTTGCTGCGGATGTCAAGAAGGAA
ACACGAAAGATGTTCATTGATGAAGTTATGGAATTGGTTGAGCTTAAGCCACTGATGAATGCTCTCGTAGGACTTCCACGAATTAACGGTCTTTCAACTG
AGCAGAGAAAGAGACTCACAATAGCTGTAGAGTTGGTTGCTAATCCCTCCATCATTTTTATGGATGAGCCCACATCAGGCCTTGATGCTAGAGCTGCTGC
CATTGTTATGCGTGCTATAAGGCATATGGTGGATACAGGACGAACTGTTGTCTGCACCATTCACCAACCAAGCATTGACATCTTTGAAACTTTTGATGAA
CTTTTGTTGATGAAAAGAGGAGGACAAGTGATATATGCTGGACCTCTTGGTCGCAATTCTCACAAGCTTGTGCAATATTTTGAGGCTGTTCCAGGGGTTC
CCAGGATCAAACAAGGCTCTAATCCTGCTACATGGATGTTGGAGATCAGCTCTGAGGCAATAGAAGCTCAACTTCAAGTTGATTTTGCAGAAGTTTATGC
CAACTCTGAACTTTACCGGAAAAATCAGGAACTCATAAAAAAACTTAGTACTCCACGACCAGGTTCCAAGGATCTTTCTTTCCCCTCTCAATACTCCCAA
AGCTTTATAACTCAATGCACGGCTTGTTTCTGGAAACAACACAAGTCATACTGGCGAAACTCAGAATTCAATTATACCCGGTTCGTTGTAGCAATCATCA
TTGGTATCCTGTTTGGTCTTGTCTTTTGGAGCAGAGGAGATCGAATATACAAAAGAAATGATCTTATTAATCTTCTGGGAGCTACATATGCTGCTGTTCT
TTTCCTTGGAGCCACTAATGCTTCTGCTGTGCAATCTGTCATAGCAACTGAGAGAACAGTTTTCTACCGTGAAAGAGCCGCAGGGATGTATTCAGAGTTG
CCTTATGCTTTTGCTCATGTGGCTATAGAGATAATTTATGTTTCAATCCAAACTTTTCTATATTCTCTTCTTTTGTACTCAATGATTGGGTTTGAATGGA
ATGTGGGCAAGTTCTTGTACTTCTACTACTTCATTTTCATGAGCTTCACCTACTTTTCAATGTATGGGATGATGATTATTTCCCTGACTCCTGGCCCTGA
GATCGCTGCCGTGTTTATGTCCTTCTTCATAAGTTTCTGGAACTTGTTCTCTGGTTATCTCATCGCTAGGCCGCTGATTCCTGTGTGGTGGAGGTGGTAT
TACTGGGCTTCTCCAGTTGCTTGGACCATTTACGGCATTTTCACCTCACAAGTTGTTGATAAAAACACCCTTCTTGAAATTCCTGGATCAGAACCAGTGC
CATTAAAGGCATTCGTTGAGAAGTACTTGGGTTATGATCACGAATTCCTTCTACCTGTAGTCCTAGCTCATGTTGGTTGGGTTCTCCTCTTCTTTTTCGC
CTTCGCATACGGCATCAAGTTCCTTAACTTCCAAAGGAGATGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.009G147100.2 pacid=42772503 polypeptide=Potri.009G147100.2.p locus=Potri.009G147100 ID=Potri.009G147100.2.v4.1 annot-version=v4.1
MMEESLKQVMRNDPRMVLDEGAKLGTEDKKQFMESPRKIVEEDYDYLRRLRKRVDRVGMELPRIEIRFQNLSVEGEAYVGTRALPTLLNTTLNAVEGVAQ
MVGLSPSKKRAVKILQDVKGIVKPSRMSLLLGPPGSGKTTLLKALAGKLDNDIKVTGKVTYCGHEFSEFVPQKTCAYISQHELHYGQMTVRETLDFSGRC
MGAGTRHQILSELLRREKEAGIKPNPRIRKEAAAMTCQDTSLITENILKILKLDSCADTKVGDDMIRGISGGEKKRVTTGELLVGPARAFVMDEISTGLD
SSTAYQIVKFMRKMVHLLDMTMVTSLLQPTPETFELFDDIILLSEGQIVYQGPRDNVLEFFEHMGFKCPERKGVADFLQEVTSKKDQERYWFRKNQPYEY
VSVPKFVRAFNSFHIGLQLSEHLKVPFNKFRVHPDALVSEKYGVSNWELFKACFSREWLLMKRNSIVSIFKIIQITIIAIIAFTAFSKTGRKAGQKNGAA
NFWGALFFGLTNFIINAMIELTMTVFRLPVFFKQRSSMLYPAWAFGLPICLFSIPVSLIESGIWVTLTYYSIGFAPAASRFFKQLLAFFSTYQMTLSLYR
FIAVVGRKLLVANILGFLTMVTVIVLGGFIITKDDIELWMRWGYYLSPIMYGQNAISINEFLDNRWGNLTGSPHESTVGKSLLKERGFFTDEYWYWICIG
VLLGFSLIFNFLFIAALEFFNAPADSRAVIADDDTENVSTRKLAPISEVKISRGEYKHSKNPNKQYKKGTVLPFQPLSLAFNNVNYYVDMPVETRKQGTE
KNRLQLLKDVSGAFRPGTLTALVGVSGAGKTTLMDVLAGRKIMGYIEGSISISGYPKNQVTFARVSGYCEQIDMHSPCVTVYESLLYSASMRLAADVKKE
TRKMFIDEVMELVELKPLMNALVGLPRINGLSTEQRKRLTIAVELVANPSIIFMDEPTSGLDARAAAIVMRAIRHMVDTGRTVVCTIHQPSIDIFETFDE
LLLMKRGGQVIYAGPLGRNSHKLVQYFEAVPGVPRIKQGSNPATWMLEISSEAIEAQLQVDFAEVYANSELYRKNQELIKKLSTPRPGSKDLSFPSQYSQ
SFITQCTACFWKQHKSYWRNSEFNYTRFVVAIIIGILFGLVFWSRGDRIYKRNDLINLLGATYAAVLFLGATNASAVQSVIATERTVFYRERAAGMYSEL
PYAFAHVAIEIIYVSIQTFLYSLLLYSMIGFEWNVGKFLYFYYFIFMSFTYFSMYGMMIISLTPGPEIAAVFMSFFISFWNLFSGYLIARPLIPVWWRWY
YWASPVAWTIYGIFTSQVVDKNTLLEIPGSEPVPLKAFVEKYLGYDHEFLLPVVLAHVGWVLLFFFAFAYGIKFLNFQRR
|
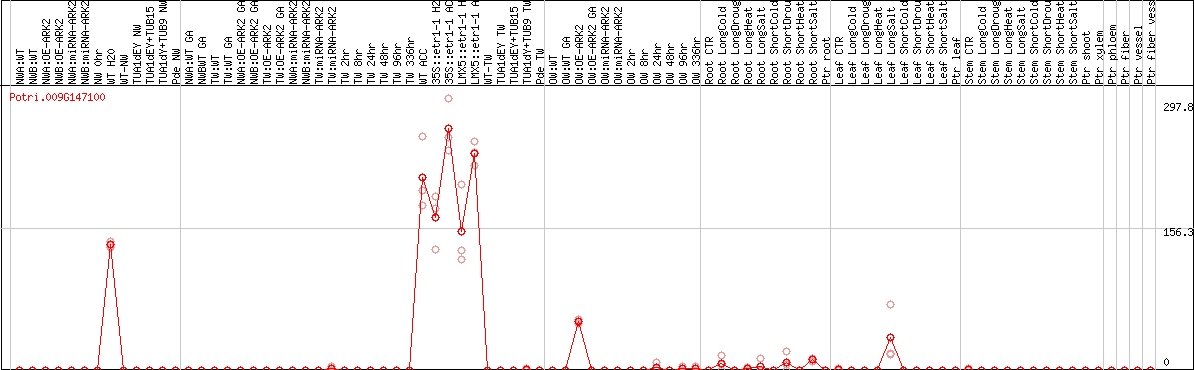
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.009G147100 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
