External link
Symbol
Pt-ATGB2.1
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G35860 412 / 6e-149
ATGB2, AtRABB1b, Atrab2C
GTP-binding 2 (.1.2)
AT4G17170 382 / 5e-137
ATRAB-B1B, AT-RAB2, AtRABB1c, AtRab2A
ARABIDOPSIS RAB GTPASE HOMOLOG B1B, RAB GTPase homolog B1C (.1)
AT4G17160 312 / 2e-109
AtRab2B, AtRABB1a
RAB GTPase homolog B1A (.1)
AT1G06400 204 / 2e-66
ARA2, AtRABA1a, AtRab11E, Ara-2
ARABIDOPSIS THALIANA RAB GTPASE HOMOLOG A1A, Ras-related small GTP-binding family protein (.1)
AT5G45750 203 / 2e-66
AtRABA1c
RAB GTPase homolog A1C (.1)
AT1G07410 196 / 2e-63
ATRAB-A2B, AtRABA2b
ARABIDOPSIS RAB GTPASE HOMOLOG A2B, RAB GTPase homolog A2B (.1)
AT5G60860 196 / 2e-63
AtRABA1f
RAB GTPase homolog A1F (.1)
AT1G02130 194 / 7e-63
ARA5, AtRABD2a, AtRab1B, Ara-5
ARABIDOPSIS THALIANA RAB D2A, ARABIDOPSIS THALIANA RESPONSIVE TO ABSCISIC ACID 1B, ARABIDOPSIS RAS 5, RAS 5 (.1)
AT3G09900 194 / 9e-63
AtRABE1e, AtRab8E
RAB GTPase homolog E1E (.1)
AT4G18430 194 / 1e-62
AtRABA1e
RAB GTPase homolog A1E (.1)
Paralogs
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10038744
381 / 1e-136
AT4G17170 434 / 2e-157
ARABIDOPSIS RAB GTPASE HOMOLOG B1B, RAB GTPase homolog B1C (.1)
Lus10039114
356 / 8e-127
AT4G17170 407 / 5e-147
ARABIDOPSIS RAB GTPASE HOMOLOG B1B, RAB GTPase homolog B1C (.1)
Lus10025184
355 / 4e-126
AT4G17170 389 / 1e-139
ARABIDOPSIS RAB GTPASE HOMOLOG B1B, RAB GTPase homolog B1C (.1)
Lus10016066
342 / 1e-120
AT4G17170 369 / 3e-131
ARABIDOPSIS RAB GTPASE HOMOLOG B1B, RAB GTPase homolog B1C (.1)
Lus10020746
204 / 9e-67
AT1G06400 375 / 3e-134
ARABIDOPSIS THALIANA RAB GTPASE HOMOLOG A1A, Ras-related small GTP-binding family protein (.1)
Lus10029253
204 / 2e-66
AT5G45750 393 / 4e-141
RAB GTPase homolog A1C (.1)
Lus10029789
202 / 6e-66
AT1G06400 373 / 3e-133
ARABIDOPSIS THALIANA RAB GTPASE HOMOLOG A1A, Ras-related small GTP-binding family protein (.1)
Lus10007306
201 / 2e-65
AT5G45750 387 / 5e-139
RAB GTPase homolog A1C (.1)
Lus10025432
199 / 2e-64
AT5G60860 422 / 1e-152
RAB GTPase homolog A1F (.1)
Lus10015297
199 / 2e-64
AT5G60860 428 / 6e-155
RAB GTPase homolog A1F (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0023
P-loop_NTPase
PF00025
Arf
ADP-ribosylation factor family
Representative CDS sequence
>Potri.009G159600.2 pacid=42771475 polypeptide=Potri.009G159600.2.p locus=Potri.009G159600 ID=Potri.009G159600.2.v4.1 annot-version=v4.1
ATGTCTTACGATTACCTCTTCAAGTACATAATCATCGGTGATACAGGCGTAGGGAAGTCATGCCTGCTCTTGCAATTCACTGACAAAAGATTCCAGCCTG
TTCATGATCTCACCATCGGTGTTGAATTCGGAGCTCGCATGGTCACCATCGATGCCCGCCCTATTAAGCTTCAAATTTGGGACACCGCTGGGCAAGAGTC
CTTCAGATCCATCACTAGATCTTACTACAGAGGAGCAGCCGGAGCACTTCTGGTTTATGACATTACCAGGAGAGAGACGTTTAATCACCTAGCAAGCTGG
CTGGAGGATGCTCGGCAGCATGCAAATCCCAACATGTCAATTATGCTTATTGGGAACAAGTGTGATCTTGCTCATCGTAGGGCTGTTAGCAAAGAGGAAG
GGGAACAGTTTGCAAAAGAAAATGGACTTCTGTTCTTGGAGGCATCTGCAAGAACAGCTCAAAATGTTGAGGAGGCCTTCATAGGGACTGCTGGAAAAAT
CCTTCAGAATATCCAGGAAGGTGTGTTTGATGTATCCAATGAGTCTTCAGGGATCAAGGTTGGATATGGGCGCCCCCAAGGTGCATCAGGTGCAAGGGAT
GGAACTGTTGCTCAAAGAGGGGGGTGCTGCAGCTAA
AA sequence
>Potri.009G159600.2 pacid=42771475 polypeptide=Potri.009G159600.2.p locus=Potri.009G159600 ID=Potri.009G159600.2.v4.1 annot-version=v4.1
MSYDYLFKYIIIGDTGVGKSCLLLQFTDKRFQPVHDLTIGVEFGARMVTIDARPIKLQIWDTAGQESFRSITRSYYRGAAGALLVYDITRRETFNHLASW
LEDARQHANPNMSIMLIGNKCDLAHRRAVSKEEGEQFAKENGLLFLEASARTAQNVEEAFIGTAGKILQNIQEGVFDVSNESSGIKVGYGRPQGASGARD
GTVAQRGGCCS
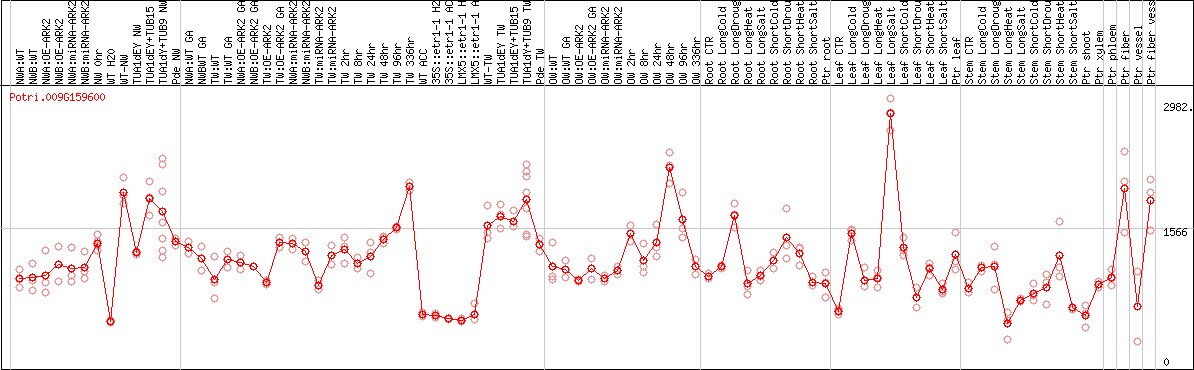
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.009G159600 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.