Pt-GLUR2.1 (Potri.009G168300) [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | Pt-GLUR2.1 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.009G168300.2 pacid=42772042 polypeptide=Potri.009G168300.2.p locus=Potri.009G168300 ID=Potri.009G168300.2.v4.1 annot-version=v4.1
ATGAATCTGGCCTGGCTTTTGTCCTTCTGGATTCTCTGTACCAGTTCTTTTTCACAAGGGGCTTTAAGCCCTGGTGGTACTGTGAATGTTGGAGCTATTT
TTACATTTAGCTCCATCAATGGAAGAGTCGCAAAGATTGCCATGGAGGCAGCTGAGGATGATATCAATTCTGATCCAAGCCTTCTTGGAGGAAGGAAGTT
GTCTATAAATATGCATGATTCCAACTTCAGTGGATTTCTGGGAATCATTGGAGCTTTGCAATTCCTGGAGACTGATACTGTAGCTGTTATTGGTCCACAA
ACTGCTGTGATGGCTCATGTACTCTCACATCTTGCTAATGAGCTCCAAGTTCCTTTCTTGTCTTTCACAGCTCTAGATCCCACTCTTTCACCTCTCCAAT
TCCCATACTTTATTCAAACAGCTCCCAACGATTTGTTTCAAATGACTGCCATTGCAGACATAGTTAGTTATTATGGTTGGTCTGAGGTTACTGCAGTTTT
CAATGATGATGATCAGAACCGAAATGGTATAACTGTACTAGGTGATAAACTTGCGGAAAGGCGTTGCAAAATTTCCTATAAAGCAGCACTTCCTCCAGAA
CCAAAAGCCACCCGAAGCAATATTCAAGATGAGTTAGCTAAGATTCTGGGGATGGAATCTCGAGTAATTGTGTTGAATACCTTCAGCAAAACAGGTCTCT
TAGTTTTTGATGTTGCAAAGGCACTTGGGATGATGGAGAATGGGTTTGTTTGGATAGTTACCAGTTGGCTGTCTACTGTTATAGATTCAGCTTCACCTCT
TCCTACTACTGCCAACTCAATCCAGGGAGTTTTAGCACTCCGTCCACATACACCAGATTCAAAAAGAAAAAGGGATTTCATCTCACGGTGGAAACAGCTG
AGTAATGGCTCTATTGGGCTAAATCCTTATGGTCTATATGCCTATGATACTGTTTGGTTACTTGCACGCGCATTGAAATCGTTTTTCGATCAGGGAAACA
CCATTTCTTTTACCAATGATTCGAGATTAGGTGGAATAGGAGGGGGATATTTGAACCTTGGTGCATTGAGCATATTTGATGGAGGAAGCCAATTGCTTAA
AAATATATTGCAAACAAGTATGACAGGTCTGACAGGACCTTTTCGATTTAATCCAGACAGGTCCATTTTACATCCCTCTTATGATATCATTAATGTATTA
GAAACTGGGTATCAGCAGGTTGGATACTGGTCAAACTATTCTGGGCTATCTGTTGTGCCCCCAGAGACACTTTATGGAAAGGCAGCTAACCGTTCTAGTT
CAAGCCAACATTTACAGAGTGTGGTATGGCCTGGGGGGACAACAGCTAGGCCACGGGGCTGGGTGTTTCCAAACAATGGAAAGGAATTACAAATTGGAAT
CCCAAACCGAGTTAGTTATCGGGATTTTGTCTCAAAAGTGAATGGCACCGACATGGTCCAAGGATATTGCATAGATGTATTTCTTGCTGCCATTAAATTG
CTTCCATACGCTGTTCCACACAAGTTCATTCCATTTGGAGATGGTCATAAGAATCCGACATACTACGATCTTGTTTACAAGATCACAACCAGGGTCTTTG
ATGCTGTTATCGGTGATGTTGCTATTGTCACCAACCGGACAAAGATTGTCGATTTCACTCAGCCATACATAGAGTCAGGGCTAGTTGTGGTGGCTCCGGT
CAAGAAGCGAAATTCAAATGCTTGGGCATTCTTGCGGCCTTTTAGTCCATTGATGTGGGCTGTCACGGCAATGTTTTTCCTTATTGTGGGAGCTGTTGTA
TGGATCCTTGAGCATAGAATTAATGATGAATTCCGAGGGCCACCTAGGAAACAGCTTGTCACAATTCTCTGGTTTAGCTTCTCCACTTTGTTTTTCTCCC
ACAGAGAAAATACTGTGAGCACACTTGGTCGCCTTGTGCTGATAATCTGGCTTTTTGTAGTTTTGATAATCAACTCAAGCTATACTGCAAGTCTGACATC
AATTCTCACAGTGCAACAGTTATCCTCCACCATTAAAGGAATTGATTCCTTGATAACTAGCAATGCTCAAATTGGATTCCAAGTAGGATCCTTTGCTGAA
AATTATCTAAATGAAGAACTCAGCATTGCTAAAACTAGACTTGTTCCTCTTGGTTCACCAGAAGAATATGCGGATGCCCTAAAAAATGGTACTGTTGCTG
CCGTGGTTGATGAGCGACCTTATGTAGACCTCTTTCTCTCAGAGCACTGTGAGTTCTCAATAATAGGTCAAGAGTTCACCAGAAGTGGTTGGGGATTTGC
ATTTCCAAGAGATTCACCCTTGGCCATTGATATGTCTACTGCCATTCTCCAACTATCTGAGAATGGCGAACTTCAGAACATCCATAACAAGTGGTTGCAA
AGAAAGCTCTGCAGTTCCCAGGACATTGGGTCGAGTGCAGATCAGCTACAGTTACAAAGCTTCTGGGGGCTTTTCCTCATTTGCGGGATTGCATGTTTGC
TTGCTCTCTTGATCTACTTTTGCACCACATTTCGCCAATTCAGCCGGCACTTCCCCGAAGAATCCGATTCTTCTGTCCAAAGTCGCTCTCGCTCCAAACG
TCTTCAAACATTCTTATCATTTGCTGATGACAAGGTGGAACAATGGAAGAAGAGCAAGTCCAAGAGAAAAAGAGAAGATGAATTATCAAATCGGAGTGGG
GAGGGAAGCATGTCAGTTAATAGATCTGAAAGAATCCAAAGGGACATTTCTCAGGAAAGAGAAAATGGTGATACTTGGCTGCATTAA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.009G168300.2 pacid=42772042 polypeptide=Potri.009G168300.2.p locus=Potri.009G168300 ID=Potri.009G168300.2.v4.1 annot-version=v4.1
MNLAWLLSFWILCTSSFSQGALSPGGTVNVGAIFTFSSINGRVAKIAMEAAEDDINSDPSLLGGRKLSINMHDSNFSGFLGIIGALQFLETDTVAVIGPQ
TAVMAHVLSHLANELQVPFLSFTALDPTLSPLQFPYFIQTAPNDLFQMTAIADIVSYYGWSEVTAVFNDDDQNRNGITVLGDKLAERRCKISYKAALPPE
PKATRSNIQDELAKILGMESRVIVLNTFSKTGLLVFDVAKALGMMENGFVWIVTSWLSTVIDSASPLPTTANSIQGVLALRPHTPDSKRKRDFISRWKQL
SNGSIGLNPYGLYAYDTVWLLARALKSFFDQGNTISFTNDSRLGGIGGGYLNLGALSIFDGGSQLLKNILQTSMTGLTGPFRFNPDRSILHPSYDIINVL
ETGYQQVGYWSNYSGLSVVPPETLYGKAANRSSSSQHLQSVVWPGGTTARPRGWVFPNNGKELQIGIPNRVSYRDFVSKVNGTDMVQGYCIDVFLAAIKL
LPYAVPHKFIPFGDGHKNPTYYDLVYKITTRVFDAVIGDVAIVTNRTKIVDFTQPYIESGLVVVAPVKKRNSNAWAFLRPFSPLMWAVTAMFFLIVGAVV
WILEHRINDEFRGPPRKQLVTILWFSFSTLFFSHRENTVSTLGRLVLIIWLFVVLIINSSYTASLTSILTVQQLSSTIKGIDSLITSNAQIGFQVGSFAE
NYLNEELSIAKTRLVPLGSPEEYADALKNGTVAAVVDERPYVDLFLSEHCEFSIIGQEFTRSGWGFAFPRDSPLAIDMSTAILQLSENGELQNIHNKWLQ
RKLCSSQDIGSSADQLQLQSFWGLFLICGIACLLALLIYFCTTFRQFSRHFPEESDSSVQSRSRSKRLQTFLSFADDKVEQWKKSKSKRKREDELSNRSG
EGSMSVNRSERIQRDISQERENGDTWLH
|
DESeq2's median of ratios [POPLAR]
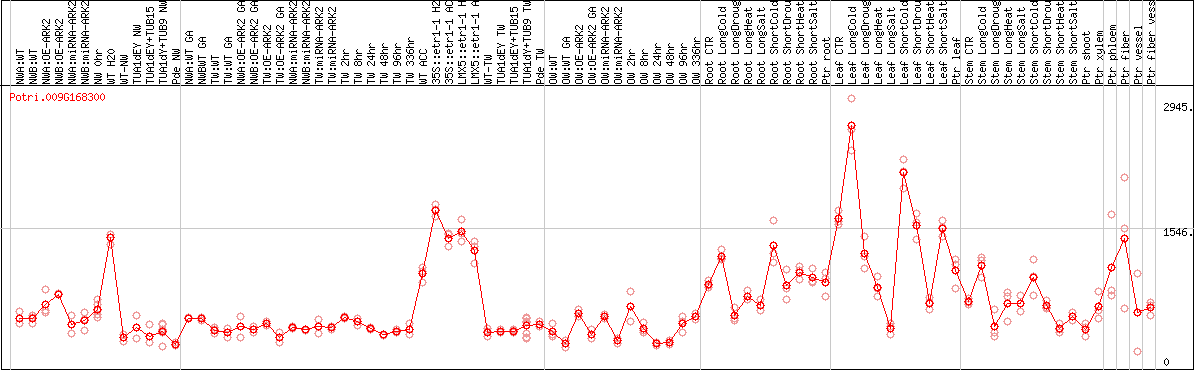
Coexpressed genes
Potri.009G168300 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
