External link
Symbol
HY5.1
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G17609 133 / 8e-40
bZIP
HYH
HY5-homolog (.1.2.3.4)
AT5G11260 97 / 3e-25
bZIP
TED5, HY5
REVERSAL OF THE DET PHENOTYPE 5, ELONGATED HYPOCOTYL 5, Basic-leucine zipper (bZIP) transcription factor family protein (.1)
AT2G40950 49 / 1e-06
bZIP
BZIP17
Basic-leucine zipper (bZIP) transcription factor family protein (.1)
AT3G56660 45 / 2e-05
bZIP
BZIP49
basic region/leucine zipper motif protein 49 (.1)
AT1G03970 42 / 0.0001
bZIP
GBF4
G-box binding factor 4 (.1)
AT5G44080 41 / 0.0004
bZIP
Basic-leucine zipper (bZIP) transcription factor family protein (.1)
AT2G36270 40 / 0.0005
bZIP
EEL, GIA1, ABI5
GROWTH-INSENSITIVITY TO ABA 1, ABA INSENSITIVE 5, Basic-leucine zipper (bZIP) transcription factor family protein (.1)
AT4G34000 40 / 0.0006
bZIP
AtABF3, ABF3, DPBF5
DC3 PROMOTER-BINDING FACTOR 5, abscisic acid responsive elements-binding factor 3 (.1.2.3)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.018G029500
102 / 3e-27
AT5G11260 185 / 2e-60
REVERSAL OF THE DET PHENOTYPE 5, ELONGATED HYPOCOTYL 5, Basic-leucine zipper (bZIP) transcription factor family protein (.1)
Potri.006G251800
102 / 3e-27
AT5G11260 143 / 6e-44
REVERSAL OF THE DET PHENOTYPE 5, ELONGATED HYPOCOTYL 5, Basic-leucine zipper (bZIP) transcription factor family protein (.1)
Potri.006G034500
50 / 6e-07
AT2G40950 475 / 1e-157
Basic-leucine zipper (bZIP) transcription factor family protein (.1)
Potri.016G032400
44 / 4e-05
AT2G40950 498 / 9e-167
Basic-leucine zipper (bZIP) transcription factor family protein (.1)
Potri.001G374200
40 / 0.0008
AT3G58120 282 / 6e-94
Basic-leucine zipper (bZIP) transcription factor family protein (.1)
Potri.005G192900
40 / 0.0009
AT5G44080 142 / 5e-40
Basic-leucine zipper (bZIP) transcription factor family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10002900
101 / 8e-27
AT5G11260 197 / 3e-65
REVERSAL OF THE DET PHENOTYPE 5, ELONGATED HYPOCOTYL 5, Basic-leucine zipper (bZIP) transcription factor family protein (.1)
Lus10002028
98 / 2e-25
AT5G11260 182 / 5e-59
REVERSAL OF THE DET PHENOTYPE 5, ELONGATED HYPOCOTYL 5, Basic-leucine zipper (bZIP) transcription factor family protein (.1)
Lus10038995
54 / 4e-08
AT2G40950 421 / 2e-137
Basic-leucine zipper (bZIP) transcription factor family protein (.1)
Lus10027290
54 / 4e-08
AT2G40950 429 / 2e-140
Basic-leucine zipper (bZIP) transcription factor family protein (.1)
Lus10012104
50 / 6e-07
AT2G40950 454 / 9e-150
Basic-leucine zipper (bZIP) transcription factor family protein (.1)
Lus10010440
50 / 7e-07
AT2G40950 466 / 5e-155
Basic-leucine zipper (bZIP) transcription factor family protein (.1)
Lus10040069
43 / 9e-05
AT1G42990 84 / 4e-18
Bbasic region/leucine zipper motif 60, basic region/leucine zipper motif 60 (.1)
Lus10002399
42 / 0.0002
AT3G19290 203 / 2e-61
ABA-RESPONSIVE ELEMENT BINDING PROTEIN 2, ABRE binding factor 4 (.1.2.3)
Lus10027897
42 / 0.0003
AT3G19290 289 / 1e-93
ABA-RESPONSIVE ELEMENT BINDING PROTEIN 2, ABRE binding factor 4 (.1.2.3)
Lus10028575
41 / 0.0003
AT5G44080 191 / 2e-59
Basic-leucine zipper (bZIP) transcription factor family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0018
bZIP
PF00170
bZIP_1
bZIP transcription factor
Representative CDS sequence
>Potri.010G004200.1 pacid=42799476 polypeptide=Potri.010G004200.1.p locus=Potri.010G004200 ID=Potri.010G004200.1.v4.1 annot-version=v4.1
ATGGATATTGGTGAATCCAACAACAAGTACCCTCATGATCATAAGAAGCAATTACTTGATGACGATGACGACGACGACGACGACGACGACGACCATCAGG
TGGAGGGCACTAGTGCTACCGCTACCTCTTCGTCTTCCTCTTCTTGGAGGATCAGGAGAAGTACTGGTAGTACAAGTGACAATCTCATTTCACACCACAA
AAACAAATTAGTAGATCATGAAGACAGTGATGAAGATCTTTTCACGGTTCCGGATGTAGAAGCTCGACCACCAGCGGAGGCTGCTGCTGCTGCTGCTGGT
AATAATACTACTACTTACAATAATAAGAACATTAATTCTAATAATCCAGAGGTTCAATCCGCCGCAGCAGCACCAGCTGCAACAGGGAGTAATAAACGCC
ATAGAGGACGAAACCCTGTTGACAAAGAATATAGAAGGCTCAAGAGCAGATTGCTAAGGAACAGGGTCTCAGCCCAACAAGCTCGAGAAAGAAAGAAGGT
TTACGTCAATGACTTGGAATCAAGAGCCAAAGAATTGCAGGAAATTAATACCAAATTAGAGGAGAAGATCTCCACATTGACCAATGAAAACACCATGCTT
CGGAAGGTCCTCATGAATACCAGGCCTAAGGTTGATGAAAGCATGGAGCAAAAGCAGTAA
AA sequence
>Potri.010G004200.1 pacid=42799476 polypeptide=Potri.010G004200.1.p locus=Potri.010G004200 ID=Potri.010G004200.1.v4.1 annot-version=v4.1
MDIGESNNKYPHDHKKQLLDDDDDDDDDDDDHQVEGTSATATSSSSSSWRIRRSTGSTSDNLISHHKNKLVDHEDSDEDLFTVPDVEARPPAEAAAAAAG
NNTTTYNNKNINSNNPEVQSAAAAPAATGSNKRHRGRNPVDKEYRRLKSRLLRNRVSAQQARERKKVYVNDLESRAKELQEINTKLEEKISTLTNENTML
RKVLMNTRPKVDESMEQKQ
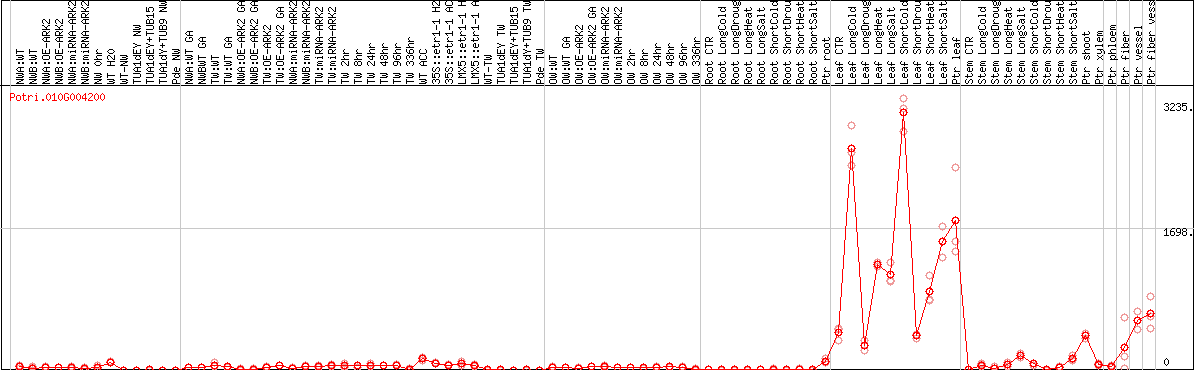
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.010G004200 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.