External link
Symbol
POR1.2
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G01280 390 / 5e-138
VDAC1, ATVDAC1
ARABIDOPSIS THALIANA VOLTAGE DEPENDENT ANION CHANNEL 1, voltage dependent anion channel 1 (.1)
AT5G15090 368 / 2e-129
VDAC3, ATVDAC3
ARABIDOPSIS THALIANA VOLTAGE DEPENDENT ANION CHANNEL 3, voltage dependent anion channel 3 (.1.2)
AT5G67500 266 / 2e-89
VDAC2, ATVDAC2
ARABIDOPSIS THALIANA VOLTAGE DEPENDENT ANION CHANNEL 2, voltage dependent anion channel 2 (.1.2)
AT5G57490 252 / 7e-84
VDAC4, ATVDAC4
ARABIDOPSIS THALIANA VOLTAGE DEPENDENT ANION CHANNEL 4, voltage dependent anion channel 4 (.1)
AT3G49920 162 / 3e-49
VDAC5, ATVDAC5
ARABIDOPSIS THALIANA VOLTAGE DEPENDENT ANION CHANNEL 5, voltage dependent anion channel 5 (.1.2)
AT5G37610 76 / 2e-16
Eukaryotic porin family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.008G194900
498 / 0
AT3G01280 418 / 5e-149
ARABIDOPSIS THALIANA VOLTAGE DEPENDENT ANION CHANNEL 1, voltage dependent anion channel 1 (.1)
Potri.017G078200
416 / 2e-148
AT3G01280 414 / 9e-148
ARABIDOPSIS THALIANA VOLTAGE DEPENDENT ANION CHANNEL 1, voltage dependent anion channel 1 (.1)
Potri.006G169400
273 / 9e-92
AT5G57490 394 / 1e-139
ARABIDOPSIS THALIANA VOLTAGE DEPENDENT ANION CHANNEL 4, voltage dependent anion channel 4 (.1)
Potri.005G146800
269 / 2e-90
AT5G67500 416 / 2e-148
ARABIDOPSIS THALIANA VOLTAGE DEPENDENT ANION CHANNEL 2, voltage dependent anion channel 2 (.1.2)
Potri.018G093900
269 / 2e-90
AT5G57490 387 / 9e-137
ARABIDOPSIS THALIANA VOLTAGE DEPENDENT ANION CHANNEL 4, voltage dependent anion channel 4 (.1)
Potri.007G055800
260 / 9e-87
AT5G67500 405 / 7e-144
ARABIDOPSIS THALIANA VOLTAGE DEPENDENT ANION CHANNEL 2, voltage dependent anion channel 2 (.1.2)
Potri.014G016300
236 / 1e-77
AT5G67500 344 / 5e-120
ARABIDOPSIS THALIANA VOLTAGE DEPENDENT ANION CHANNEL 2, voltage dependent anion channel 2 (.1.2)
Potri.012G070300
174 / 2e-53
AT3G01280 163 / 3e-49
ARABIDOPSIS THALIANA VOLTAGE DEPENDENT ANION CHANNEL 1, voltage dependent anion channel 1 (.1)
Potri.001G294100
172 / 1e-51
AT5G67500 225 / 3e-72
ARABIDOPSIS THALIANA VOLTAGE DEPENDENT ANION CHANNEL 2, voltage dependent anion channel 2 (.1.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10013271
463 / 5e-167
AT3G01280 400 / 3e-142
ARABIDOPSIS THALIANA VOLTAGE DEPENDENT ANION CHANNEL 1, voltage dependent anion channel 1 (.1)
Lus10030794
447 / 3e-160
AT3G01280 379 / 3e-133
ARABIDOPSIS THALIANA VOLTAGE DEPENDENT ANION CHANNEL 1, voltage dependent anion channel 1 (.1)
Lus10015357
380 / 3e-134
AT3G01280 385 / 5e-136
ARABIDOPSIS THALIANA VOLTAGE DEPENDENT ANION CHANNEL 1, voltage dependent anion channel 1 (.1)
Lus10007267
380 / 4e-134
AT3G01280 379 / 1e-133
ARABIDOPSIS THALIANA VOLTAGE DEPENDENT ANION CHANNEL 1, voltage dependent anion channel 1 (.1)
Lus10037212
310 / 6e-106
AT3G01280 286 / 1e-96
ARABIDOPSIS THALIANA VOLTAGE DEPENDENT ANION CHANNEL 1, voltage dependent anion channel 1 (.1)
Lus10036716
281 / 4e-94
AT3G01280 265 / 6e-88
ARABIDOPSIS THALIANA VOLTAGE DEPENDENT ANION CHANNEL 1, voltage dependent anion channel 1 (.1)
Lus10019284
242 / 1e-79
AT5G67500 362 / 9e-127
ARABIDOPSIS THALIANA VOLTAGE DEPENDENT ANION CHANNEL 2, voltage dependent anion channel 2 (.1.2)
Lus10011539
222 / 2e-71
AT5G67500 340 / 4e-118
ARABIDOPSIS THALIANA VOLTAGE DEPENDENT ANION CHANNEL 2, voltage dependent anion channel 2 (.1.2)
Lus10007739
217 / 6e-70
AT5G67500 317 / 5e-109
ARABIDOPSIS THALIANA VOLTAGE DEPENDENT ANION CHANNEL 2, voltage dependent anion channel 2 (.1.2)
Lus10018683
213 / 2e-68
AT5G67500 309 / 6e-106
ARABIDOPSIS THALIANA VOLTAGE DEPENDENT ANION CHANNEL 2, voltage dependent anion channel 2 (.1.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0193
MBB
PF01459
Porin_3
Eukaryotic porin
Representative CDS sequence
>Potri.010G033500.1 pacid=42797284 polypeptide=Potri.010G033500.1.p locus=Potri.010G033500 ID=Potri.010G033500.1.v4.1 annot-version=v4.1
ATGGGAAAAGGCCCAGGTCTCTACTTCGATATTGGCAAGAAAGCCAGAGATCTTCTCTACAAGGATTACCAGAGCGACCACAAGTTCACTGTCACTACTT
ACACTTCCGCTGGAGTTGCCATTACTTCAACTGGGATCAAGAAAGGGGAGCTGTTTTTAGCGGATATCAGCAGTCAGCTAAAGAACAAGAACATCACAAC
TGATGTAAAAGTGGACACAAACTCCAATCTTCTCACAACTATTACTATTGACGAACCTGCTCCTGGGCTCAAGACAATCTTTAGCTTCAAAGTTCCTGAT
CAGAGATCTGGCAAGGTGGAACTCCAGTACCAGCATGAGTATGCTGGAATTAGTACTAGCCTTGGGTTGACTGCCAATCCCATTGTTAACTTCTCAGGTG
TGGTTGGGAGCAATGTTGTTGCTCTGGGGACTGATCTCTCATTTGATACTGCCACTGGGAACTTCACTAAATGCAACGCAGGGTTGAGTTATACGAATTC
TGACCTAATTGCTTCCCTAACAGTTAATGATAAGGGAGACACCCTTAGTGCTTCCTACTACCACACAGTGAGTCCATTGACTAGTACAGCTGTTGGTGCG
GAGCTGACCCATAGCTTTTCAAGCAATGAAAACATATTGACCATTGGCACGCAGCATGCACTTGACCCGCTGACTACAGTGAAGGCTCGGTTGAACAACT
ATGGGAAAGTGAGTGCTCTTATCCAGAATGAGTGGCGCCCTAAGTCTCTATTCACAATCTCAGGGGAGGTTGATACGAAGGCAATTGAAAAGAGTGCTAA
GGTTGGATTGGCCTTGTCTCTCAAGCCCTAG
AA sequence
>Potri.010G033500.1 pacid=42797284 polypeptide=Potri.010G033500.1.p locus=Potri.010G033500 ID=Potri.010G033500.1.v4.1 annot-version=v4.1
MGKGPGLYFDIGKKARDLLYKDYQSDHKFTVTTYTSAGVAITSTGIKKGELFLADISSQLKNKNITTDVKVDTNSNLLTTITIDEPAPGLKTIFSFKVPD
QRSGKVELQYQHEYAGISTSLGLTANPIVNFSGVVGSNVVALGTDLSFDTATGNFTKCNAGLSYTNSDLIASLTVNDKGDTLSASYYHTVSPLTSTAVGA
ELTHSFSSNENILTIGTQHALDPLTTVKARLNNYGKVSALIQNEWRPKSLFTISGEVDTKAIEKSAKVGLALSLKP
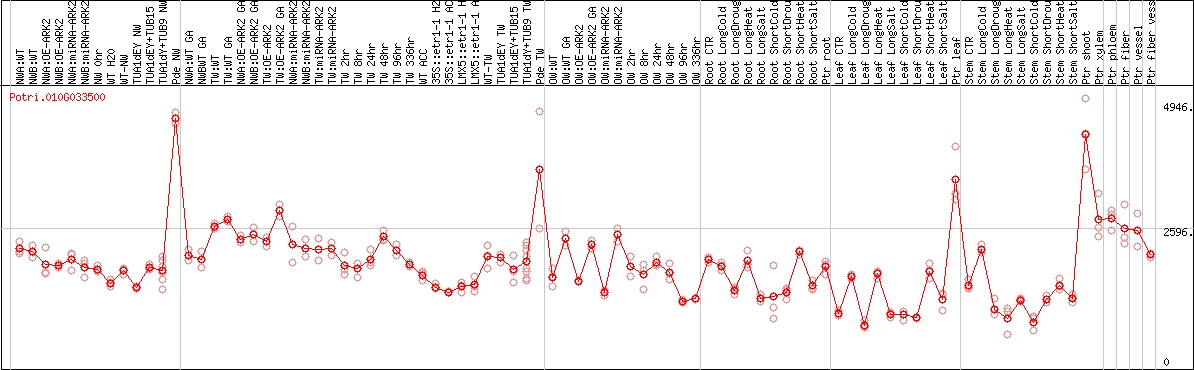
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.010G033500 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.