External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G70460 257 / 3e-82
AtPERK13, RHS10
proline-rich extensin-like receptor kinase 13, root hair specific 10 (.1)
AT1G23540 257 / 3e-82
IGI1, AtPERK12
proline-rich extensin-like receptor kinase 12, INFLORESCENCE GROWTH INHIBITOR 1, proline-rich extensin like receptor kinase, Protein kinase superfamily protein (.1)
AT1G10620 232 / 1e-72
AtPERK11
proline-rich extensin-like receptor kinase 11, Protein kinase superfamily protein (.1)
AT1G70450 218 / 2e-70
Protein kinase superfamily protein (.1)
AT1G68690 213 / 1e-65
AtPERK9
proline-rich extensin-like receptor kinase 9, Protein kinase superfamily protein (.1)
AT1G26150 213 / 2e-65
ATPERK10
proline-rich extensin-like receptor kinase 10 (.1)
AT5G38560 212 / 2e-65
AtPERK8
proline-rich extensin-like receptor kinase 8, Protein kinase superfamily protein (.1)
AT3G24550 181 / 4e-54
ATPERK1
proline-rich extensin-like receptor kinase 1, proline extensin-like receptor kinase 1 (.1)
AT1G52290 177 / 1e-53
AtPERK15
proline-rich extensin-like receptor kinase 15, Protein kinase superfamily protein (.1)
AT2G18470 171 / 2e-50
AtPERK4, PERK4
proline-rich extensin-like receptor kinase 4, roline-rich extensin-like receptor kinase 4 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.008G189700
306 / 4e-101
AT1G23540 642 / 0.0
proline-rich extensin-like receptor kinase 12, INFLORESCENCE GROWTH INHIBITOR 1, proline-rich extensin like receptor kinase, Protein kinase superfamily protein (.1)
Potri.010G132900
219 / 2e-67
AT1G26150 605 / 0.0
proline-rich extensin-like receptor kinase 10 (.1)
Potri.008G111600
218 / 2e-67
AT1G68690 622 / 0.0
proline-rich extensin-like receptor kinase 9, Protein kinase superfamily protein (.1)
Potri.004G105200
215 / 3e-66
AT5G38560 650 / 0.0
proline-rich extensin-like receptor kinase 8, Protein kinase superfamily protein (.1)
Potri.017G110400
214 / 6e-66
AT5G38560 654 / 0.0
proline-rich extensin-like receptor kinase 8, Protein kinase superfamily protein (.1)
Potri.018G081300
195 / 4e-59
AT3G24550 629 / 0.0
proline-rich extensin-like receptor kinase 1, proline extensin-like receptor kinase 1 (.1)
Potri.005G124400
182 / 1e-54
AT2G18470 597 / 0.0
proline-rich extensin-like receptor kinase 4, roline-rich extensin-like receptor kinase 4 (.1)
Potri.007G027000
175 / 6e-52
AT2G18470 599 / 0.0
proline-rich extensin-like receptor kinase 4, roline-rich extensin-like receptor kinase 4 (.1)
Potri.001G183000
164 / 3e-48
AT3G24550 560 / 0.0
proline-rich extensin-like receptor kinase 1, proline extensin-like receptor kinase 1 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10030879
252 / 4e-82
AT1G70460 542 / 0.0
proline-rich extensin-like receptor kinase 13, root hair specific 10 (.1)
Lus10013014
244 / 5e-80
AT1G23540 572 / 0.0
proline-rich extensin-like receptor kinase 12, INFLORESCENCE GROWTH INHIBITOR 1, proline-rich extensin like receptor kinase, Protein kinase superfamily protein (.1)
Lus10029151
243 / 1e-79
AT1G23540 575 / 0.0
proline-rich extensin-like receptor kinase 12, INFLORESCENCE GROWTH INHIBITOR 1, proline-rich extensin like receptor kinase, Protein kinase superfamily protein (.1)
Lus10013013
243 / 2e-76
AT1G70460 596 / 0.0
proline-rich extensin-like receptor kinase 13, root hair specific 10 (.1)
Lus10030610
234 / 1e-73
AT1G23540 354 / 9e-112
proline-rich extensin-like receptor kinase 12, INFLORESCENCE GROWTH INHIBITOR 1, proline-rich extensin like receptor kinase, Protein kinase superfamily protein (.1)
Lus10034082
224 / 2e-69
AT5G38560 700 / 0.0
proline-rich extensin-like receptor kinase 8, Protein kinase superfamily protein (.1)
Lus10003066
221 / 2e-68
AT5G38560 707 / 0.0
proline-rich extensin-like receptor kinase 8, Protein kinase superfamily protein (.1)
Lus10023645
190 / 5e-57
AT3G24550 691 / 0.0
proline-rich extensin-like receptor kinase 1, proline extensin-like receptor kinase 1 (.1)
Lus10011098
184 / 6e-55
AT3G24550 700 / 0.0
proline-rich extensin-like receptor kinase 1, proline extensin-like receptor kinase 1 (.1)
Lus10043219
181 / 4e-54
AT3G24550 688 / 0.0
proline-rich extensin-like receptor kinase 1, proline extensin-like receptor kinase 1 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0016
PKinase
PF07714
PK_Tyr_Ser-Thr
Protein tyrosine and serine/threonine kinase
Representative CDS sequence
>Potri.010G041401.1 pacid=42797259 polypeptide=Potri.010G041401.1.p locus=Potri.010G041401 ID=Potri.010G041401.1.v4.1 annot-version=v4.1
ATGGTTTTCAGGTACTTGGCACCTGAGTATGCATCAAGTGGAAAACTAACAGATAGGTCTGATGTATTTTCATTTGGAGTTGTGCTTCTAGAACTGATAA
CCGGGCGCAAACCAGTCGATGCAAGTCAGCCTTTGGGGGATGAAAGCTTGGTTGAATGGGCTCGTCCACTCCTGATTCATGCCCTTGAAACCGGCGAATT
TGGTGAGTTGGTTGATACACGGCTCGAAAAGCATTACGTGGAGAGTGAATTATTTAGAATGGTTGAGACAGCTGCTGCTTGTGTTCGCCATTCAGCTCCA
AAGCGGCCACGCATGGTGCAGGTGGTGAGAGCATTAGATAGTGAAGGTGAACTCCCAGATCTCAGTAATGGTGTGAGATTTGGTCAGAGCGCTGTATATG
ATTCAGTTCAGTACAGTCAAGACATTAGGAAATTCAGGGGAATGGCTATTGGCAGTAATGGAAGCTCAGAATTTGACACTTCCAGTGGAGATTACTCTGG
TAGACACATATCCCGTGAACCCCCAGCATCAAGTTAG
AA sequence
>Potri.010G041401.1 pacid=42797259 polypeptide=Potri.010G041401.1.p locus=Potri.010G041401 ID=Potri.010G041401.1.v4.1 annot-version=v4.1
MVFRYLAPEYASSGKLTDRSDVFSFGVVLLELITGRKPVDASQPLGDESLVEWARPLLIHALETGEFGELVDTRLEKHYVESELFRMVETAAACVRHSAP
KRPRMVQVVRALDSEGELPDLSNGVRFGQSAVYDSVQYSQDIRKFRGMAIGSNGSSEFDTSSGDYSGRHISREPPASS
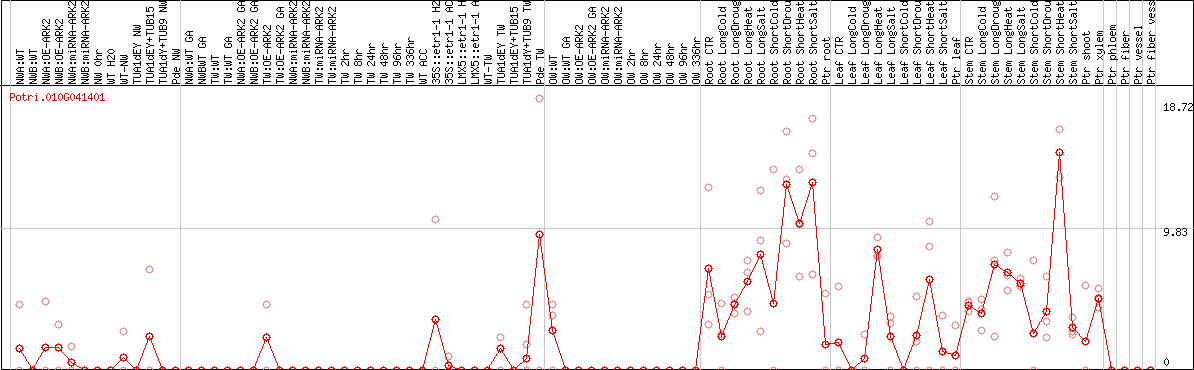
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.010G041401 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.