Potri.010G070200 [POPLAR]

| External link |
|
||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | |||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| ||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info | |||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.010G070200.4 pacid=42797227 polypeptide=Potri.010G070200.4.p locus=Potri.010G070200 ID=Potri.010G070200.4.v4.1 annot-version=v4.1
ATGAAGTCTTCAACTCGGCTTGACTCGGCCATTTTTCAACTCACTCCAACTCGAACCAGGTGTGATTTGATTATATGTGTGAATGGTAAGACAGAGAAAA
TAGCTTCAGGGTTGGCACAACCATTTCTTGACCACTTGAAAACTGCTCAAGATCAATTGGCCAAGGGGGGCTACTCAATTATTCTTGAGCCTGGAACTGA
TGCAGCATGGTTCACGAAAGGAACAATCGAGAGGTTTGTTCGCTTTGTGAGTACACCAGAGGTCTTGGAACGAGTTTACAATTTAGAATCTGAGATCTTG
CAGATCGAAAAGGGAATTGCAATTCAAAGCAACAATGATATAGGATTGAGCTCGGTAGAAGATAATCGAGCAAAACCTGCAGAGTGCATTGAAGGTAGCA
GGCCTCCAACAGATTCTAGTGAGGAAAAAGCCATTGTCCTTTACAAGCCTGGTTCACATCCACCAGAAGCAAATGGGTCGACTGTGCAGGAAGGAAATTC
CAAGGTTCAGCTTCTGAAAGTTCTGGAGACACGAAAAACTTCACTGCAGAAAGAGCAGGGCATGGCCTTTGCACGTGCTGTAGCTGCTGGTTTTGACATT
GATCACATGGCACATTTAATGTCATTTGCTGAAAGTTTTGGAGCCTTACGCTTGATGGATGCTTGTGTAAGATTTATGGAATTATGGAAAAGAAAACATG
AAACTGGCCAGTGGGTTGAAATTGAAGCAGCTGAAGCAATGTCTAGCAGGACAGATTTTTCAGCCATGAATGCGTCAGGCATCGATCTCTCTAATACAAT
CAACAAGCAGTGGCCCGAAACTCCTGATAGTAACAGGAAAGCAGGTGTTGACCCAAATGCAGATGAGAGGCCTCCAACGGATCAACAACCATCACCAGGC
CAACAGGAATATTTTCAAGCCCAGTTTCCGCATCCTATGTTCCCTCCCTGGCCCATTCATTCGCCACCTGGAGCAGTGCCTGTCTTTCCAGGGTATCCAA
TGCAAGGCATTGCTTACTACCAGAATTATCCGGGAAATAACCCAGTTTTTCAGCCACCATATCCATCAGGCGAGGATCCTAGAATCCATGCTGGTCAAAG
GATGAGGCAAAGAAGGCATTCTATGGATAGCAACACTGAAACTGAGGCTTGGGAAGTGGATGCTTTGAGAACAGGATCTCAGGATGAGGCAGAATTGGAG
AAGGAAACTTCGAGAGGTCGAGGACGAGGGAGGAAGGGAAGTCATTCAGGTAAAAAAAAATCAGGCACGGTTGTTATTCGGAACATCAACTACATCACTT
CAAAGAGGCAGGACTCATCTGTTAGTGAATCACGATCAGCTTCTGGTTCTGAAAATGACGAGGAAGATGAAATTTTATCAGACACAGCTCCGAATGTGAA
GCACAGGAACTCTCTGAGATCCTCAAAAAGGAAGGGAAGCCATACAAAATCTACAGATGAACTGAAATTATCAGATATGGCTGGAACATCTTACGGGAAG
GAGGAGGAAGGTGGACACTGGAAAGCATTTCAAAATTATTTGCTGAAAGATGCTGATGAAGCTGAACGTGTTGTTGACCAAGGTATGTTTGCAATGGAAA
AGAATGTTCGAGCAAAAAGGCGACAAAATACAATGGGTGATGATCCTTTAGTTTTTGATGGACGAGATCCAGTTGATAATCAAGAAGGGGATGTGACAGT
TATGCAGAAAATTAGTGGAAACTTGACCCGCATGACTAAGGCATCAAAAGATGAATTGTTGCTATCCATAAAAATGGGTCAGCCTAATGATGATAGAAGG
TTGATAAATGGTCAAATGGATCTACAGTCTGCAGAAATAGATGGGAGGAGAGGTCAATATAGGATGAATGCAAATGATGATTTTATAATTCATGGACGAG
AAAACAAGTCAGGTTATAGAAGTTTAGCATCAGATCCTCTGGCTGTTAATGGATTTGAGACTGCAAAAAATGATCTAGACAGGAGGTCATCCGTCAACAT
GGATGATGATTCATACATAGTCTCACTAAGGTCAACTTCACTAGATCAAGCTGGAACTGAGGGGAGAAACACAATCGACATGGACTCTGAGTTCCCATCA
ACAGTTCAGAGGACAGAGAGCTTATCTAACAGGAGCCAAGTCAAATATGAACCGGATGATTTAAGTTTGATGCCTGAGCGTGGGACAGAAAAGGGGTCTA
TTGGTTATGATCCTGCTTTAGATTATGATATGCAGGCTTCGCTACATAAGAAAAATAATGAGGCTGTAGCTGGTCAAGGCTCTAAGAAGTCAGACAAGGA
CCGGAAATCAAAACTCATTCCAGATACTTCAGATAGGAAGAAGCCAGCAGGACCGATAAGGAAAGGAAAGCCTTCGAAGTTGAGTCCTTTGGATGAAGCA
AAGGCACGTGCTGAGAGGCTAAGAACATTTAAAGCTGATCTCCAGAAAATGAAGAAAGAGAAGGAAGAAGAAGAGATTAAACGACTGGAAGCTTTGAAGT
TAGAGAGACAAAAGAGGATTGCTGCCAGAGGCAGTTCCACTACTGCTCAGTCAGCATCACAGCGAACCAGTAAACAATTGCCAATAAAGCTTTCGCCAGG
CTCTCAAAGAGGATCAAAATTCAGTGATTCAGAGCCGGGATCCTCATCACCTCTACAAAGGTTCTCCATCAAAACTGTTTCTGCAGGATCGGGTGATTCT
CAGAAAGTCTCAAGGTCCAGCAAATTGAGCACTGGAACAACTAGCACAGTTGGAAATAGGTTAACTCAGTCAGTATCATCATTGTCTGAACCAAAGAAAG
ACAACAGTGGTGTGACACCTGATTCAAAGGCATCCGTGGCCAGGATTAGGAGGTTATCAGAGCCAAAAATAAGTAGCCGCGATCATACTTCTTCAACTAA
ACCACGAAACAGTGAATCAGTATCAAAGCCAAAATTATCCAGTGGGGCTGACAGCAAGAAAATATCTGCACTGATGAACCATGATAAAAGCAAGGTGGCA
TCTCTTCCAGAACTGAAAACTAAAACCACTAAAGGACATGATGTTGTGCCTGGCAATTCAGCAGCAAAAGAAATTCCACAGAAGATGAACAAAAGTAAGT
CTATTTCAACTTCTAAAAGCACTGAATTGAAACAGAATGGCAACAAAATTTCACATCATAGTGATGGGGATGACAACCCAATAATTGAGAAAACTGTTGT
GCTTGAGTGTGAGAAGCCCACCATTCCCTCTGTACACGCATCAGAACAGAATATAGAAGTACAAGATGGACACTCTAATAACTATAAAATACCGGAGAAA
ACAGAGACTGTGGTGGATTATGCTAATTTTCAAGCGCCAGTTTCACCATTTACCATGGATGGGATTGATAGAAACCACACTGAGCATCAATTACCAAAGC
ACCCCGGGGTTCATGAGGCTGCATCTGAGCATGCAAGTCATGCAGAGAAAGAATTACCAAAGCTTTCAAGCACTCATATTGCTGAGAAGCCCTACCATGC
CCCTTATGCTCGGGTCTCTTTTATGGAAGATCCTTGTACTGAAAATTCTGAACATGGCAAAGCAACTCCAACAAGCTTACAGACACATTCAGCAGGTGCA
GAGACCATTAAAGCTCATGTATCTGATTTAAAAAGCTTGAAACTAGAACAGATCCCTGAAGTATTGGAGAAGCCTCAGACAAAGGAATCATCTAAAGGGT
TTAGACGATTGCTGAAGTTTGGAAGAAAGAGCCAAACTGCAGGTGAACGTAATGTAGAATTGGATAATGTTAGTTTGAATGGTTCCGAGATGGATGACAA
CGCTGCTTTTTCTAGTGAAGTTCATACATTGAAGAATCTAATCTCTCAAGATGAGACCCCTACTGCTGGCCCCAATCAAAAGACTTCTCGCCATTTCTCA
TTGTTATCGCCCTTCCGAAGCAAGTCAGGTGAAAAGAAGATGACTACTTGA
|
||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.010G070200.4 pacid=42797227 polypeptide=Potri.010G070200.4.p locus=Potri.010G070200 ID=Potri.010G070200.4.v4.1 annot-version=v4.1
MKSSTRLDSAIFQLTPTRTRCDLIICVNGKTEKIASGLAQPFLDHLKTAQDQLAKGGYSIILEPGTDAAWFTKGTIERFVRFVSTPEVLERVYNLESEIL
QIEKGIAIQSNNDIGLSSVEDNRAKPAECIEGSRPPTDSSEEKAIVLYKPGSHPPEANGSTVQEGNSKVQLLKVLETRKTSLQKEQGMAFARAVAAGFDI
DHMAHLMSFAESFGALRLMDACVRFMELWKRKHETGQWVEIEAAEAMSSRTDFSAMNASGIDLSNTINKQWPETPDSNRKAGVDPNADERPPTDQQPSPG
QQEYFQAQFPHPMFPPWPIHSPPGAVPVFPGYPMQGIAYYQNYPGNNPVFQPPYPSGEDPRIHAGQRMRQRRHSMDSNTETEAWEVDALRTGSQDEAELE
KETSRGRGRGRKGSHSGKKKSGTVVIRNINYITSKRQDSSVSESRSASGSENDEEDEILSDTAPNVKHRNSLRSSKRKGSHTKSTDELKLSDMAGTSYGK
EEEGGHWKAFQNYLLKDADEAERVVDQGMFAMEKNVRAKRRQNTMGDDPLVFDGRDPVDNQEGDVTVMQKISGNLTRMTKASKDELLLSIKMGQPNDDRR
LINGQMDLQSAEIDGRRGQYRMNANDDFIIHGRENKSGYRSLASDPLAVNGFETAKNDLDRRSSVNMDDDSYIVSLRSTSLDQAGTEGRNTIDMDSEFPS
TVQRTESLSNRSQVKYEPDDLSLMPERGTEKGSIGYDPALDYDMQASLHKKNNEAVAGQGSKKSDKDRKSKLIPDTSDRKKPAGPIRKGKPSKLSPLDEA
KARAERLRTFKADLQKMKKEKEEEEIKRLEALKLERQKRIAARGSSTTAQSASQRTSKQLPIKLSPGSQRGSKFSDSEPGSSSPLQRFSIKTVSAGSGDS
QKVSRSSKLSTGTTSTVGNRLTQSVSSLSEPKKDNSGVTPDSKASVARIRRLSEPKISSRDHTSSTKPRNSESVSKPKLSSGADSKKISALMNHDKSKVA
SLPELKTKTTKGHDVVPGNSAAKEIPQKMNKSKSISTSKSTELKQNGNKISHHSDGDDNPIIEKTVVLECEKPTIPSVHASEQNIEVQDGHSNNYKIPEK
TETVVDYANFQAPVSPFTMDGIDRNHTEHQLPKHPGVHEAASEHASHAEKELPKLSSTHIAEKPYHAPYARVSFMEDPCTENSEHGKATPTSLQTHSAGA
ETIKAHVSDLKSLKLEQIPEVLEKPQTKESSKGFRRLLKFGRKSQTAGERNVELDNVSLNGSEMDDNAAFSSEVHTLKNLISQDETPTAGPNQKTSRHFS
LLSPFRSKSGEKKMTT
|
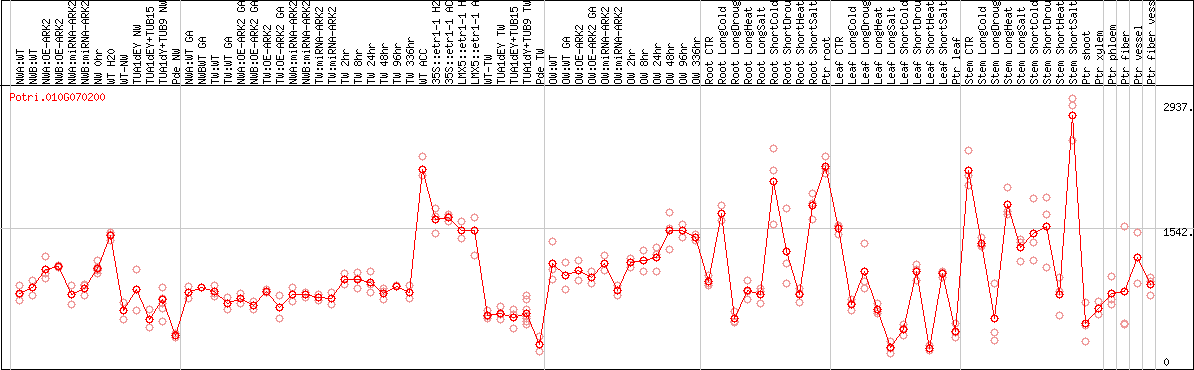
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.010G070200 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
