External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G27520 449 / 8e-160
AtPNC2, PNC2
peroxisomal adenine nucleotide carrier 2 (.1)
AT3G05290 448 / 1e-159
AtPNC1, PNC1
peroxisomal adenine nucleotide carrier 1 (.1)
AT2G39970 96 / 1e-22
PXN, PMP38, APEM3
peroxisomal NAD carrier, peroxisomal membrane protein 38, ABERRANT PEROXISOME MORPHOLOGY 3, Mitochondrial substrate carrier family protein (.1)
AT5G66380 86 / 7e-19
ATFOLT1
folate transporter 1 (.1)
AT5G07320 64 / 2e-11
APC3
ATP/phosphate carrier 3, Mitochondrial substrate carrier family protein (.1)
AT1G34065 61 / 2e-10
SAMC2
S-adenosylmethionine carrier 2 (.1)
AT2G37890 60 / 6e-10
Mitochondrial substrate carrier family protein (.1)
AT4G32400 59 / 1e-09
EMB42, EMB104, ATBT1, SHS1
SODIUM HYPERSENSITIVE 1, EMBRYO DEFECTIVE 42, EMBRYO DEFECTIVE 104, ARABIDOPSIS THALIANA BRITTLE 1, Mitochondrial substrate carrier family protein (.1)
AT5G61810 57 / 4e-09
APC1
ATP/phosphate carrier 1, Mitochondrial substrate carrier family protein (.1.2)
AT1G14560 54 / 4e-08
Mitochondrial substrate carrier family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.005G033800
449 / 3e-160
AT3G05290 433 / 1e-153
peroxisomal adenine nucleotide carrier 1 (.1)
Potri.013G023200
448 / 1e-159
AT3G05290 449 / 5e-160
peroxisomal adenine nucleotide carrier 1 (.1)
Potri.010G192600
106 / 2e-26
AT2G39970 460 / 7e-164
peroxisomal NAD carrier, peroxisomal membrane protein 38, ABERRANT PEROXISOME MORPHOLOGY 3, Mitochondrial substrate carrier family protein (.1)
Potri.002G000200
93 / 4e-21
AT2G39970 389 / 3e-135
peroxisomal NAD carrier, peroxisomal membrane protein 38, ABERRANT PEROXISOME MORPHOLOGY 3, Mitochondrial substrate carrier family protein (.1)
Potri.007G019000
88 / 9e-20
AT5G66380 463 / 8e-166
folate transporter 1 (.1)
Potri.008G064800
86 / 6e-19
AT2G39970 393 / 2e-137
peroxisomal NAD carrier, peroxisomal membrane protein 38, ABERRANT PEROXISOME MORPHOLOGY 3, Mitochondrial substrate carrier family protein (.1)
Potri.002G200900
67 / 2e-12
AT2G47490 471 / 1e-168
NAD+ transporter 1, ARABIDOPSIS THALIANA NAD+ TRANSPORTER 1, NAD+ transporter 1 (.1)
Potri.015G108900
65 / 1e-11
AT5G51050 725 / 0.0
ATP/phosphate carrier 2, Mitochondrial substrate carrier family protein (.1)
Potri.012G110700
65 / 1e-11
AT5G51050 714 / 0.0
ATP/phosphate carrier 2, Mitochondrial substrate carrier family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10010517
471 / 2e-168
AT3G05290 437 / 2e-155
peroxisomal adenine nucleotide carrier 1 (.1)
Lus10034055
443 / 1e-157
AT3G05290 427 / 3e-151
peroxisomal adenine nucleotide carrier 1 (.1)
Lus10015188
422 / 4e-149
AT3G05290 480 / 4e-172
peroxisomal adenine nucleotide carrier 1 (.1)
Lus10031510
421 / 1e-148
AT3G05290 474 / 8e-170
peroxisomal adenine nucleotide carrier 1 (.1)
Lus10014839
109 / 3e-27
AT2G39970 481 / 5e-172
peroxisomal NAD carrier, peroxisomal membrane protein 38, ABERRANT PEROXISOME MORPHOLOGY 3, Mitochondrial substrate carrier family protein (.1)
Lus10009886
106 / 3e-26
AT2G39970 484 / 2e-173
peroxisomal NAD carrier, peroxisomal membrane protein 38, ABERRANT PEROXISOME MORPHOLOGY 3, Mitochondrial substrate carrier family protein (.1)
Lus10028339
79 / 1e-16
AT5G66380 384 / 2e-135
folate transporter 1 (.1)
Lus10001463
72 / 4e-14
AT2G47490 415 / 1e-146
NAD+ transporter 1, ARABIDOPSIS THALIANA NAD+ TRANSPORTER 1, NAD+ transporter 1 (.1)
Lus10039847
66 / 1e-11
AT4G28390 545 / 0.0
ADP/ATP carrier 3 (.1)
Lus10013518
61 / 2e-10
AT5G56450 485 / 8e-174
Mitochondrial substrate carrier family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF00153
Mito_carr
Mitochondrial carrier protein
Representative CDS sequence
>Potri.010G092100.1 pacid=42798884 polypeptide=Potri.010G092100.1.p locus=Potri.010G092100 ID=Potri.010G092100.1.v4.1 annot-version=v4.1
ATGGCCTTCGATTTGGAGTCTTTGTCAGAGGCAACATCCGGGGCAATTGGTGCTCTGGTTAGCACCACTATATCGTACCCACTTGACACCTGCAAGACCA
AATATCAAGCTGAGGTTAGAGCTCATCATCAACAAAAGTATAGGAACATATCTGATGTTTTCTGGGAGGCAATTGCTTCCAGGCAGGTTCTTTCATTGTA
TCAAGGCCTTGGGACGAAGAACTTGCAGTCTTTTATTTCACAATTTGTATATTTCTATGGGTACAGCTTCTTTAAGCGGTTATATTTGGAGAAAAGTAGG
AACAAAACCATTGGAACAAAGGCAAATTTGATAGTGGCTGCTGCTGCTGGGGCTTGCACTGTCATAGTGACACAGCCTCTGGACACAGCATCCTCTAAAA
TGCAAACAAGTGAATTTGGAAAATCCAGAGGACTTTGGAAAACCCTCTCGGAGGGCACTTGGAGTGAGGCATTTGATGGTCTTGGCATATCTCTTCTTCT
GACATCAAACCCATCAATCCAGTACACAGTTTTTGATCAGTTGAAACGAAGACTACTGGAGCGGCAGCTCAGCAAAAGATCAAGCATAGAATCATCTCCA
GAAGCACTCTCTGCCTTTTCTGCTTTTGTATTGGGTGCAGTTTCAAAGTGCATCGCGACGTGTGTCACATACCCAGCTATCAGATGTAAAGTGACGCTTC
AGGCTGCTGAATCAGATGAAAGTGAGATCGAGGAAGTTCAAGCAAAAACCAAGACAATTTCAGGTGCACTCTATTCTATTTGGAAAAATGAAGGTTCGGC
TGGTTTCTTCAAAGGGTTGCTGGCCCAGAACCTGAAGACTGTGCTGAGCTCAGCATTGCACTTGATGATAAAGGAGAAGATATCAAAGACTACATGGTTC
CTGATGCTTGCACTTAAAAGGTATCTGTTTGTCACACGCAGCAGAATCAAGAGCACTTTATGA
AA sequence
>Potri.010G092100.1 pacid=42798884 polypeptide=Potri.010G092100.1.p locus=Potri.010G092100 ID=Potri.010G092100.1.v4.1 annot-version=v4.1
MAFDLESLSEATSGAIGALVSTTISYPLDTCKTKYQAEVRAHHQQKYRNISDVFWEAIASRQVLSLYQGLGTKNLQSFISQFVYFYGYSFFKRLYLEKSR
NKTIGTKANLIVAAAAGACTVIVTQPLDTASSKMQTSEFGKSRGLWKTLSEGTWSEAFDGLGISLLLTSNPSIQYTVFDQLKRRLLERQLSKRSSIESSP
EALSAFSAFVLGAVSKCIATCVTYPAIRCKVTLQAAESDESEIEEVQAKTKTISGALYSIWKNEGSAGFFKGLLAQNLKTVLSSALHLMIKEKISKTTWF
LMLALKRYLFVTRSRIKSTL
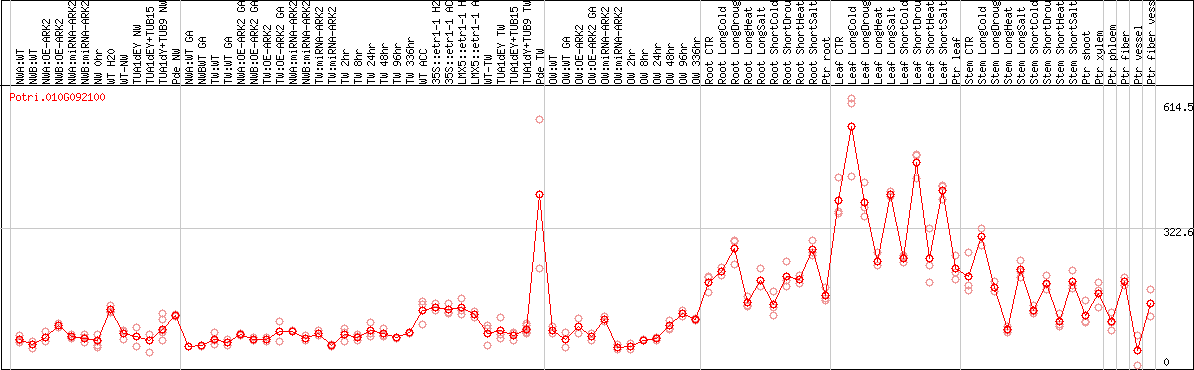
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.010G092100 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.