RDR903 (Potri.010G104900) [POPLAR]

| External link |
|
||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | RDR903 | ||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| ||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| ||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.010G104900.1 pacid=42797881 polypeptide=Potri.010G104900.1.p locus=Potri.010G104900 ID=Potri.010G104900.1.v4.1 annot-version=v4.1
ATGGCAAAGACAATTAGCTTGTATGGTTTTCCTACTGTTGAACCTCCAGATGACATAAAGGAATTTCTGGAAGAATACTCTGGTGAAGGTACAGTTGTAG
CTGTTCATGTTGCACAACCAAAAAGTGCAGGGTCAAGAACAAATGCTATAGTTGAATTCACAACCAGTGAAGCTGCTCAAAATATCAAGCTCAAGTCCTT
AGATGATGAAGGTCTATGGTATGGTAATTCTTATCTGAAAGCGCGGGATGTGTATCGCGATACTGTTCCAATGAGAAAGGCCTTTCAGACTCAGTATAGC
ATGGATAACATAACACTGCACTTCGGATGTCAGGTTTCAAAGGAAAAATTCTCAGTTCTTTGGACTCAAAAAGATGTTTCGGTAAAATTTGGTGTTGAAC
TGAGGAAATTCCATTTCTTTCTGACGCATCTTTCAAAAAATTACAAGCTCGAGCTATCATATGAGAACATTTTTCAAATTGAGCTGCATCGACCTCGTGG
AAAAACTAAGAAATATCTTCTCGTCCAGATGCTCGGAGGCCCCAAGATTTTTAAGAAAGACACGAATAAGCTCAGCAATTTTAAGGAAGCTACTGATGAT
CAGTGGGTTCGAGATGTTGATTTTTCTCCATCTTACTGCATAGGCCAATCTTCTGCTTTGTGCTTGGAGCTTCCGCCCAACAGCCAACTTCCGAATTTTC
AGGAAAATTTTGTCTGTTACAAAGAAGATGAAGGCCACTTCATTCTGGAAAAAGGATCAACCTTCTGTTGCAAGTCAGATCTTGTTCCTATTTTGAGTGC
AGCTCAAGTATCTGAACTTCCATATGACATTGTGTTTAAAGTCAATTCCTTGGTCCAGCATGGTTGTCTTCCTGGACCAGCACTTGATGCAAGGTTCTTT
CGTTTGATCAATCCATCTAAAATTCGTATTGCGCATGCATACATACAACATGCCCTTGAGAAGCTAAGTCATTTGAAGGAATGCTGTTATGATCCTGCAC
GATGGCTCCGGGAGCAATACCAGAAATACCTCACCACTGGCAGACTTCCTACACCACCTGCTATTGCTGTAGATGATGGGTTGGTATTTTTACGTAGGGT
TCAGATAACCCCTACTAAATTGTACTTCTGTGGCCCAGACGTGAATCTCTCCAATCGTGTTTTACGCAAATATCCTGGTGATATTGATAACTTTCTTCGG
GTTTCTTTTGTTGACGAGGATCTTGATAAACTTTTTTCTGCAAATATATCGCCACGGACATTTTCTGCAATTGAGGGGAGGCAAACCAACATTTATCAAA
GGATACTGTCAGTTCAAAGAAATGGCATAACTATCGGATCTAAGAAGTTTGAGTTTCTTGGTTTCTCACAGAGTCAAGTACGGGAGAGTTCTCTTTGGAT
GTTTGCTTCTAGACCTGGGCTCACAGCAGCAGACATTAGAGAATGGATGGGTGATTTTCGTGAAATAAAGAATGTGGCCAAATATGCTGCAAGGCTTGGC
CAGTCTTTCAGCTCATCGAGGGAAAGTTTCAACATTGATAGGCATGAAATAGAAATTATTCCTGATATAGAGGTTAAAAGCGGTGGAGTCAACTATGTAT
TCTCTGATGGTATTGGAAAAATTTCAGCAGAATTGGCAGATAGCATAGCTCGGAAACTTGGCTTTAGAAGTTTTACTCCTTCTGCCTTTCAGATTAGGTA
CGGAGGCTACAAAGGTGTTGTGGCTGTTGATCCTACTTCATCAATGAAATTGTCACTGAGGAGAAGTATGTCCAAGTACAAATCAACTAACACTAGCTTG
GATATTTTGGATTGGAGCAAGTATCGGGCTTGTTTTCTCAATCGTGAAGTGATCATTCTTTTATCTACCCTTGGGGTTAAAGATCAAGTTTTTGAAAGGA
AGCAAAAGGAGGCTATAGCTCAACTAGATGCTATTTTAACCGATCCAATAAAAGCACAGGAGGCACTTGAGTTGATGGCTGGTGGAGAGAACGCACGTGT
TCTGAATGGAATGCTTGCATGCGGTTATAAGCCGGGTGCAGAGCCATTTCTAGCAATGATGCTGGAAACACTCCGCGCATTGAAGTTGCTTGATTTGCGG
ACAAAATCAAGAATATTTGTTCCAAACGGAAGAGCAATGACGGGATGTCTGGATGAAACCAGGACGTTGGAATATGGTCAAGCATTTGTCCAGTATTCTC
GTGCTAGATTTAGGAAGTTACATGATCATTTCAAAGGTGGCAAAACAGATCAACATACTCTGATTTTTAGGGGAAAGATAGTTGTTGCGAAAAACCCCTG
TCTACACCCAGGAGACGTGCAGATCTTAGAGGCTGTGGATGTGCCGGCTTTACATCACATGGTGGATTGCATCGTTTTCCCCCAAAAAGGAAAGAGACCT
CATACAAATGAATGCTCGGGAAGTGATCTAGATGGAGATGTGTACTTTGTATGTTGGGATTCGGATCTCATTCCTCCTCAGAAATTTCCACCCATGGATT
ACGCTGCACCACAAACTACGATTTTGGACCACGATGTTACAATAGAGGAAGTTCAGGAGTATTTTACAGATTACCTACTCAACGACAACTTAGGGATCAT
TTGCAGTGCTCACGTGGTCTTTGCAGACAGTGAGCCTGACATGGCAAGAAGTGAAAAATGTATTGAGCTTGCTCAGCTATCCTCAATCGCTGTTGACTTC
CCAAAAACCGGTGTGCCTGCTAAAATTCCTAAAGAACTACGCGTCAAAGAATATCCAGATTTTATGGAGAAGGCTGCATATAAACATACTACCTATGTAT
CTCAGCGTGTACTTGGAAAGCTTTTTCGAGCTGTAAGAGACATCGCACCGGACACAAGTCCTGTAAGACCCTTCACTAAGGAAGCGGCAAAGCGGTCTTA
TGACCCTGACATGGAAGTTGATGGATTTAAGGATTATATCAACGAGGCTTTCTATTACAAAAGTGAATACGACAACAAACTGGGGAACATGATGGACTAT
TACGGGATCAAGACTGAGGCTGAAATAATCGGTGGTTGCATCATGAGAATGGGAAGATCTTTTGATAAGAAAAAGGACTTGGAAGGCATTAATTTTTCTG
TAAGTTCCTTGAGGAAGCAAGCTAGAGCCTGGTTCAACGAGAGCGGCTCAAAGGAAAGCCCTGATGATGTTTATCCAAAGGCATCAGCTTGGTACCACGT
CACATATCATCCTCGTTTTTGGGGCCTCTACAACGAGGGAATGGATCGAGTTCATTTCCTCAGTTTTCCCTGGTGTGTTCATGACAAACTTTTTGAGATT
AAGAAGGGCACGACATCACTGGTGCACCAGCTCAAACGAATGGCGAGCTTTCATGTAGAGATCGGAGACTAG
|
||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.010G104900.1 pacid=42797881 polypeptide=Potri.010G104900.1.p locus=Potri.010G104900 ID=Potri.010G104900.1.v4.1 annot-version=v4.1
MAKTISLYGFPTVEPPDDIKEFLEEYSGEGTVVAVHVAQPKSAGSRTNAIVEFTTSEAAQNIKLKSLDDEGLWYGNSYLKARDVYRDTVPMRKAFQTQYS
MDNITLHFGCQVSKEKFSVLWTQKDVSVKFGVELRKFHFFLTHLSKNYKLELSYENIFQIELHRPRGKTKKYLLVQMLGGPKIFKKDTNKLSNFKEATDD
QWVRDVDFSPSYCIGQSSALCLELPPNSQLPNFQENFVCYKEDEGHFILEKGSTFCCKSDLVPILSAAQVSELPYDIVFKVNSLVQHGCLPGPALDARFF
RLINPSKIRIAHAYIQHALEKLSHLKECCYDPARWLREQYQKYLTTGRLPTPPAIAVDDGLVFLRRVQITPTKLYFCGPDVNLSNRVLRKYPGDIDNFLR
VSFVDEDLDKLFSANISPRTFSAIEGRQTNIYQRILSVQRNGITIGSKKFEFLGFSQSQVRESSLWMFASRPGLTAADIREWMGDFREIKNVAKYAARLG
QSFSSSRESFNIDRHEIEIIPDIEVKSGGVNYVFSDGIGKISAELADSIARKLGFRSFTPSAFQIRYGGYKGVVAVDPTSSMKLSLRRSMSKYKSTNTSL
DILDWSKYRACFLNREVIILLSTLGVKDQVFERKQKEAIAQLDAILTDPIKAQEALELMAGGENARVLNGMLACGYKPGAEPFLAMMLETLRALKLLDLR
TKSRIFVPNGRAMTGCLDETRTLEYGQAFVQYSRARFRKLHDHFKGGKTDQHTLIFRGKIVVAKNPCLHPGDVQILEAVDVPALHHMVDCIVFPQKGKRP
HTNECSGSDLDGDVYFVCWDSDLIPPQKFPPMDYAAPQTTILDHDVTIEEVQEYFTDYLLNDNLGIICSAHVVFADSEPDMARSEKCIELAQLSSIAVDF
PKTGVPAKIPKELRVKEYPDFMEKAAYKHTTYVSQRVLGKLFRAVRDIAPDTSPVRPFTKEAAKRSYDPDMEVDGFKDYINEAFYYKSEYDNKLGNMMDY
YGIKTEAEIIGGCIMRMGRSFDKKKDLEGINFSVSSLRKQARAWFNESGSKESPDDVYPKASAWYHVTYHPRFWGLYNEGMDRVHFLSFPWCVHDKLFEI
KKGTTSLVHQLKRMASFHVEIGD
|
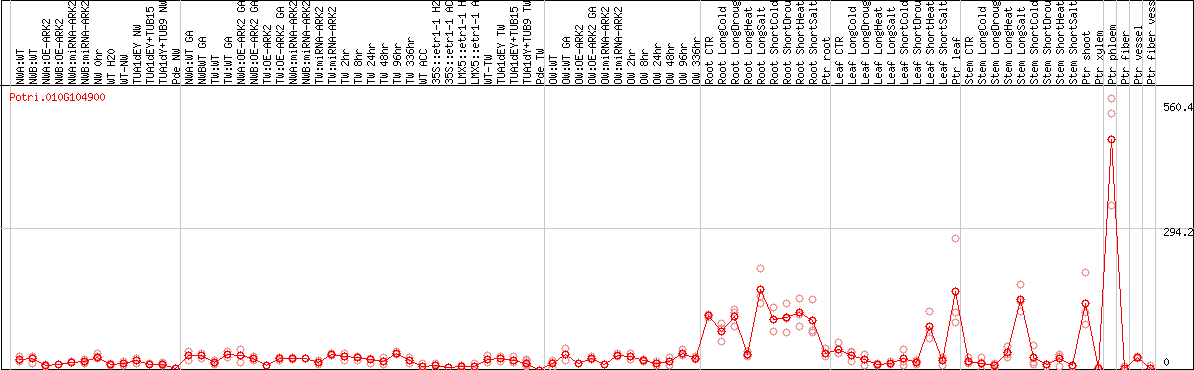
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.010G104900 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
