External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G23260 269 / 5e-94
MMZ1 ,UEV1A
UBIQUITIN E2 VARIANT 1A, MMS ZWEI homologue 1 (.1)
AT1G70660 263 / 2e-91
MMZ2 ,UEV1B
UBIQUITIN E2 VARIANT 1B, MMS ZWEI homologue 2 (.1.2)
AT1G70650 255 / 5e-83
Ran BP2/NZF zinc finger-like superfamily protein (.1.2)
AT3G52560 233 / 5e-80
MMZ4 ,UEV1D ,UEV1D-4
MMS2 ZWEI HOMOLOGUE 4, ubiquitin E2 variant 1D-4 (.1.2.3.4)
AT2G36060 227 / 1e-77
MMZ3 ,UEV1C
UBIQUITIN E2 VARIANT 1C, MMS ZWEI homologue 3 (.1.2.3)
AT3G46460 55 / 7e-10
UBC13
ubiquitin-conjugating enzyme 13 (.1)
AT5G59300 52 / 1e-08
ATUBC7, UBC7
ARABIDOPSIS THALIANA UBIQUITIN CARRIER PROTEIN 7, ubiquitin carrier protein 7 (.1)
AT1G50490 48 / 3e-07
UBC20
ubiquitin-conjugating enzyme 20 (.1)
AT5G25760 46 / 1e-06
PEX4, UBC21
ubiquitin-conjugating enzyme 21, peroxin4 (.1.2)
AT3G20060 45 / 1e-06
UBC19
ubiquitin-conjugating enzyme19 (.1.2)
Paralogs
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10021797
275 / 6e-96
AT1G23260 268 / 3e-93
UBIQUITIN E2 VARIANT 1A, MMS ZWEI homologue 1 (.1)
Lus10034611
273 / 2e-95
AT1G23260 266 / 1e-92
UBIQUITIN E2 VARIANT 1A, MMS ZWEI homologue 1 (.1)
Lus10029415
233 / 5e-80
AT3G52560 290 / 7e-103
MMS2 ZWEI HOMOLOGUE 4, ubiquitin E2 variant 1D-4 (.1.2.3.4)
Lus10004211
223 / 6e-76
AT2G36060 273 / 8e-96
UBIQUITIN E2 VARIANT 1C, MMS ZWEI homologue 3 (.1.2.3)
Lus10016573
54 / 3e-09
AT5G59300 291 / 6e-102
ARABIDOPSIS THALIANA UBIQUITIN CARRIER PROTEIN 7, ubiquitin carrier protein 7 (.1)
Lus10040844
53 / 4e-09
AT5G59300 290 / 1e-101
ARABIDOPSIS THALIANA UBIQUITIN CARRIER PROTEIN 7, ubiquitin carrier protein 7 (.1)
Lus10005014
47 / 1e-06
AT3G20060 292 / 1e-102
ubiquitin-conjugating enzyme19 (.1.2)
Lus10019034
47 / 1e-06
AT3G20060 292 / 1e-102
ubiquitin-conjugating enzyme19 (.1.2)
Lus10033611
45 / 4e-06
AT1G16890 311 / 5e-111
UBIQUITIN CONJUGATING ENZYME 13B, ubiquitin-conjugating enzyme 36 (.1.2.3)
Lus10017654
45 / 4e-06
AT1G16890 311 / 5e-111
UBIQUITIN CONJUGATING ENZYME 13B, ubiquitin-conjugating enzyme 36 (.1.2.3)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0208
UBC
PF00179
UQ_con
Ubiquitin-conjugating enzyme
Representative CDS sequence
>Potri.010G107700.2 pacid=42798886 polypeptide=Potri.010G107700.2.p locus=Potri.010G107700 ID=Potri.010G107700.2.v4.1 annot-version=v4.1
ATGGGTTCTGAAGGTTCAAGCGTTGTTGTGCCGAGGAACTTCAGACTACTGGAGGAGCTTGAGAGAGGGGAAAAGGGGATCGGAGATGGAACTGTCAGTT
ATGGAATGGACGACGCTGATGATATCTTCATGCAGTCATGGACAGGAACTATAATTGGTCCCCCCAATACTGTTCACGAGGGGCGTATCTACCAGTTAAA
ATTGTTTTGTGGCAAGGAATATCCCGATAATCCACCAAGTGTCAGGTTCCAAACTCGGATAAACATGACCTGCGTCAATCCTGAAAGTGGAGTGGTCGAG
CCCAGTCTTTTCCCTATGCTTGCTAATTGGCAGAGGGAGCATACAATGGAGGATATATTAACTCAGTTGAAGAAAGAAATGATGACTTCACAGAACAGGA
AGCTCGCTCAGCCTCCTGAAGGAAACGAGGAGGCAAGGTTGGATCAAAAGGGGCTAGTTCTTAAGTGTTGTATTCTCTAA
AA sequence
>Potri.010G107700.2 pacid=42798886 polypeptide=Potri.010G107700.2.p locus=Potri.010G107700 ID=Potri.010G107700.2.v4.1 annot-version=v4.1
MGSEGSSVVVPRNFRLLEELERGEKGIGDGTVSYGMDDADDIFMQSWTGTIIGPPNTVHEGRIYQLKLFCGKEYPDNPPSVRFQTRINMTCVNPESGVVE
PSLFPMLANWQREHTMEDILTQLKKEMMTSQNRKLAQPPEGNEEARLDQKGLVLKCCIL
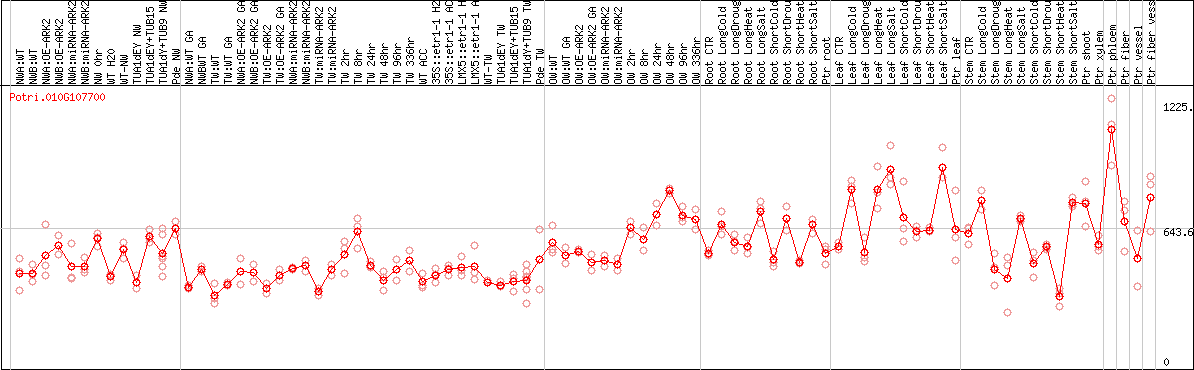
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.010G107700 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.