External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G01200 203 / 9e-67
AUX_IAA
IAA32, MEE10
maternal effect embryo arrest 10, indole-3-acetic acid inducible 32 (.1.2)
AT1G15050 174 / 2e-55
AUX_IAA
IAA34
indole-3-acetic acid inducible 34 (.1)
AT2G33310 99 / 2e-25
AUX_IAA
IAA13
auxin-induced protein 13 (.1.2.3)
AT1G04550 92 / 6e-23
AUX_IAA
BDL, IAA12
indole-3-acetic acid inducible 12, BODENLOS, AUX/IAA transcriptional regulator family protein (.1.2)
AT1G51950 75 / 2e-16
AUX_IAA
IAA18
indole-3-acetic acid inducible 18 (.1)
AT4G28640 74 / 5e-16
AUX_IAA
IAA11
indole-3-acetic acid inducible 11 (.1.2.3)
AT1G52830 72 / 1e-15
AUX_IAA
SHY1, IAA6
SHORT HYPOCOTYL 1, indole-3-acetic acid 6 (.1)
AT3G16500 72 / 2e-15
AUX_IAA
IAA26, PAP1
indole-3-acetic acid inducible 26, phytochrome-associated protein 1 (.1)
AT3G62100 70 / 3e-15
AUX_IAA
IAA30
indole-3-acetic acid inducible 30 (.1)
AT2G46990 68 / 2e-14
AUX_IAA
IAA20
indole-3-acetic acid inducible 20 (.1)
Paralogs
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10014464
92 / 4e-22
AT2G33310 231 / 4e-75
auxin-induced protein 13 (.1.2.3)
Lus10023719
89 / 4e-21
AT2G33310 223 / 4e-72
auxin-induced protein 13 (.1.2.3)
Lus10034962
86 / 4e-20
AT4G29080 289 / 4e-97
indole-3-acetic acid inducible 27, phytochrome-associated protein 2 (.1)
Lus10022868
82 / 8e-19
AT4G28640 198 / 1e-62
indole-3-acetic acid inducible 11 (.1.2.3)
Lus10012984
81 / 3e-18
AT4G29080 280 / 8e-94
indole-3-acetic acid inducible 27, phytochrome-associated protein 2 (.1)
Lus10028222
74 / 6e-16
AT5G65670 322 / 3e-110
indole-3-acetic acid inducible 9 (.1.2)
Lus10039488
72 / 8e-16
AT5G43700 247 / 3e-84
indole-3-acetic acid inducible 4, AUXIN INDUCIBLE 2-11, AUX/IAA transcriptional regulator family protein (.1)
Lus10039413
72 / 1e-15
AT1G04240 250 / 2e-85
SHORT HYPOCOTYL 2, indole-3-acetic acid inducible 3, AUX/IAA transcriptional regulator family protein (.1)
Lus10042929
74 / 2e-15
AT5G65670 356 / 3e-122
indole-3-acetic acid inducible 9 (.1.2)
Lus10019241
72 / 5e-15
AT5G65670 356 / 4e-123
indole-3-acetic acid inducible 9 (.1.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0072
Ubiquitin
PF02309
AUX_IAA
AUX/IAA family
Representative CDS sequence
>Potri.010G119900.1 pacid=42797020 polypeptide=Potri.010G119900.1.p locus=Potri.010G119900 ID=Potri.010G119900.1.v4.1 annot-version=v4.1
ATGGATTCTAATGCATCAGGCTTCCTTCTAAACCCTCCAGCTCTTCACTCTACTTACTATCAGCCTAGGGAAGATGATGGTATTATTGATCTGGGTCTCA
GTCTCAGAACTTTGAAGCCTGAGGCTTATCATCCATCTGGACATCTGGTAGGCCTGGAGGGATATGGTGACCTGATGGACTGGCCTCGGGCAAACTCACC
ACTAAAACACTCCACTTCAAGCTACATTAGATTTACCCCACAAGATTGTGACGAGGAGGCAGAGGGAGTTCAGAGCAAAGACAGGTGGGCTTATGTGAAG
GTAAACATGGATGGGGTGATTGTTGGTAGGAAAATATGCATGCTAGATCATGGTGGTTACTCAAGCCTCGCACTTCAGCTCGAAGACATGTTTGGAAGAC
AATCTGCATCTGGGTTAAGGCTGTTCCAAGCAGGTTCTGAATTCTGCCTGTTCTATAAGGACAGAGAGGAGAATTGGAGAACCGTTGGTGATGTTCCTTG
GAAGGAGTTTGTAGAAAGCGTCAAGCGGCTGAGGATTGCACGAAAAAGCGAACCTCTTCTTCCATATTCACCGGCCTTCTCCTGA
AA sequence
>Potri.010G119900.1 pacid=42797020 polypeptide=Potri.010G119900.1.p locus=Potri.010G119900 ID=Potri.010G119900.1.v4.1 annot-version=v4.1
MDSNASGFLLNPPALHSTYYQPREDDGIIDLGLSLRTLKPEAYHPSGHLVGLEGYGDLMDWPRANSPLKHSTSSYIRFTPQDCDEEAEGVQSKDRWAYVK
VNMDGVIVGRKICMLDHGGYSSLALQLEDMFGRQSASGLRLFQAGSEFCLFYKDREENWRTVGDVPWKEFVESVKRLRIARKSEPLLPYSPAFS
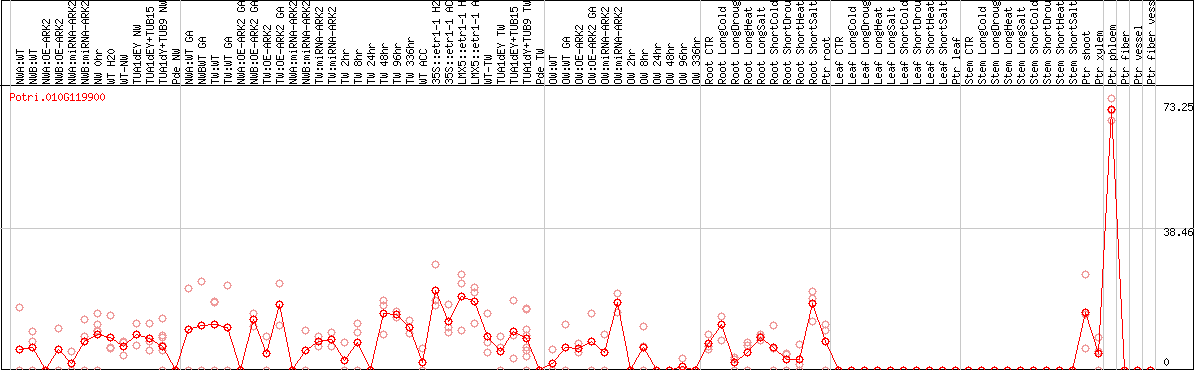
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.010G119900 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.