External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G68640 84 / 4e-19
bZIP
PAN
PERIANTHIA, bZIP transcription factor family protein (.1)
AT3G12250 58 / 3e-10
bZIP
BZIP45, TGA6
TGACG motif-binding factor 6 (.1.2.3.4.5)
AT5G06960 46 / 5e-06
bZIP
TGA5, OBF5
TGACG MOTIF-BINDING FACTOR 5, OCS-element binding factor 5 (.1.2)
AT5G06950 46 / 6e-06
bZIP
TGA2, AHBP-1B
bZIP transcription factor family protein (.1.2.3.4)
AT5G06839 45 / 1e-05
bZIP
TGA10, bZIP65
TGACG \(TGA\) motif-binding protein 10, bZIP transcription factor family protein (.1.2.3)
AT1G08320 43 / 4e-05
bZIP
TGA9, bZIP21
TGACG \(TGA\) motif-binding protein 9, bZIP transcription factor family protein (.1.2.3)
AT5G65210 43 / 4e-05
bZIP
TGA1
bZIP transcription factor family protein (.1.2.3.4.5.6)
AT5G10030 42 / 9e-05
bZIP
OBF4, TGA4
OCS ELEMENT BINDING FACTOR 4, TGACG motif-binding factor 4 (.1.2)
Paralogs
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10041475
81 / 6e-18
AT1G68640 444 / 4e-153
PERIANTHIA, bZIP transcription factor family protein (.1)
Lus10034296
69 / 7e-14
AT1G68640 427 / 1e-146
PERIANTHIA, bZIP transcription factor family protein (.1)
Lus10024140
53 / 3e-08
AT3G12250 532 / 0.0
TGACG motif-binding factor 6 (.1.2.3.4.5)
Lus10026166
49 / 5e-07
AT3G12250 514 / 0.0
TGACG motif-binding factor 6 (.1.2.3.4.5)
Lus10008658
47 / 3e-06
AT3G12250 521 / 0.0
TGACG motif-binding factor 6 (.1.2.3.4.5)
Lus10020610
46 / 9e-06
AT1G08320 633 / 0.0
TGACG \(TGA\) motif-binding protein 9, bZIP transcription factor family protein (.1.2.3)
Lus10037550
44 / 2e-05
AT5G65210 444 / 3e-155
bZIP transcription factor family protein (.1.2.3.4.5.6)
Lus10004876
44 / 2e-05
AT1G08320 635 / 0.0
TGACG \(TGA\) motif-binding protein 9, bZIP transcription factor family protein (.1.2.3)
Lus10011445
44 / 3e-05
AT5G65210 449 / 2e-158
bZIP transcription factor family protein (.1.2.3.4.5.6)
Lus10023822
43 / 7e-05
AT5G06839 491 / 2e-171
TGACG \(TGA\) motif-binding protein 10, bZIP transcription factor family protein (.1.2.3)
PFAM info
Representative CDS sequence
>Potri.010G128001.5 pacid=42797534 polypeptide=Potri.010G128001.5.p locus=Potri.010G128001 ID=Potri.010G128001.5.v4.1 annot-version=v4.1
ATGTTCTGTCTAAAATCAGGCAATGTCACCGTTGTGTCCCATAACTTACCTTATGCAACTGCCTTGAACACGAGCATTGGGTCAGCAGAGATAGCAACAA
CTGGAGCAGGGTGTCTGGACACAGGGCAATACGTGTACCAGAAAGGGACTGGGTTTGATTCCTCGTTGGGAAACGGGCAGAGTTTTGAGAATTGGGGAGA
CTCAGGGATGGCAGATAATAGCCTACAGACGGACACCTCAACAGATGTTAATACTGATGATAAAAACCAGCTTCGTGGAGTTCCGCATGGTGCAGTCATG
GTTGTGAATTCTATGGATCAATCCAAGGGAAGAACTAGTGACCAAAAGACGCAGACCCTTCGAAGGCTGGCTCAGAATCGTGAAGCTGCAAGGAGGGGTC
GACTTAGGAAGAAAGTAATGTCTTTGCTAGTTATGTTATATTGTTTCCTCACGCTGATGATTGGGATTCTTAAAATCTTTTTTTTTTTTTTTTTTTGGAA
TCTTGTTTTTCCAAGTCTTACATTCTCTTTATCCTTCAGTTTTGCCCTTGAATTCCCCTACTGTAGGAAATAA
AA sequence
>Potri.010G128001.5 pacid=42797534 polypeptide=Potri.010G128001.5.p locus=Potri.010G128001 ID=Potri.010G128001.5.v4.1 annot-version=v4.1
MFCLKSGNVTVVSHNLPYATALNTSIGSAEIATTGAGCLDTGQYVYQKGTGFDSSLGNGQSFENWGDSGMADNSLQTDTSTDVNTDDKNQLRGVPHGAVM
VVNSMDQSKGRTSDQKTQTLRRLAQNREAARRGRLRKKVMSLLVMLYCFLTLMIGILKIFFFFFFWNLVFPSLTFSLSFSFALEFPYCRK
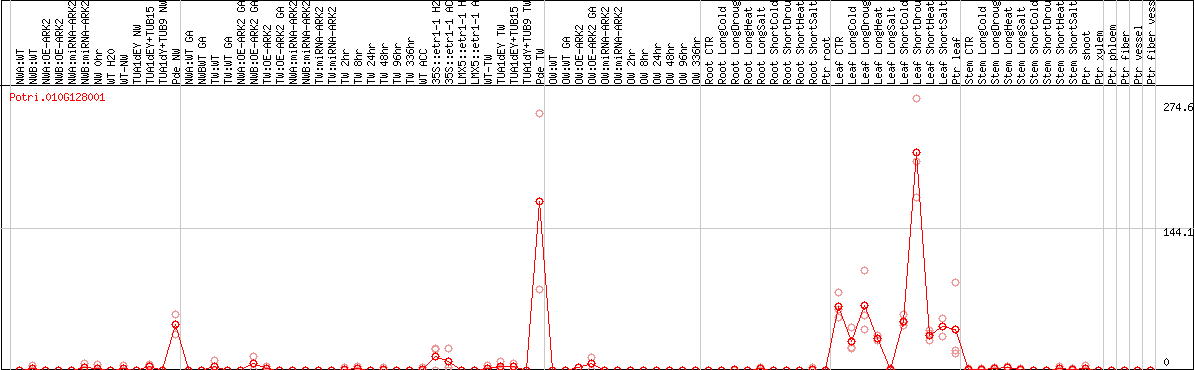
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.010G128001 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.