External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G25520 334 / 1e-117
Uncharacterized protein family (UPF0016) (.1)
AT1G68650 330 / 6e-116
Uncharacterized protein family (UPF0016) (.1)
AT5G36290 156 / 1e-46
Uncharacterized protein family (UPF0016) (.1), Uncharacterized protein family (UPF0016) (.2)
AT4G13590 129 / 2e-35
Uncharacterized protein family (UPF0016) (.1), Uncharacterized protein family (UPF0016) (.2)
AT1G64150 95 / 1e-22
Uncharacterized protein family (UPF0016) (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.008G117900
391 / 5e-140
AT1G25520 344 / 2e-121
Uncharacterized protein family (UPF0016) (.1)
Potri.019G116100
391 / 6e-140
AT1G25520 344 / 2e-121
Uncharacterized protein family (UPF0016) (.1)
Potri.013G086700
160 / 2e-48
AT5G36290 322 / 8e-111
Uncharacterized protein family (UPF0016) (.1), Uncharacterized protein family (UPF0016) (.2)
Potri.019G053400
159 / 5e-48
AT5G36290 328 / 4e-113
Uncharacterized protein family (UPF0016) (.1), Uncharacterized protein family (UPF0016) (.2)
Potri.003G174600
124 / 7e-34
AT4G13590 369 / 2e-127
Uncharacterized protein family (UPF0016) (.1), Uncharacterized protein family (UPF0016) (.2)
Potri.001G053600
124 / 1e-33
AT4G13590 417 / 5e-146
Uncharacterized protein family (UPF0016) (.1), Uncharacterized protein family (UPF0016) (.2)
Potri.003G134300
103 / 6e-26
AT1G64150 344 / 6e-117
Uncharacterized protein family (UPF0016) (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10034293
363 / 5e-129
AT1G25520 349 / 2e-123
Uncharacterized protein family (UPF0016) (.1)
Lus10035040
355 / 8e-123
AT1G25520 339 / 4e-116
Uncharacterized protein family (UPF0016) (.1)
Lus10041478
343 / 4e-121
AT1G25520 327 / 1e-114
Uncharacterized protein family (UPF0016) (.1)
Lus10026995
164 / 8e-50
AT5G36290 401 / 5e-142
Uncharacterized protein family (UPF0016) (.1), Uncharacterized protein family (UPF0016) (.2)
Lus10020195
141 / 5e-42
AT5G36290 321 / 5e-112
Uncharacterized protein family (UPF0016) (.1), Uncharacterized protein family (UPF0016) (.2)
Lus10023685
127 / 5e-35
AT4G13590 443 / 1e-156
Uncharacterized protein family (UPF0016) (.1), Uncharacterized protein family (UPF0016) (.2)
Lus10011763
125 / 3e-34
AT4G13590 429 / 6e-151
Uncharacterized protein family (UPF0016) (.1), Uncharacterized protein family (UPF0016) (.2)
Lus10021697
100 / 2e-25
AT1G25520 94 / 5e-23
Uncharacterized protein family (UPF0016) (.1)
Lus10032330
91 / 4e-21
AT1G64150 333 / 8e-113
Uncharacterized protein family (UPF0016) (.1)
Lus10024713
88 / 4e-20
AT1G64150 322 / 5e-108
Uncharacterized protein family (UPF0016) (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0292
LysE
PF01169
UPF0016
Uncharacterized protein family UPF0016
Representative CDS sequence
>Potri.010G128400.2 pacid=42798457 polypeptide=Potri.010G128400.2.p locus=Potri.010G128400 ID=Potri.010G128400.2.v4.1 annot-version=v4.1
ATGACCTCGGTTGCTCAGGGTTTCACTAAGTCCCTTGCAATGACCGTGCTATCGGAGATCGGCGACAAGACTTTTTTTGCTGCTGCGATCTTGGCCATGC
GTCACCCAAGAAGACTTGTTTTGTCAGGTTGCTTGGCAGCTTTGATTGTGATGACTATCCTTTCTGCTATCGTTGGCTGGGCTGCTCCAAATCTGATCTC
TCGTACTTGGACTCATCACATAACAACAATACTGTTTTTTGGATTTGGGTTTTGGTCATTGTGGGATGGATTTAATGACAAAGGGGAGGCTGAAGAATTG
GCAGAAGTTGAAGCTAAATTGGATGCTGATTGGAAAGCTAACACAGGAACCACCAAAGGTGGTAGCAAGGACGATGATGAGTTGAAAAAGCGGAGGCGGC
CATTTCTTTCACAGTTGTTCTCGCCAATACTCTTGAAAGCATTTTCCATTACATTCTTTGGTGAATGGGGTGACAAGAGCCAGATAGCCACCATTGGCTT
GGCTGCTGATGAGAACCCATTAGGTGTAGTTCTTGGTGGAATTGTAGGGCAAGCATTGTGTACTACTGCCGCTGTTTTTGGTGGAAAGAGCTTGGCATCT
CAAATATCTGAGAAAATAGTTGCATTGTCAGGTGGAGTTCTTTTTATTATTTTTGGAATCCAATCCTTCCTATCAACGGTTGAGTAG
AA sequence
>Potri.010G128400.2 pacid=42798457 polypeptide=Potri.010G128400.2.p locus=Potri.010G128400 ID=Potri.010G128400.2.v4.1 annot-version=v4.1
MTSVAQGFTKSLAMTVLSEIGDKTFFAAAILAMRHPRRLVLSGCLAALIVMTILSAIVGWAAPNLISRTWTHHITTILFFGFGFWSLWDGFNDKGEAEEL
AEVEAKLDADWKANTGTTKGGSKDDDELKKRRRPFLSQLFSPILLKAFSITFFGEWGDKSQIATIGLAADENPLGVVLGGIVGQALCTTAAVFGGKSLAS
QISEKIVALSGGVLFIIFGIQSFLSTVE
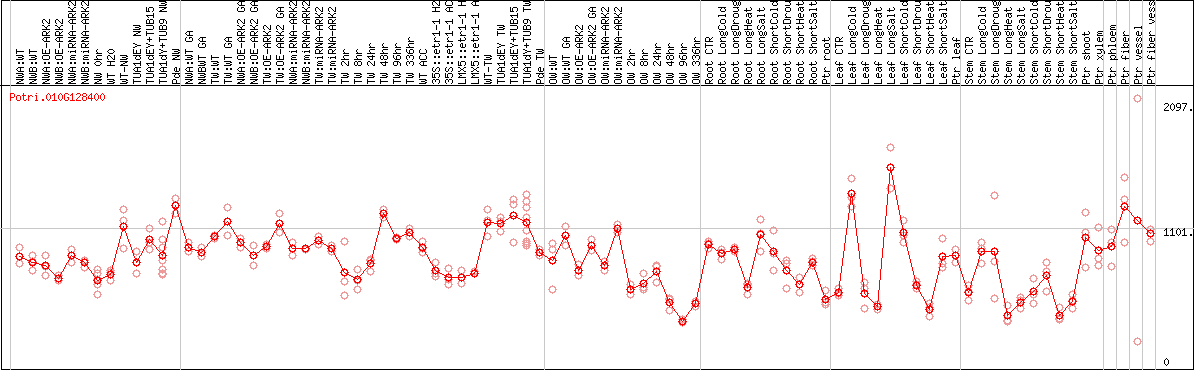
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.010G128400 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.