Potri.010G131800 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.010G131800.1 pacid=42797723 polypeptide=Potri.010G131800.1.p locus=Potri.010G131800 ID=Potri.010G131800.1.v4.1 annot-version=v4.1
ATGACGGACATAACAGACGATATAGCTGAGGAAATCTCCTTCCAAGGATTTGATGATTACTGTAAACTGCTTAAAAATTTATTAAACGATGTGTTGCAGA
GAGAAGTTGGTACTGAATTCGTCGACAAACTTGAACGCAACCTCACTCTCGCTCAGAGTGCTTGTAATTTGAGATTGGCGGGAATAGAAGACACTGCCGA
GTTGCTGGAGAAGCAGCTCGCGTCAGAGATATCAAAAATGACATTAGAGGAAGCATTGACACTTGCTCGTGCTTTTAGCCATTATCTTAATTTGATGGGA
ATTGCTGAGACCCATCACAGGGCACGTAAGACAAGGAATTTGGCGAATCTATCAAAATCTTGTGATGAAGTCTTTAATCAGCTGTTGCACGGTGGAAAAT
CGGGTGATGAACTTTATGCCTCTGTTTGCATGCAGGAGGTTGAAATTGTTCTTACTGCACATCCCACTCAAATTAATCGCCGTACTTTGCAATACAAGCA
TGTTAGAATTGCTCATCTTTTAGAATACAACGATCGACCTGATCTTACACAGGAAGATAGAGAAATCCTTATTGAAGATTTGGTAAGAGAAATTACTTCA
ATATGGCAGACAGATGAGCTTAGGCGCCACAAACCTACACCAGTAGATGAAGCCAGGGCTGGCTTGCACATTGTGGAGCAGTCTCTATGGAAAGCAGTAC
CTCATTTTTTACGACGTGTCAGTAATGCACTAAAGAAGCATACAGGAAAGCCACTTCCATTAACTTGCACGCCAATAAAATTTGGGTCTTGGATGGGGGG
TGATAGAGATGGAAATCCAAATGTAACAGCGAAGGTAACACGAGATGTCTCTCTCTTATCAAGGTGGATGGCTATTGACCTTTACATTAGAGAAGTTGAT
AGTCTCAGATTTGAACTGTCCATGACTCGCTGCAGTGATAAGTTGTCAAGAGAGGCTCATGAAATTCTAGAGCGAGAAACTTCACCAGAGGATCGGCATG
AGAGTTGGAATCAACCCACAAGCAGAAATCAAACAAAGCTTCATCAGCATGCTCCACCACTTCCCACACAACTTCCAGCTAGAGCTGATCTTCCTGCCTG
CACTGAATGCGGTGATGACGGTGGTTCACATCCCAAACTAGAACTTCCAGGGACTGATTACATGCCATTGAGTCGTCAGGATGTTCAGGGTTCTTCAAAT
TCAGAATCTTCATTCCACAAGTCTGGTCATGGTTCTAGCAAATCAATTGCTAATGGAAGCATTGCTAATTCTAACGGCCATCAATCAGCTCCAAGTCCAC
GTGGATCTTTCACCTCCAGTCAACTCCTTGCTCAGAGAAAATGTTTTGCAGAATCTAAGATAGGAAGATCCAGCTTCCAGAAGCTTCTAGAGCCAAGTCC
ACCTGAACGTCCTGGAATCGCTCCTTATAGAATTGTTCTTGGCCATGTGAAAGATAAGCTTATGAAGGCTAGAAGACGTCTGGAACTTCTTCTTGAGGAT
CTTCCCTGTGAACATGAGCCTTGGGATTACTATGAGACAACAGACCAATTGTTGGAACCACTGCTTCTATGCTACGAGTCTCTGCAATCTTGTGGAGCTG
GGGTGCTTGCAGATGGCCGGCTGGTTGACCTGATAAGAAGAGTTGCGACTTTTGGAATGGTATTAATGAAGCTTGACTTGCGTCAGGAATCTGGTAGACA
TTCTGAAGCACTAGATGCAATCACCAAATATTTGGATATGGGAACATATAGTGAGTGGGATGAAGAAAAGAAACTGGAATTCCTAACAAGAGAGCTCAAA
AGCAAGAGGCCACTAGTTCCTCCCACCATTCAGGTTGCTCCTGATGTTAAAGAAGTCTTGGATACATTCCGTGTTGCTGCTGAGCTAGGGAGTGATTCAC
TTGGAGCATATGTGATCTCTATGGCATCCAATGCGAGCGATGTCCTTGCTGTGGAGCTTTTACAAAAAGATGCCCGTCTTGCTGTTAGTGGCGAGCTAGG
GAGACCATGTCCTAGAGGAACGCTGCGGGTGGTTCCATTGTTTGAAACTGTGAAAGATCTTAGAGGGGCTGGTTCGGTTATCAGGAAATTATTATCAATT
GATTGGTACTCAGAGCACATTGTAAAGAACCATAACGGGCATCAGGAGGTGATGGTTGGATATTCTGATTCTGGTAAAGATGCTGGGCGGTTCACTGCTG
CATGGGAACTTTACAAAGCCCAAGAGGATGTTGCGGCAGCTTGCAAAGATCATAAAGTTAAAGTAACTCTATTCCATGGGCGAGGAGGGAGTATCGGTCG
TGGTGGTGGACCGACATATCTTGCCATTCAATCCCAACCACCAGGTTCAGTGATGGGTACTCTTAGGTCAACTGAGCAAGGAGAAATGGTGCAAGCCAAG
TTCGGGCTGCCACACACAGCTGTTAGACAGTTAGAGATATACACAACTGCAGTTCTGCTTGCAACATTAAAACCCCCAGAGCTACCTCGAGAAGAAAAAT
GGCGTAATCTCATGGATGAGATTTCAACAATCAGCTGCCAAAGTTACCGGAGCACAGTATATGAAAACCCGGAATTTCTTGCCTACTTTCACGAGGCTAC
TCCACAAGCTGAGCTTGGTTTCCTTAACATAGGAAGCCGGCCCACAAGAAGAAAGAGCTCAACAGGAATCGGACATCTTCGTGCCATTCCATGGGTATTT
GCGTGGACCCAAACAAGATTTGTTCTTCCAGCCTGGCTAGGAGTTGGGGCAGGTTTGAAGGGTGTTTGTGAGAAGGGACACACTCAAGAGTTGAAAGCAA
TGTACAAAGAATGGCCTTTCTTCCAATCTACCATAGACCTGATAGAGATGATTCTGGGGAAGGCAGACATTCATATAGCGAAGCACTATGATGAAGTGCT
CGTATCTGATAAGAAGAGGAGAGAACTTGGTGCTGAATTAAGAAGGGAGCTCTTAACAACTGAGAAGTGTGTATTGGTGGTTAGTGGGCATGAGAAGTTG
TCGGAAAACAACCGGAGCTTGAGGCGGCTTATCGAGAGCAGGCTACCGTATCTTAATCCTATGAATTTGTTACAGGTTGAGATTCTAAAGAGGTTGAGAA
GTGATGATGACAACCACAAACTCAGAGATGCTCTGCTTATTACTATAAATGGTATTGCTGCTGGGATGAGGAACACAGGTTGA
|
|||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.010G131800.1 pacid=42797723 polypeptide=Potri.010G131800.1.p locus=Potri.010G131800 ID=Potri.010G131800.1.v4.1 annot-version=v4.1
MTDITDDIAEEISFQGFDDYCKLLKNLLNDVLQREVGTEFVDKLERNLTLAQSACNLRLAGIEDTAELLEKQLASEISKMTLEEALTLARAFSHYLNLMG
IAETHHRARKTRNLANLSKSCDEVFNQLLHGGKSGDELYASVCMQEVEIVLTAHPTQINRRTLQYKHVRIAHLLEYNDRPDLTQEDREILIEDLVREITS
IWQTDELRRHKPTPVDEARAGLHIVEQSLWKAVPHFLRRVSNALKKHTGKPLPLTCTPIKFGSWMGGDRDGNPNVTAKVTRDVSLLSRWMAIDLYIREVD
SLRFELSMTRCSDKLSREAHEILERETSPEDRHESWNQPTSRNQTKLHQHAPPLPTQLPARADLPACTECGDDGGSHPKLELPGTDYMPLSRQDVQGSSN
SESSFHKSGHGSSKSIANGSIANSNGHQSAPSPRGSFTSSQLLAQRKCFAESKIGRSSFQKLLEPSPPERPGIAPYRIVLGHVKDKLMKARRRLELLLED
LPCEHEPWDYYETTDQLLEPLLLCYESLQSCGAGVLADGRLVDLIRRVATFGMVLMKLDLRQESGRHSEALDAITKYLDMGTYSEWDEEKKLEFLTRELK
SKRPLVPPTIQVAPDVKEVLDTFRVAAELGSDSLGAYVISMASNASDVLAVELLQKDARLAVSGELGRPCPRGTLRVVPLFETVKDLRGAGSVIRKLLSI
DWYSEHIVKNHNGHQEVMVGYSDSGKDAGRFTAAWELYKAQEDVAAACKDHKVKVTLFHGRGGSIGRGGGPTYLAIQSQPPGSVMGTLRSTEQGEMVQAK
FGLPHTAVRQLEIYTTAVLLATLKPPELPREEKWRNLMDEISTISCQSYRSTVYENPEFLAYFHEATPQAELGFLNIGSRPTRRKSSTGIGHLRAIPWVF
AWTQTRFVLPAWLGVGAGLKGVCEKGHTQELKAMYKEWPFFQSTIDLIEMILGKADIHIAKHYDEVLVSDKKRRELGAELRRELLTTEKCVLVVSGHEKL
SENNRSLRRLIESRLPYLNPMNLLQVEILKRLRSDDDNHKLRDALLITINGIAAGMRNTG
|
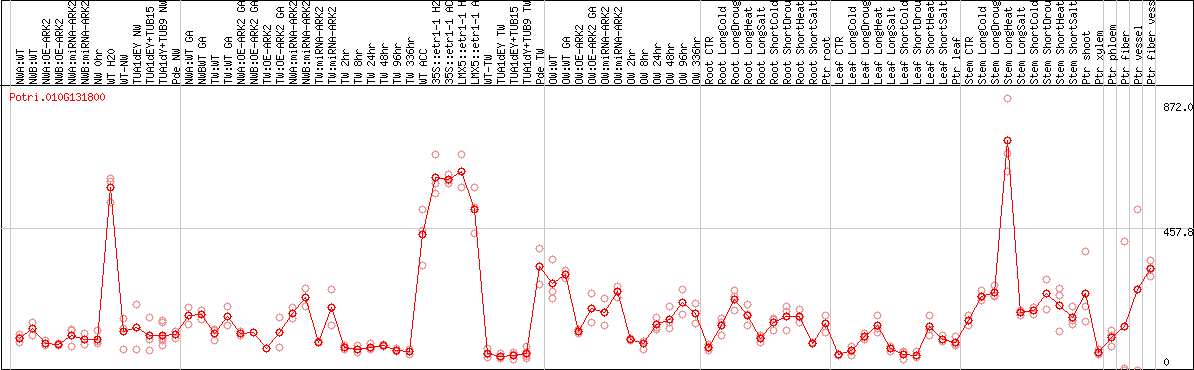
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.010G131800 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
