External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G49600 152 / 2e-47
Protein of unknown function, DUF538 (.1)
AT1G09310 123 / 9e-36
Protein of unknown function, DUF538 (.1)
AT1G56580 119 / 1e-34
SVB
SMALLER WITH VARIABLE BRANCHES, Protein of unknown function, DUF538 (.1)
AT5G46230 108 / 1e-30
Protein of unknown function, DUF538 (.1)
AT4G24130 81 / 1e-19
Protein of unknown function, DUF538 (.1)
AT1G30020 72 / 2e-16
Protein of unknown function, DUF538 (.1)
AT3G07460 40 / 0.0003
Protein of unknown function, DUF538 (.1.2)
AT5G19860 39 / 0.0004
Protein of unknown function, DUF538 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.005G011200
137 / 3e-41
AT1G56580 192 / 4e-63
SMALLER WITH VARIABLE BRANCHES, Protein of unknown function, DUF538 (.1)
Potri.013G007000
134 / 4e-40
AT1G56580 176 / 5e-57
SMALLER WITH VARIABLE BRANCHES, Protein of unknown function, DUF538 (.1)
Potri.004G064700
119 / 1e-34
AT5G46230 184 / 4e-61
Protein of unknown function, DUF538 (.1)
Potri.001G084200
103 / 2e-28
AT4G24130 208 / 5e-70
Protein of unknown function, DUF538 (.1)
Potri.003G146400
103 / 3e-28
AT4G24130 211 / 5e-71
Protein of unknown function, DUF538 (.1)
Potri.004G056700
68 / 1e-14
AT1G09310 99 / 4e-27
Protein of unknown function, DUF538 (.1)
Potri.004G084600
65 / 1e-13
AT1G56580 96 / 9e-26
SMALLER WITH VARIABLE BRANCHES, Protein of unknown function, DUF538 (.1)
Potri.017G133300
53 / 3e-09
AT1G56580 69 / 1e-15
SMALLER WITH VARIABLE BRANCHES, Protein of unknown function, DUF538 (.1)
Potri.001G008240
40 / 0.0002
AT1G55265 137 / 8e-42
Protein of unknown function, DUF538 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10037804
140 / 3e-42
AT5G49600 144 / 2e-44
Protein of unknown function, DUF538 (.1)
Lus10017087
139 / 9e-41
AT5G49600 145 / 3e-43
Protein of unknown function, DUF538 (.1)
Lus10018630
120 / 2e-34
AT1G09310 195 / 5e-64
Protein of unknown function, DUF538 (.1)
Lus10039866
118 / 2e-33
AT1G09310 196 / 5e-64
Protein of unknown function, DUF538 (.1)
Lus10031428
111 / 2e-31
AT1G09310 185 / 2e-60
Protein of unknown function, DUF538 (.1)
Lus10010904
111 / 2e-31
AT1G09310 185 / 1e-60
Protein of unknown function, DUF538 (.1)
Lus10017330
98 / 4e-26
AT4G24130 209 / 3e-70
Protein of unknown function, DUF538 (.1)
Lus10001674
89 / 8e-23
AT4G24130 212 / 2e-71
Protein of unknown function, DUF538 (.1)
Lus10000355
82 / 4e-20
AT5G46230 117 / 1e-34
Protein of unknown function, DUF538 (.1)
Lus10017655
81 / 1e-19
AT1G56580 115 / 1e-33
SMALLER WITH VARIABLE BRANCHES, Protein of unknown function, DUF538 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF04398
DUF538
Protein of unknown function, DUF538
Representative CDS sequence
>Potri.010G149100.2 pacid=42798743 polypeptide=Potri.010G149100.2.p locus=Potri.010G149100 ID=Potri.010G149100.2.v4.1 annot-version=v4.1
ATGTCAGCGCCAGAACCAACTGCAAACCTCTTGACATCAAGACATCGTAGGAGGAAAGACATGTCTATGGTGACACAAGATATCAGGGCCAAGGCAGAGG
TTTGTTATGGAGATGAAACATGCAGAGAGAAGATTATTTCCTTGCTGACAGAAAAGGGCTTGCCAAGTGGTCTGATAACAGTATTAGAAGAGATAGAGGA
ATACCGCTACATTAAGGACACAGGGTTTGTTTCGCTCAAGCACAACAGCAAGAGAAAAGATCACAAGTTCGACAAGGTAGCCGTATGCTATGACAATGAA
GTCACGGCATATTTTGAGCCCAACAGGATCAGGAATTTGACTGGTGTTAAGGCCAAGGAGTTCTTGATTTGGATTACTTTGAGTGAAATTTATGTAAGCG
GTGACATTCCAGTTGCGTTGATTACATTCAAGACTCCGGCAGGATTCTCCAAATCTTTCCCATTATCATCGTTCACTTTTAAAGATGAGGAGGAGGCGAA
TGAAGTTCATGAAGCAAAAATTGGAAAAATAAAGGGGGTGAAACTGTAA
AA sequence
>Potri.010G149100.2 pacid=42798743 polypeptide=Potri.010G149100.2.p locus=Potri.010G149100 ID=Potri.010G149100.2.v4.1 annot-version=v4.1
MSAPEPTANLLTSRHRRRKDMSMVTQDIRAKAEVCYGDETCREKIISLLTEKGLPSGLITVLEEIEEYRYIKDTGFVSLKHNSKRKDHKFDKVAVCYDNE
VTAYFEPNRIRNLTGVKAKEFLIWITLSEIYVSGDIPVALITFKTPAGFSKSFPLSSFTFKDEEEANEVHEAKIGKIKGVKL
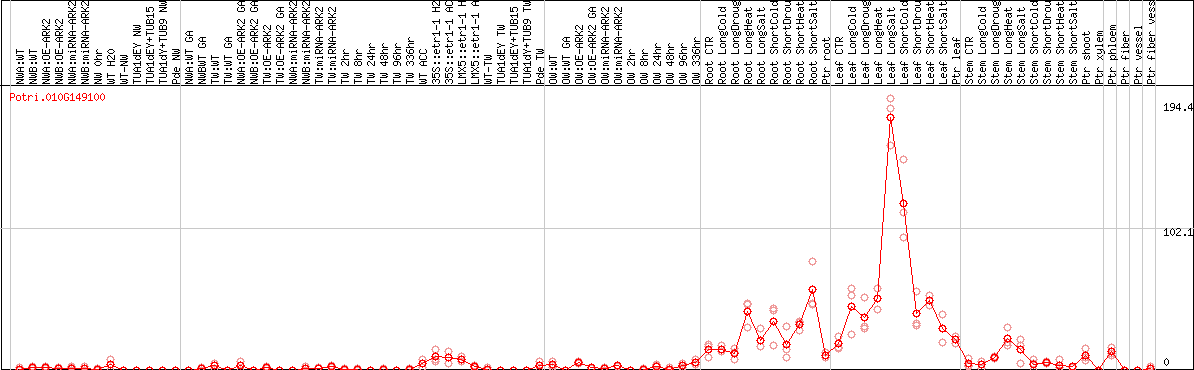
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.010G149100 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.