Potri.010G152400 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.010G152400.4 pacid=42797245 polypeptide=Potri.010G152400.4.p locus=Potri.010G152400 ID=Potri.010G152400.4.v4.1 annot-version=v4.1
ATGGCTACTGCTGAACCAGCTTCAGCTACCGTTGGGCCACGGTATGCTCCAGAAGACCCAACCCTTCCCAAACCATGGACAGGACTGATTGATGGAAGTA
CAGGTCTTTTATATTACTGGAATCCTGAAACAAACATCACACAGTATGAAAAGCCCCCATCTGTGCCGCCACAGATGCCTCCTGCTCTACCTCCTAATGC
TTCTACGCCTAAGTTGGCCCAGATACCCATGGCACACTCATCGCAACCAAATGGCTTGGTGGCTCAAGCTGCACAACAGACTATGCAAGCATCACAGCAA
CAAGGGCAACAGATTAGCCAGTTGCATCAACAATATGGACAAGTAACTTCACAACAGCAGGGTCCCCAAGTAGTGCAGGTTTCTAACCAGCAGGGTGCAC
CACAGCAAGGTTCACAGTTGGCACAGGCCATTCAACAACCGGGGCAGTTGAGATCAATGCAGCATCCAGGTCAGCATCTGCTGTCCCATGCAGGGCAGCA
AATGCCTCAACAAGGAGGTCAGCAGGTGCCCCAGCAGGTAGGACAGTACATTATGCAGCATCAAAGCTTGCAAATGCCACAACCACAAGGACATCAGTAT
GCATATCAGCAGCCAATGCAGTACATGACATATCAGCAAAACGTGCTTCCACAAGGCCAACAAAGCTCGCAGCAGCAGGCACATCTTAGTGCACAAGGGT
TGCAATTTCCTAATCAACATGAATACAAGGCAGTGTTGCCGAAGAGAGAGGAAGTTGAGTTCCAGCAAGGAAATCAAACTGGGTTTTCTCCAGCTCGGTT
TCAACAGACTGGTGGATCATCCTCTCAAAACCTGTCTTCTGCTGGAAGCAATCCGGTTTGCATGCCGCATTCTGGTGTTCAGTTGGGTCAGGCTCAGCAG
TTTGGTAGCTCTTCTGTTAATATTCAACATCCTCCTTCCATGGTTCAGATGCAGCAAATTGGGACTGATTTTCATCAGCAGCATGGTCCTAGGTTTCAAA
ATCATATGGGTCCCTCAGTGATGCACAATCAGCAGTCTAATCTGCCCCCTGCTGGGTTAAATATGGGTTATGAGAACAATGCACATGGTAGGTCTGGCAA
TGATCATTATATTAATGCTAAAGTGGAAGGGCCTGTGATGAGTCCCCATCAGCCTGAGTTTACAGCTATGTCTAGGGCAAGAAAGCAGCAGGATTCAAGG
ACAGGGAGCGTCCCATTTCAGAATGTCGGGCCTGGTCATGGTGGCGGATTTAATGCAGATGGGCATCCCAACCATAATATGTATAGTCATGCAACCAGTG
GTCCACCTTTTCCAAACAATGCATTGATGAGACCATCTTTCATCGAAACTGCAGACATTTCTAACCTCTCGCCTGCTGAAGTTTACCGCCAGGAGCATGA
AGTCTCAGCTACGGGTGACAATGTTCCTGCCCCATTCATGACTTTTGAAGCCACTGGTTTCCCTTCGGAGATATTACGAGATATACATTCTGCTGGATTC
GTATCTCCGACACCAATTCAGGCACAAACATGGCCAATTGCGCTACAAAGTAGGGATATAGTGGCCATTGCCAAAACTGGTTCCGGTAAAACATTGGGTT
ACTTGATTCCTGCCTTCATTCTTCTTCAACAGCGACGAAATAATGCTCAGAATGGACCTACAGTGTTAGTCTTGGCTCCAACCAGGGAGCTTGCCACGCA
AATACAAGATGAGGTGATGAAATTTGGTCGATCTTCTAGGGTCTCTTGCACGTGCTTGTATGGTGGAGCTCCAAAGATTCCTCAACTGAAAGAATTAGAA
AGGGGAGCAGATATAGTTGTGGCAACACCTGGTCGACTCAATGACATCCTTGAAATGAAAAGGATTGACTTTAGGCAAGTTTCACTCCTTGTGCTTGATG
AGGCTGATCGAATGCTTGACATGGGATTTGAGCCTCAGATCCGTAAGATTGTGAACGAAATACCTCCTCAAAGGCAAACACTTATGTTCACTGCAACCTG
GCCAAAAGAAGTGAGGAAAATAGCAAGTGATCTTCTTGTTCATCCTGTCCAGGTGAACATTGGCAGTGTTGATGTGCTTGCTGCCAACAAGTCAATCACA
CAGTATGTTGAAGTGGTCCCACAAATGGAAAAGGATCGGCGCTTGGAGCAAATCCTTAGAACACAAGAGAGAGGTTCCAAGGCTATCATTTTCTGCTCTA
CAAAGAGATTGTGTGACCAGCTTGCACGGAGTATTGGACGGAACTTTGGAGCCGCTGCAATCCATGGAGACAAGTCTCAAGGTGAGAGAGACTGGGCTTT
AAACCAATTCCGGAGTGGAAAATCTCCAATTTTAGTTGCCACTGACGTTGCTGCCCGTGGTCTTGATATCAAAGACATAAGGATTGTTATCAACTACGAC
TTCCCCTCTGGGATTGAGGACTATGTTCACCGAATTGGTAGAACTGGGAGGGCTGGTGCAACTGGGGTGTCATACACCTTTTTCTCTGAGCAGGACTGGA
AATATGCAGCTGACCTGGTTAAACTGCTGGAGGGAGCTAATCAGCATGTGCCCGTAGAGGTGCGAGAGATGGCGTTGCGTGGTGGACCTAGCTTTGGGAA
GGATCGAGGAGGGTTGAACCGTTTTGATGCTGGCAGGGGTGGTGGGCGGTGGGCTTCTGGTGGCCGTGGTGGGATGAGAGATGGTGGCTTTGGTGGTCGT
GGTGGGATGAGAGATGGTGGCTTTGGTGGCCGTGGTGGGATGCGGGAAAACAGCTTTGGGGGACGTGGTGGGATGAGGGAAAACAGCTTTGGGGGACGTG
GTGGGATGAGGGATGGGAACTTTGGGGGACGTGGTGGGAGAAACGACTTGTTTTCTGGGCGGGAAAACAGGGGAAGGGGTTTTGGTGGCTCTGGTGGTGG
ACATGTAGGTTGGGGTAGGAATGATCGAGGCCCAAATGATCGTCATAGTTACATGGATGGCCGTGGACGTGGTCATGGAAGATTCGATGGTAGAAGGGAT
ATGACTGATAAGAGTAGGGGCAGAAGTTTCAGTCGAAGCCCTGATAGGGCTCGAACACGGGGTTATAGCCGGAGCCGGAGCCGGAGCCGTAGTAGGAGTC
GGAGCTGGACCAGAAGCCGCAGCCGCAGCCCTCGAGACAGCCGCAGCCGCAGCCCCCGACACAGCCGCAGCCGCAGCCGCAGCCCCCGACGCAGCCGCAG
CCGCAGCGGCATCCGCAGCCATAGTCCTCAACATGCCAGTAGTCACAGCCTTAGCCCTCGAAGGAGCCCTGACTATGATAGATTTGAAAGGCCTCGAGAA
AGAAACATTGATGAGAAGGATATGACAGTTCCAGAATTGAAGGCTTCAGAACAAGAAATGTCTCCAATGTCGCCGGGCACACATGGTAATGCATTTTCTG
GAGCAGAGCTGCTGCCAGCCACTGGGAGTGCTGAGCCAGTGCAACAGGAGGCTACAGGAGATCCAGGGCTTCCTTCAGTTGCTGAAGCTTGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.010G152400.4 pacid=42797245 polypeptide=Potri.010G152400.4.p locus=Potri.010G152400 ID=Potri.010G152400.4.v4.1 annot-version=v4.1
MATAEPASATVGPRYAPEDPTLPKPWTGLIDGSTGLLYYWNPETNITQYEKPPSVPPQMPPALPPNASTPKLAQIPMAHSSQPNGLVAQAAQQTMQASQQ
QGQQISQLHQQYGQVTSQQQGPQVVQVSNQQGAPQQGSQLAQAIQQPGQLRSMQHPGQHLLSHAGQQMPQQGGQQVPQQVGQYIMQHQSLQMPQPQGHQY
AYQQPMQYMTYQQNVLPQGQQSSQQQAHLSAQGLQFPNQHEYKAVLPKREEVEFQQGNQTGFSPARFQQTGGSSSQNLSSAGSNPVCMPHSGVQLGQAQQ
FGSSSVNIQHPPSMVQMQQIGTDFHQQHGPRFQNHMGPSVMHNQQSNLPPAGLNMGYENNAHGRSGNDHYINAKVEGPVMSPHQPEFTAMSRARKQQDSR
TGSVPFQNVGPGHGGGFNADGHPNHNMYSHATSGPPFPNNALMRPSFIETADISNLSPAEVYRQEHEVSATGDNVPAPFMTFEATGFPSEILRDIHSAGF
VSPTPIQAQTWPIALQSRDIVAIAKTGSGKTLGYLIPAFILLQQRRNNAQNGPTVLVLAPTRELATQIQDEVMKFGRSSRVSCTCLYGGAPKIPQLKELE
RGADIVVATPGRLNDILEMKRIDFRQVSLLVLDEADRMLDMGFEPQIRKIVNEIPPQRQTLMFTATWPKEVRKIASDLLVHPVQVNIGSVDVLAANKSIT
QYVEVVPQMEKDRRLEQILRTQERGSKAIIFCSTKRLCDQLARSIGRNFGAAAIHGDKSQGERDWALNQFRSGKSPILVATDVAARGLDIKDIRIVINYD
FPSGIEDYVHRIGRTGRAGATGVSYTFFSEQDWKYAADLVKLLEGANQHVPVEVREMALRGGPSFGKDRGGLNRFDAGRGGGRWASGGRGGMRDGGFGGR
GGMRDGGFGGRGGMRENSFGGRGGMRENSFGGRGGMRDGNFGGRGGRNDLFSGRENRGRGFGGSGGGHVGWGRNDRGPNDRHSYMDGRGRGHGRFDGRRD
MTDKSRGRSFSRSPDRARTRGYSRSRSRSRSRSRSWTRSRSRSPRDSRSRSPRHSRSRSRSPRRSRSRSGIRSHSPQHASSHSLSPRRSPDYDRFERPRE
RNIDEKDMTVPELKASEQEMSPMSPGTHGNAFSGAELLPATGSAEPVQQEATGDPGLPSVAEA
|
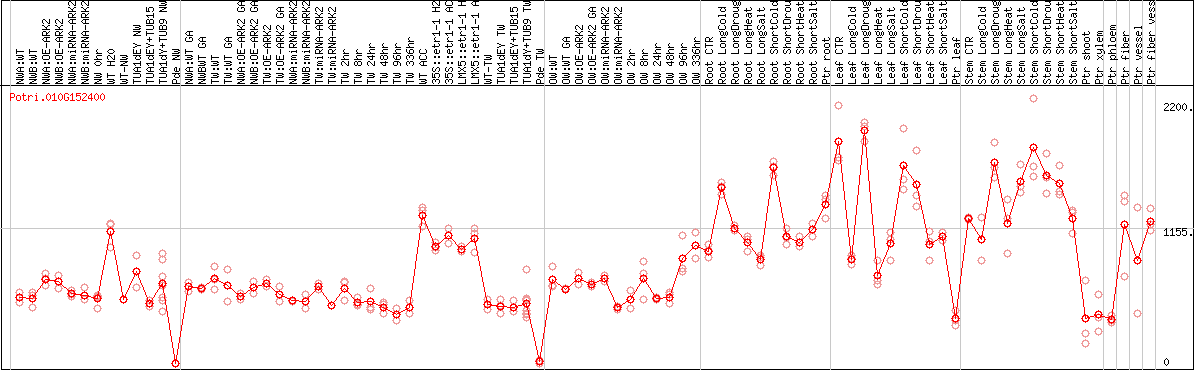
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.010G152400 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
