Potri.010G153800 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.010G153800.4 pacid=42797620 polypeptide=Potri.010G153800.4.p locus=Potri.010G153800 ID=Potri.010G153800.4.v4.1 annot-version=v4.1
ATGGCTCAGGTGGCATACTCTGATGAGACAGAGTCAAAGAAGATTGACTCAGCAGAGCTTGGAGGAAGCATAAGAACCTCATTCAGAAGTCACGAGCCTA
GTTTTCATAGCCTTTCAATTGGGAATGCCAACCATCGTAGAAATGAAAATGAAGAAGACGCATCACAATGCTTGGCTACTATTGAGAGATTACCTTCTTT
TGAACGGATTTCTACTGCTTTATCGGAGGAGAAAGATGGCACAAATGGAAAAGGAGATGCAATGGGGGGAAAAGTTGTTAATGTTGCCAAACTTCGAGCT
CAAGAAGGGCATGTGTTCAATGAAAAGCTGATCAAGCACGTTGAAAATGATAATCTCCGATTGTTGCAGAAGCTTAGGAAAAGAATTGACATCGCTGGTA
TACAATTGCCCACCGTAGAAGTGAAATACAGAAATGTTTGTGTTGAAGCAGACTGCGAGGTAGTTCGTGGCAAACCTCTTCCCACTCTGTGGAGCACTGC
TAAGAGCATTCTCTCTGGGTTTGCCAATCTGTCACGTTCGAAACAAAGAACAAAGATAAGCATCATAAAAGATGTCAGCGGCATCATTAAGCCAGGAAGG
ATGACTTTACTGCTTGGCCCTCCAGGTTGTGGCAAAACCACTCTGTTGAAGGCTCTCTCTGGGAAACCTAGCAATTCTCTCAAGGTTGCAGGGGAAATAT
CTTACAATGGTCACAGATTGGAAGAGTTTGTTCCTCAGAAAACCGCAGCTTATGTAAGCCAATATGATTTGCATATACCAGAGATGACAGTGAGGGAGAC
GGTTGATTTTTCAGCACGCTGTCAGGGCACAGGAAGCCAAGCTGAGATCTTGATGGAAATCAGTAGAAAGGAGAAGCAGGCAGGGATTCTCCAAGATACT
GATTTGGATACATACATGAAGGGAATATCAGAAGAAGGAGCGAAAATTACCCTTCAAACAGACTACGTTTTGGAGATTCTAGGACTTGATATTTGTGCCG
ACACAATGGTCGGGGATACTATGAGAAGAGGTATCTCAGGTGGACAGAAGAAAAGACTAAGTACAGGAGAAATGGTTGTGGGTCCAATGAAAGCCCTATT
CATGGATGAAATATCAAATGGCTTAGACAGTTCGACCACTTTCCAGATTGTGTCTTGTATGCAGCATTTGGCACATATTACCGATGCCACTGTGTTGATT
TCACTTCTTCAGCCAGCACCAGAGATCTTTGACCTCTTTGATGACATCATGTTGATGGCAGAAGGAATGGTTGTTTATCATGGCCCACGTTCTAGTGTTT
GCAGATTCTTCGAGGACTCTGGGTTTAGGTGCCCTGAGAGAAAAGAAGTTGCAGACTTCCTTCAAGAGGTTATCTCTAGAAAAGATCAAAGGCAGTATTG
GTACTGCACAGAACAACCTCACAGTTATGTTTCTGTAGAACAGTTTGTTAAGAAATTCAAAGAAAGCCAGCTTGGTCAGATGCTAGATGAGGAGATCATG
AAGCCATTTGATAAGTCTAATAGCCACAAGACTGCTTTATGCTTCAGAAAATACTCATTATCAAAATGGGAACTGTTCAAAGTTTGTAGTACGAGAGAAT
TTGTTCTCATGAAGAGGAATTCTTTCATTTATGTTTTTAAATGTACACAGCTAGTGATCACTGCTTCAATAACAATGACTGTCTTCCTGCGCACAAGGAT
GGCTGTTGATGCGATCCATGCAAGTTATTACATGAGCGCTTTATTCTTTGCATTGACTATATTATTTTCTGATGGAATCCCAGAGTTGCATATGACTGTA
TCCCGACTTGCAGTTTTTTATAAGCAGAGAGAGTTATGCTTTTACCCTGCTTGGGCTTATGTTGTTCCCACAGCTATCCTTAAAGTTCCACTTTCTCTCG
TGGAAGCATTTGTTTGGACAACTCTTACTTATTATGTTGTTGGTTACAGCCCAGAGTTTGGAAGGTTCTTCCGCCAGTTCCTTCTACTCTTTCTGGTACA
TTCAACATCAATATCCATGTTTCGCTTTGTCGCCTCCCTCTTCCAAACTATGGTTGCTTCAGTGACAGCTGGAGGACTCGCTTTATTGATTACTTTACTA
TTTGGTGGCTTTTTAATCCCAAAGCCCTCTATGCCTGTGTGGTTGGGATGGGGATTTTGGATTTCTCCGTTGGCGTATGGTGAAATAGGCTTAAGTCTGA
ATGAGTTTCTTACTCCTAGGTGGGCAAAGACCGTGTCAGGTAACACAACCATACAACAGCAAACTCTGGAAAGCCGTGGATTGAACTTTCACGGTTATTT
TTACTGGATATCAGTTGGAGCTTTAATTGGGTTGACAGTGCTCTTCAACGTTGGCTTTGCTTTGGCTTTAACCTTCTTGAAATCCCCAGGAAACTCTCGA
GCTATTATTTCTTATGAAAGATACTATCAACAGCAAGGAAAACTAGATGATGGTGCTTCTTTTGACATAAACAACGACAAGAAGACCCTCACCTGTGCAT
GTCCTAAATCTAGTCCAGGAGATAAAAAGGGAAGGATGGCTTTACCTTTTGAGCCCCTTACTATGACATTTAAGGATGTGCGGTACTATGTTGATACTCC
TTTGGAAATGAGAAAACGTGGTTTTCCACAGAAGAAGCTCCAGCTTCTTTCAGACATTACTGGTGCATTCAGACCTGGTATTCTTACAGCATTAATGGGT
GTTAGTGGAGCCGGGAAAACAACTCTTATGGATGTTCTTTCCGGGAGGAAAACTGGAGGCACTATTGAAGGAGAGATTAGAATTGGTGGGTACCCAAAGG
TTCAGCATTCATTCGCTAGAGTATCAGGTTACTGCGAACAAACAGACATACATTCCCCCCAGATTACTGTAGAAGAATCTGTGATATATTCTGCTTGGTT
GCGGCTTCCTCCTGAAATTGATACGAAAACTAAATATGAATTTGTAAATCAAGTCCTTGAAACTATCGAGCTTGATGAAATCAAAGATTCTTTAGTAGGC
ATTCCCGGAATTAGTGGTTTATCAATTGAACAGCGTAAACGACTGACAGTAGCTGTGGAACTTGTTGCCAATCCATCCATCATATTCATGGATGAACCGA
CTTCAGGTTTGGATGCAAGAGCAGCTGCGATTGTCATGAGAGTAGTGAAGAACATAGTTGAAACAGGAAGAACAATTGTTTGCACCATCCACCAGCCAAG
TATAGATATCTTCGAGGCATTTGATGAGTTGATTCTTATGAAAATTGGTGGACGCATAATATATTCTGGACCACTGGGTCAGCGGTCAAGCAAGGTCATT
GAGTATTTTGAGAATATTCCTGGAGTGCCTAAGATTAAGAACAGATACAATCCAGCAACATGGATGTTAGAGGTTTCTTCTAAAACTGCAGAGGCTGATC
TTGGTGTTGATTTTGGTGAAGCTTATGAAGGGTCCACCCTATATGAGGAGAACAAAGAGTTGGTTAAGCAATTGAGCTCACCAACTCCTGGCTCAAAAGA
CTTGCATTTTCCTACCTGTTTTCCGCAGAATGGCTGGGAGCAACTCAAAGCATGCTTGTGGAAACAGCACTTGTCTTATTGGCGAAGTCCTTCGTATAAT
CTTTTGCGCATTGTCTTTATGTCCTTTGGAGCTCTCCTATTTGGCCTACTGTTCTGGCAGCAAGGAAATAAAATAAACAACCAGCAAGACTTGTTCAGTA
TCGCTGGCTCGATGTACTCTATAATAATATTTTTCGGCATAAACAATTGTTCTCCAGTCCTAGCATTTGTTGCAAGAGAGCGGACTGTTTTCTACCGTGA
AAGATTTGCAGGAATGTACTCTTCATGGGCCTATTCATTTGCACAGGTGCTGGTTGAGGTTCCATACTTGCTCATTGAAGGGATTCTTTATGTGATCATC
ACATATCCTATGATTGGCTACTCCTTGTCTGCCTATAAGATATTCTGGTCTTTCTATAGCATGTTTTGTATGCTGCTGTTTTTCAACTACCTAGGAATGC
TGCTCGTATCATTGACACCAAACATTCAAGTCGCCTCCAATTTAGCTGCGTTTGCGTACACAACGCTGAATTTCTTCTCGGGGTTTATCGTGCCGAAACC
ATACATCCCAAAGTGGTGGGTTTGGTTGTATTATATATGCCCCTCATCTTGGACATTAAATGCTATGCTCACCTCGCAGTATGGAGATGTGAACAAAGAA
ATATCAGTGTTTGGAGAGACGATGACTGTTGCTGATTTTGTAGGTGATTACTTTGGGTTTCACCACAACTTTTTGGGTGTTGTCGGTGTTGTGCTAATCA
TCTTTCCCATTATTACCGCATCTCTTTTTGCTTATTTCTTTGGAAGACTGAACTTCCAGAGAAGGTAG
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.010G153800.4 pacid=42797620 polypeptide=Potri.010G153800.4.p locus=Potri.010G153800 ID=Potri.010G153800.4.v4.1 annot-version=v4.1
MAQVAYSDETESKKIDSAELGGSIRTSFRSHEPSFHSLSIGNANHRRNENEEDASQCLATIERLPSFERISTALSEEKDGTNGKGDAMGGKVVNVAKLRA
QEGHVFNEKLIKHVENDNLRLLQKLRKRIDIAGIQLPTVEVKYRNVCVEADCEVVRGKPLPTLWSTAKSILSGFANLSRSKQRTKISIIKDVSGIIKPGR
MTLLLGPPGCGKTTLLKALSGKPSNSLKVAGEISYNGHRLEEFVPQKTAAYVSQYDLHIPEMTVRETVDFSARCQGTGSQAEILMEISRKEKQAGILQDT
DLDTYMKGISEEGAKITLQTDYVLEILGLDICADTMVGDTMRRGISGGQKKRLSTGEMVVGPMKALFMDEISNGLDSSTTFQIVSCMQHLAHITDATVLI
SLLQPAPEIFDLFDDIMLMAEGMVVYHGPRSSVCRFFEDSGFRCPERKEVADFLQEVISRKDQRQYWYCTEQPHSYVSVEQFVKKFKESQLGQMLDEEIM
KPFDKSNSHKTALCFRKYSLSKWELFKVCSTREFVLMKRNSFIYVFKCTQLVITASITMTVFLRTRMAVDAIHASYYMSALFFALTILFSDGIPELHMTV
SRLAVFYKQRELCFYPAWAYVVPTAILKVPLSLVEAFVWTTLTYYVVGYSPEFGRFFRQFLLLFLVHSTSISMFRFVASLFQTMVASVTAGGLALLITLL
FGGFLIPKPSMPVWLGWGFWISPLAYGEIGLSLNEFLTPRWAKTVSGNTTIQQQTLESRGLNFHGYFYWISVGALIGLTVLFNVGFALALTFLKSPGNSR
AIISYERYYQQQGKLDDGASFDINNDKKTLTCACPKSSPGDKKGRMALPFEPLTMTFKDVRYYVDTPLEMRKRGFPQKKLQLLSDITGAFRPGILTALMG
VSGAGKTTLMDVLSGRKTGGTIEGEIRIGGYPKVQHSFARVSGYCEQTDIHSPQITVEESVIYSAWLRLPPEIDTKTKYEFVNQVLETIELDEIKDSLVG
IPGISGLSIEQRKRLTVAVELVANPSIIFMDEPTSGLDARAAAIVMRVVKNIVETGRTIVCTIHQPSIDIFEAFDELILMKIGGRIIYSGPLGQRSSKVI
EYFENIPGVPKIKNRYNPATWMLEVSSKTAEADLGVDFGEAYEGSTLYEENKELVKQLSSPTPGSKDLHFPTCFPQNGWEQLKACLWKQHLSYWRSPSYN
LLRIVFMSFGALLFGLLFWQQGNKINNQQDLFSIAGSMYSIIIFFGINNCSPVLAFVARERTVFYRERFAGMYSSWAYSFAQVLVEVPYLLIEGILYVII
TYPMIGYSLSAYKIFWSFYSMFCMLLFFNYLGMLLVSLTPNIQVASNLAAFAYTTLNFFSGFIVPKPYIPKWWVWLYYICPSSWTLNAMLTSQYGDVNKE
ISVFGETMTVADFVGDYFGFHHNFLGVVGVVLIIFPIITASLFAYFFGRLNFQRR
|
DESeq2's median of ratios [POPLAR]
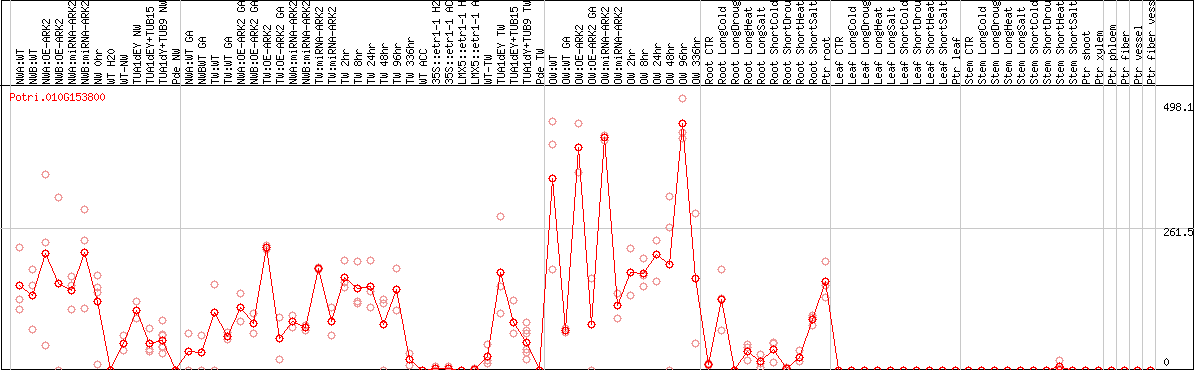
Coexpressed genes
Potri.010G153800 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
