External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G61240 517 / 0
Leucine-rich repeat (LRR) family protein (.1), Leucine-rich repeat (LRR) family protein (.2)
AT1G13910 445 / 4e-158
Leucine-rich repeat (LRR) family protein (.1)
AT5G46330 155 / 5e-42
FLS2
FLAGELLIN-SENSITIVE 2, Leucine-rich receptor-like protein kinase family protein (.1)
AT1G35710 149 / 1e-39
Protein kinase family protein with leucine-rich repeat domain (.1)
AT4G08850 147 / 5e-39
Leucine-rich repeat receptor-like protein kinase family protein (.1.2)
AT3G11080 146 / 6e-39
AtRLP35
receptor like protein 35 (.1)
AT4G20140 143 / 7e-38
GSO1
GASSHO1, Leucine-rich repeat transmembrane protein kinase (.1)
AT4G36180 143 / 9e-38
Leucine-rich receptor-like protein kinase family protein (.1)
AT5G61480 142 / 3e-37
PXY, TDR
TDIF receptor, PHLOEM INTERCALATED WITH XYLEM, Leucine-rich repeat protein kinase family protein (.1)
AT5G44700 141 / 3e-37
GSO2, EDA23
GASSHO 2, EMBRYO SAC DEVELOPMENT ARREST 23, Leucine-rich repeat transmembrane protein kinase (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.008G093400
568 / 0
AT5G61240 513 / 0.0
Leucine-rich repeat (LRR) family protein (.1), Leucine-rich repeat (LRR) family protein (.2)
Potri.006G014232
151 / 1e-40
AT1G53440 504 / 5e-161
Leucine-rich repeat transmembrane protein kinase (.1)
Potri.001G075000
151 / 1e-40
AT4G20140 1371 / 0.0
GASSHO1, Leucine-rich repeat transmembrane protein kinase (.1)
Potri.012G124300
149 / 6e-40
AT4G08850 727 / 0.0
Leucine-rich repeat receptor-like protein kinase family protein (.1.2)
Potri.018G045500
148 / 2e-39
AT2G25790 925 / 0.0
Leucine-rich receptor-like protein kinase family protein (.1)
Potri.006G237400
146 / 9e-39
AT2G25790 885 / 0.0
Leucine-rich receptor-like protein kinase family protein (.1)
Potri.004G065400
145 / 2e-38
AT5G46330 1176 / 0.0
FLAGELLIN-SENSITIVE 2, Leucine-rich receptor-like protein kinase family protein (.1)
Potri.002G008375
144 / 4e-38
AT4G08850 557 / 0.0
Leucine-rich repeat receptor-like protein kinase family protein (.1.2)
Potri.002G008100
143 / 8e-38
AT4G08850 565 / 0.0
Leucine-rich repeat receptor-like protein kinase family protein (.1.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10037108
528 / 0
AT5G61240 550 / 0.0
Leucine-rich repeat (LRR) family protein (.1), Leucine-rich repeat (LRR) family protein (.2)
Lus10026651
526 / 0
AT5G61240 549 / 0.0
Leucine-rich repeat (LRR) family protein (.1), Leucine-rich repeat (LRR) family protein (.2)
Lus10004662
508 / 0
AT5G61240 543 / 0.0
Leucine-rich repeat (LRR) family protein (.1), Leucine-rich repeat (LRR) family protein (.2)
Lus10036961
472 / 4e-169
AT5G61240 488 / 1e-175
Leucine-rich repeat (LRR) family protein (.1), Leucine-rich repeat (LRR) family protein (.2)
Lus10008712
152 / 5e-41
AT4G08850 1009 / 0.0
Leucine-rich repeat receptor-like protein kinase family protein (.1.2)
Lus10020935
151 / 2e-40
AT4G08850 1011 / 0.0
Leucine-rich repeat receptor-like protein kinase family protein (.1.2)
Lus10027688
149 / 9e-40
AT4G20140 707 / 0.0
GASSHO1, Leucine-rich repeat transmembrane protein kinase (.1)
Lus10038488
142 / 2e-37
AT5G46330 726 / 0.0
FLAGELLIN-SENSITIVE 2, Leucine-rich receptor-like protein kinase family protein (.1)
Lus10023681
142 / 2e-37
AT2G33170 1152 / 0.0
Leucine-rich repeat receptor-like protein kinase family protein (.1)
Lus10006045
142 / 3e-37
AT2G25790 858 / 0.0
Leucine-rich receptor-like protein kinase family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0022
LRR
PF13855
LRR_8
Leucine rich repeat
Representative CDS sequence
>Potri.010G160700.3 pacid=42798255 polypeptide=Potri.010G160700.3.p locus=Potri.010G160700 ID=Potri.010G160700.3.v4.1 annot-version=v4.1
ATGGCGCGTCCATCACTCTCCCTCCCTCTCATTCTCATTTTCCCTCTTCTTTTCCATCTTGCTCTTGCCAAAACCCTTAAACGCGACGTGAAGGCATTGA
ATGAAATAAAGGCATCTCTAGGCTGGAGAGTGGTATACGCTTGGGTCGGCGACGATCCTTGTGGTGACGGTGACCATCCGCCGTGGTCTGGTGTCACATG
CTCTACTGTAGGAGATTATAGAGTCGTCACCGAATTGGAAGTATACGCGGTATCTATTGTGGGGCCATTTCCTACTGCTGTAACGAATTTGCTGGATCTT
ACTAGACTGGATCTTCACAATAACAAGCTCACAGGGCCCATTCCTCCACAAATTGGCCGATTGAAGCGTCTTAAGATACTTAATTTGAGGTGGAATAAAC
TACAAGATGTTATTCCTCCCGAAATTGGTGAACTGAAGAGTTTAACTCATCTTTATCTGAGTTTCAATGCTTTCAAAGGAGAAATCCCCAAGGAGCTTGC
AATTCTTCCTGAGCTTCGGTATCTTTATCTACATGAGAATCGCTTTTCTGGGCGAATTCCTGCAGAGCTGGGGACACTGAAAAATCTTCGGCACCTTGAT
GTTGGCAACAATCATTTGGTGGGTACTATAAGGGAACTAATACGTCTTGATGGTTGCTTCCCTGCTCTTCGCAACCTTTATTTAAATGATAATTATCTTA
CTGGAGGAGTCCCTGCACAACTTTCAAACCTGACCAGCTTGGAGATCTTGCACCTATCACACAATAGAATGACTGGAATAATACCTGTTGGGCTTGCTCA
TATGCCTAGATTGACTTACTTGTACTTAGATCACAACAATTTTAATGGGAGAATTCCTGACGCCTTCTATAAGCATCCATATTTGAAAGAATTGTATATC
GAAGGAAATGCATTCAAGCCAGGTGTAAACCCAATAGGTGTTCACAAAGTCCTTGAAGTGTCTGATACCGAGTTTGTAGTTTAG
AA sequence
>Potri.010G160700.3 pacid=42798255 polypeptide=Potri.010G160700.3.p locus=Potri.010G160700 ID=Potri.010G160700.3.v4.1 annot-version=v4.1
MARPSLSLPLILIFPLLFHLALAKTLKRDVKALNEIKASLGWRVVYAWVGDDPCGDGDHPPWSGVTCSTVGDYRVVTELEVYAVSIVGPFPTAVTNLLDL
TRLDLHNNKLTGPIPPQIGRLKRLKILNLRWNKLQDVIPPEIGELKSLTHLYLSFNAFKGEIPKELAILPELRYLYLHENRFSGRIPAELGTLKNLRHLD
VGNNHLVGTIRELIRLDGCFPALRNLYLNDNYLTGGVPAQLSNLTSLEILHLSHNRMTGIIPVGLAHMPRLTYLYLDHNNFNGRIPDAFYKHPYLKELYI
EGNAFKPGVNPIGVHKVLEVSDTEFVV
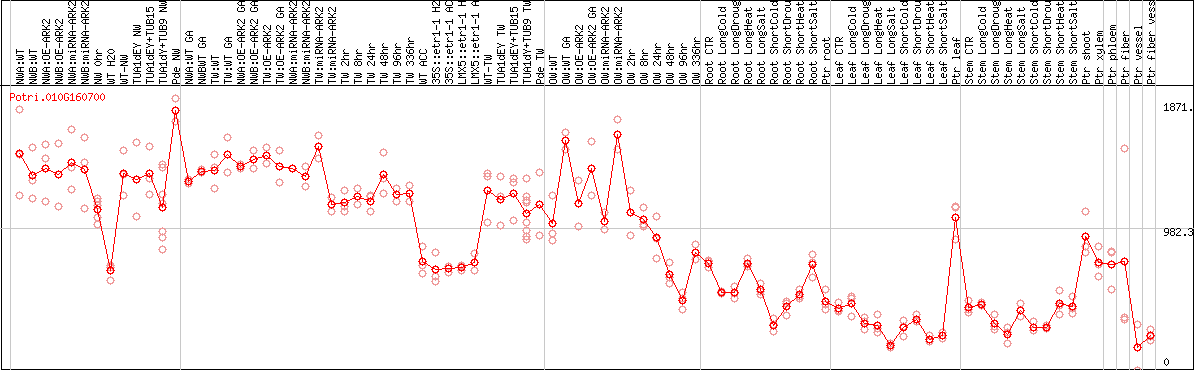
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.010G160700 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.