External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G40200 131 / 4e-37
bHLH
bHLH051
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
AT3G25710 105 / 3e-26
bHLH
TMO5, BHLH32, bHLH032, ATAIG1
TARGET OF MONOPTEROS 5, basic helix-loop-helix 32 (.1)
AT3G56770 92 / 3e-22
bHLH
bHLH107
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.2)
AT1G68810 94 / 4e-22
bHLH
bHLH030
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
AT2G41130 75 / 6e-16
bHLH
bHLH106
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
AT4G38070 65 / 1e-11
bHLH
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
AT1G12860 52 / 1e-07
bHLH
SCRM2, ICE2, bHLH033
SCREAM 2, INDUCER OF CBF EXPRESSION 2, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
AT3G26744 52 / 2e-07
bHLH
SCRM, ATICE1, ICE1, bHLH116
SCREAM, A. THALIANA INDUCER OF CBP EXPRESSION 1, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.2), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.4)
AT5G65640 50 / 6e-07
bHLH
bHLH093
beta HLH protein 93 (.1.2)
AT2G16910 50 / 7e-07
bHLH
AMS, bHLH021
ABORTED MICROSPORES, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.008G070800
429 / 3e-154
AT2G40200 132 / 2e-37
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Potri.019G089000
100 / 3e-25
AT1G68810 122 / 2e-32
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Potri.002G105300
100 / 5e-25
AT3G56770 110 / 1e-29
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.2)
Potri.010G130000
98 / 2e-23
AT1G68810 269 / 1e-87
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Potri.013G117600
96 / 3e-23
AT3G56770 112 / 8e-30
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.2)
Potri.006G037100
92 / 2e-22
AT2G41130 229 / 2e-75
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Potri.008G116000
93 / 7e-22
AT1G68810 259 / 5e-84
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Potri.016G035400
89 / 6e-21
AT2G41130 231 / 2e-76
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Potri.004G055700
82 / 4e-18
AT2G41130 120 / 1e-32
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10028978
189 / 4e-60
AT2G40200 157 / 8e-48
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Lus10007502
189 / 1e-59
AT2G40200 163 / 8e-50
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Lus10018020
104 / 2e-26
AT2G41130 206 / 1e-65
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Lus10042017
103 / 3e-26
AT2G41130 204 / 2e-65
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Lus10013917
97 / 7e-24
AT2G40200 122 / 2e-33
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Lus10035056
94 / 5e-22
AT1G68810 321 / 6e-108
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Lus10001885
92 / 5e-22
AT1G68810 124 / 1e-33
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Lus10021711
93 / 1e-21
AT1G68810 310 / 1e-103
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Lus10034271
91 / 4e-21
AT1G68810 218 / 2e-68
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Lus10041493
91 / 5e-21
AT3G25710 201 / 2e-62
TARGET OF MONOPTEROS 5, basic helix-loop-helix 32 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF00010
HLH
Helix-loop-helix DNA-binding domain
Representative CDS sequence
>Potri.010G186700.1 pacid=42798699 polypeptide=Potri.010G186700.1.p locus=Potri.010G186700 ID=Potri.010G186700.1.v4.1 annot-version=v4.1
ATGGAAGATCACTATTCTCCTTGCTGGCCTGCTGCTCCTGCAGAGGCAAACTGGGATCAAACAAGTGCTGCAGTATATGATGAGTCCTTTTTAGTCCCAT
GCCCATCTCATGCTTCTGCCTCTGCCAACTTTCAAGTATACGGATTCCCTTCGTGGTCAGTACCGCTTCAGGAGGCTTCAGAAGATAAAGCTGCTTCGAG
TTCCAAGAGTCATAGCCAAGCTGAGAAGCGGCGACGGGACAGGATTAATGCACAACTGGGAATTCTTAGGAAACTCGTTCCTAAATCAGAAAAGATGGAC
AAGGCAGCTCTCCTAGGAAGTGCGATTGATCATGTGAAGGATCTCAAACAAAAAGCAACGGAAATCAGCAGAACTTTCACAATCCCAACTGAAGTTGATG
AAGTGACGGTTGATTGTGATGTTTCTCAAGTTACAAGCCCTCCCAGCACCAACAAAGACAAGGACAACACATTTATCAGGGCATCTGTTTGCTGCGATGA
TCGGCCAGAACTGTTTTCGGAGCTTATTACGGTGCTCAAAGGCCTTAGACTAACCATAGTTAGAGCTGATATTGCTAGTGTAGGTGGAAGAGTTAAGAGC
ATATTAGTACTTTGCAGTGAGTGCAGTGAAGAGGGGAGTGTTTCAATTAGCACTATTAAACAGTCGCTTAATCTAGTTTTGAGTAGAATCGCTTCATCAT
CAGTGCCATCAAATTATCGTATTAGAAGCAAGAGGCAAAGGTTCTTTTTGCCTTCTCATTTGTCAGAACAATATGAATAA
AA sequence
>Potri.010G186700.1 pacid=42798699 polypeptide=Potri.010G186700.1.p locus=Potri.010G186700 ID=Potri.010G186700.1.v4.1 annot-version=v4.1
MEDHYSPCWPAAPAEANWDQTSAAVYDESFLVPCPSHASASANFQVYGFPSWSVPLQEASEDKAASSSKSHSQAEKRRRDRINAQLGILRKLVPKSEKMD
KAALLGSAIDHVKDLKQKATEISRTFTIPTEVDEVTVDCDVSQVTSPPSTNKDKDNTFIRASVCCDDRPELFSELITVLKGLRLTIVRADIASVGGRVKS
ILVLCSECSEEGSVSISTIKQSLNLVLSRIASSSVPSNYRIRSKRQRFFLPSHLSEQYE
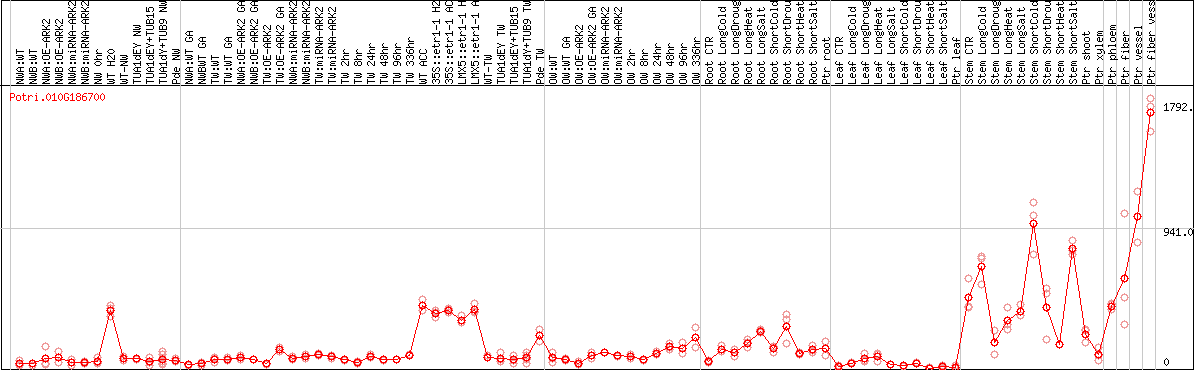
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.010G186700 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.