External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G55470 163 / 3e-52
Calcium-dependent lipid-binding (CaLB domain) family protein (.1), Calcium-dependent lipid-binding (CaLB domain) family protein (.2)
AT1G63220 96 / 7e-26
Calcium-dependent lipid-binding (CaLB domain) family protein (.1), Calcium-dependent lipid-binding (CaLB domain) family protein (.2)
AT4G00467 77 / 7e-18
Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
AT5G47710 60 / 1e-11
Calcium-dependent lipid-binding (CaLB domain) family protein (.1), Calcium-dependent lipid-binding (CaLB domain) family protein (.2)
AT4G21160 56 / 1e-09
ZAC, AGD12
ARF-GAP domain 12, Calcium-dependent ARF-type GTPase activating protein family (.1.2.3.4)
AT3G07940 55 / 3e-09
Calcium-dependent ARF-type GTPase activating protein family (.1)
AT1G70800 52 / 7e-09
EHB1
ENHANCED BENDING 1, Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
AT4G05330 50 / 8e-08
AGD13
ARF-GAP domain 13 (.1)
AT1G05500 50 / 2e-07
SYT5, NTMCTYPE2.1, ATSYTE ,NTMC2T2.1 ,NTMC2TYPE2.1
synaptotagmin 5, ARABIDOPSIS THALIANA SYNAPTOTAGMIN HOMOLOG E, Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
AT5G37740 49 / 2e-07
Calcium-dependent lipid-binding (CaLB domain) family protein (.1), Calcium-dependent lipid-binding (CaLB domain) family protein (.2)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.010G203000
213 / 5e-72
AT3G55470 167 / 6e-54
Calcium-dependent lipid-binding (CaLB domain) family protein (.1), Calcium-dependent lipid-binding (CaLB domain) family protein (.2)
Potri.002G155600
112 / 3e-32
AT1G63220 202 / 1e-67
Calcium-dependent lipid-binding (CaLB domain) family protein (.1), Calcium-dependent lipid-binding (CaLB domain) family protein (.2)
Potri.014G084100
78 / 3e-18
AT4G00467 194 / 4e-63
Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
Potri.001G372000
60 / 5e-11
AT4G21160 506 / 0.0
ARF-GAP domain 12, Calcium-dependent ARF-type GTPase activating protein family (.1.2.3.4)
Potri.017G125500
56 / 3e-10
AT3G17980 249 / 6e-86
Arabidopsis thaliana C2 domain, Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
Potri.004G089500
55 / 8e-10
AT3G17980 245 / 4e-84
Arabidopsis thaliana C2 domain, Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
Potri.008G131500
54 / 2e-09
AT3G17980 202 / 5e-67
Arabidopsis thaliana C2 domain, Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
Potri.011G098500
54 / 7e-09
AT4G21160 506 / 0.0
ARF-GAP domain 12, Calcium-dependent ARF-type GTPase activating protein family (.1.2.3.4)
Potri.016G005300
52 / 1e-08
AT5G47710 264 / 1e-91
Calcium-dependent lipid-binding (CaLB domain) family protein (.1), Calcium-dependent lipid-binding (CaLB domain) family protein (.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10040291
178 / 6e-58
AT3G55470 191 / 4e-63
Calcium-dependent lipid-binding (CaLB domain) family protein (.1), Calcium-dependent lipid-binding (CaLB domain) family protein (.2)
Lus10023410
136 / 1e-41
AT3G55470 149 / 5e-47
Calcium-dependent lipid-binding (CaLB domain) family protein (.1), Calcium-dependent lipid-binding (CaLB domain) family protein (.2)
Lus10029565
100 / 3e-27
AT1G63220 221 / 1e-75
Calcium-dependent lipid-binding (CaLB domain) family protein (.1), Calcium-dependent lipid-binding (CaLB domain) family protein (.2)
Lus10002235
100 / 4e-27
AT1G63220 216 / 2e-73
Calcium-dependent lipid-binding (CaLB domain) family protein (.1), Calcium-dependent lipid-binding (CaLB domain) family protein (.2)
Lus10014780
78 / 6e-18
AT4G00467 168 / 1e-52
Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
Lus10039113
55 / 9e-10
AT5G47710 239 / 6e-82
Calcium-dependent lipid-binding (CaLB domain) family protein (.1), Calcium-dependent lipid-binding (CaLB domain) family protein (.2)
Lus10038745
54 / 2e-09
AT5G47710 241 / 1e-82
Calcium-dependent lipid-binding (CaLB domain) family protein (.1), Calcium-dependent lipid-binding (CaLB domain) family protein (.2)
Lus10023265
53 / 1e-08
AT4G21160 495 / 3e-177
ARF-GAP domain 12, Calcium-dependent ARF-type GTPase activating protein family (.1.2.3.4)
Lus10038539
53 / 1e-08
AT4G21160 491 / 4e-176
ARF-GAP domain 12, Calcium-dependent ARF-type GTPase activating protein family (.1.2.3.4)
Lus10016347
53 / 2e-08
AT3G07940 478 / 8e-164
Calcium-dependent ARF-type GTPase activating protein family (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0154
C2
PF00168
C2
C2 domain
Representative CDS sequence
>Potri.010G203100.1 pacid=42798779 polypeptide=Potri.010G203100.1.p locus=Potri.010G203100 ID=Potri.010G203100.1.v4.1 annot-version=v4.1
ATGGCCGTTGGAATATTGGAAGTGAAGCTGGTGAAGGCAAAAGGCCTTGGAAACCCTGATTTCTTTGGTTTATCATGTTACTGCTCCAGCTCTTTGACTA
ACATGGACCCGTATGTTCTGGTAAAATACAAAAGCCAAGAGCGCAAGAGCAAAGTAGCAAGAGGTCAAGGTGGGCGTCCAGTGTGGAATGAGACTCTCAC
ATTCAAGGTAGAGTATCCAGGGCAAGGTGGCAACTATAAGCTCATTCTTAAAATCATGGACAAGGACACCTTCTCTGCTGATGATTCTGTCGGAGAAGCC
ACGATTTACGTTAAAGATTTGTTGGCATTAGGAGTGGAGAAAGGAACTGCTGAACTACAAACTCAGAAGTATAGAGTGGTTAATGCTGACAAATCTTATC
GTGGAGAGATTCAAGTTGGCGTCACTTTTACCCTCAAGGCAGAAGAGGAAGATGCTGGAGAAGATTATGGTGGATGGAGGCAAAGCTCTGTTTAG
AA sequence
>Potri.010G203100.1 pacid=42798779 polypeptide=Potri.010G203100.1.p locus=Potri.010G203100 ID=Potri.010G203100.1.v4.1 annot-version=v4.1
MAVGILEVKLVKAKGLGNPDFFGLSCYCSSSLTNMDPYVLVKYKSQERKSKVARGQGGRPVWNETLTFKVEYPGQGGNYKLILKIMDKDTFSADDSVGEA
TIYVKDLLALGVEKGTAELQTQKYRVVNADKSYRGEIQVGVTFTLKAEEEDAGEDYGGWRQSSV
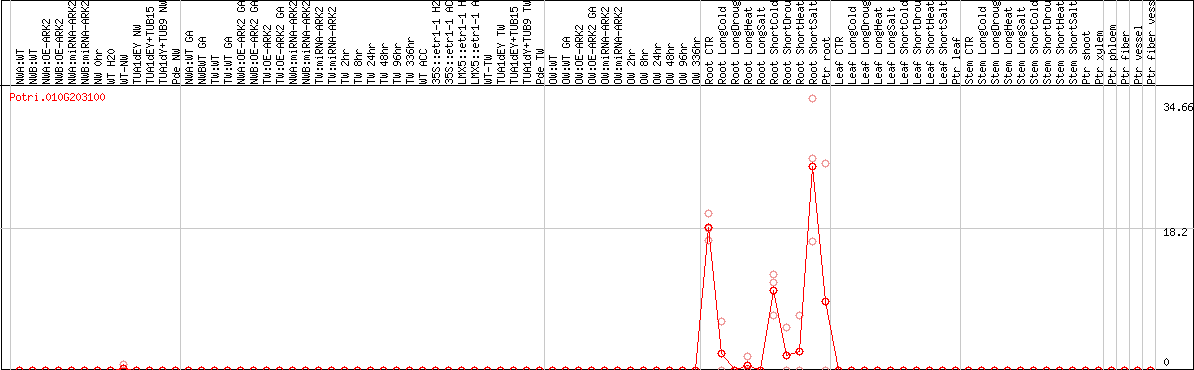
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.010G203100 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.