External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G39550 261 / 3e-86
GGB, ATGGT-IB, PGGT-I
GERANYLGERANYLTRANSFERASE-I BETA SUBUNIT, Prenyltransferase family protein (.1)
AT3G12070 72 / 1e-14
AtRGTB2
RAB geranylgeranyl transferase beta subunit 2 (.1.2)
AT5G40280 69 / 2e-13
ATFTB, WIG, ERA1
WIGGUM, ENHANCED RESPONSE TO ABA 1, ARABIDOPSIS THALIANA FARNESYL TRANSFERASE BETA SUBUNIT, Prenyltransferase family protein (.1)
AT5G12210 62 / 5e-11
AtRGTB1
RAB geranylgeranyl transferase beta subunit 1 (.1.2)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.008G054500
370 / 3e-129
AT2G39550 481 / 8e-171
GERANYLGERANYLTRANSFERASE-I BETA SUBUNIT, Prenyltransferase family protein (.1)
Potri.013G012500
75 / 1e-15
AT5G12210 525 / 0.0
RAB geranylgeranyl transferase beta subunit 1 (.1.2)
Potri.017G073200
64 / 2e-11
AT5G40280 511 / 5e-180
WIGGUM, ENHANCED RESPONSE TO ABA 1, ARABIDOPSIS THALIANA FARNESYL TRANSFERASE BETA SUBUNIT, Prenyltransferase family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10040303
270 / 4e-89
AT2G39550 436 / 4e-152
GERANYLGERANYLTRANSFERASE-I BETA SUBUNIT, Prenyltransferase family protein (.1)
Lus10023422
149 / 6e-44
AT2G39550 175 / 1e-52
GERANYLGERANYLTRANSFERASE-I BETA SUBUNIT, Prenyltransferase family protein (.1)
Lus10018502
80 / 3e-17
AT5G12210 510 / 0.0
RAB geranylgeranyl transferase beta subunit 1 (.1.2)
Lus10039714
79 / 4e-17
AT3G12070 509 / 0.0
RAB geranylgeranyl transferase beta subunit 2 (.1.2)
Lus10000944
66 / 9e-14
AT2G39550 76 / 2e-17
GERANYLGERANYLTRANSFERASE-I BETA SUBUNIT, Prenyltransferase family protein (.1)
Lus10032149
58 / 1e-09
AT5G40280 465 / 1e-161
WIGGUM, ENHANCED RESPONSE TO ABA 1, ARABIDOPSIS THALIANA FARNESYL TRANSFERASE BETA SUBUNIT, Prenyltransferase family protein (.1)
Lus10014536
45 / 4e-05
AT5G40280 422 / 1e-144
WIGGUM, ENHANCED RESPONSE TO ABA 1, ARABIDOPSIS THALIANA FARNESYL TRANSFERASE BETA SUBUNIT, Prenyltransferase family protein (.1)
Lus10017310
0 / 1
AT2G39550 86 / 4e-23
GERANYLGERANYLTRANSFERASE-I BETA SUBUNIT, Prenyltransferase family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0059
6_Hairpin
PF00432
Prenyltrans
Prenyltransferase and squalene oxidase repeat
Representative CDS sequence
>Potri.010G206001.8 pacid=42797093 polypeptide=Potri.010G206001.8.p locus=Potri.010G206001 ID=Potri.010G206001.8.v4.1 annot-version=v4.1
ATGACAAATGACACCAACCATCTCTCTCTTGCCTACTTCGTTATATCCGGCCTCGACATCCTCGGTTCCCTTGATCGCGTTGACAAGGATGCGGTTGCTG
CTTGGGTTTTATCTTTGCAAGCTAATTCTGGAGATAAAGCTGAATTGAAAAATGGACAGTTTTACGGGTTCCATGGTTCCCATTCTTCTCAATTCCGTTC
TGACAATAACGGGATTTTAATTCTCAATCACAGCCATTTGGCAAGCACTTATTGCGCACTATCCATATTGAAGACTGTTGGCTCTAATTTGTTAAATATC
GACTCCAAATTGATATCCATGTCAATAAGGAACCTTCAGCAACCTGATGGGAGTTTTCTGCCTATCCATATCGGTGCAGAGACAGATCTTCGATTCATAT
ACTGTGCATCGCCCATCTGTTATATGTTGGAGGACTGGAGTAGCATGGATAGGGTGAAAACCAAGGAGTGCATCTTAAAATGCCAGTCATATGGTGGGGG
CTTTGGGATGATTCCTGGTTCAGAATCTCATGGTGGTGGAACTTACTGTGCAGTTGCTTCTCTTCATCTGATGGGATTTATTGAACATGATGTCCAGTCA
AAAAGTGCAGCATCGTCTATCATTGACATTCCTTTGTATGGTGCTGACAGAGGCAAGCAGCTGAGGGTGGATTCCAAGGGAGAGCTAACAAGCCTAGTGA
CACATGCTATGCATTTTGGGTTGGGGCAGTTTTGA
AA sequence
>Potri.010G206001.8 pacid=42797093 polypeptide=Potri.010G206001.8.p locus=Potri.010G206001 ID=Potri.010G206001.8.v4.1 annot-version=v4.1
MTNDTNHLSLAYFVISGLDILGSLDRVDKDAVAAWVLSLQANSGDKAELKNGQFYGFHGSHSSQFRSDNNGILILNHSHLASTYCALSILKTVGSNLLNI
DSKLISMSIRNLQQPDGSFLPIHIGAETDLRFIYCASPICYMLEDWSSMDRVKTKECILKCQSYGGGFGMIPGSESHGGGTYCAVASLHLMGFIEHDVQS
KSAASSIIDIPLYGADRGKQLRVDSKGELTSLVTHAMHFGLGQF
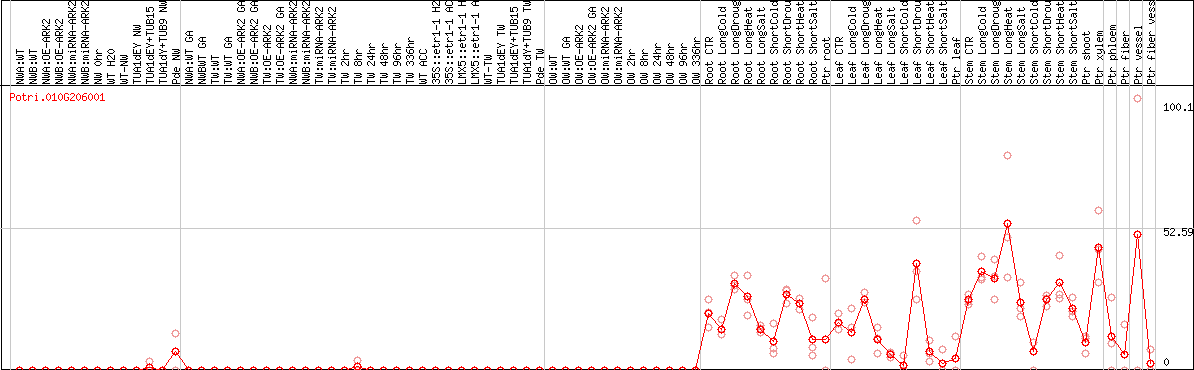
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.010G206001 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.