Potri.010G209300 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.010G209300.6 pacid=42799712 polypeptide=Potri.010G209300.6.p locus=Potri.010G209300 ID=Potri.010G209300.6.v4.1 annot-version=v4.1
ATGTCTTTCCAGCAGAAAAGCTTTTGGATGACCAGAGATGTGGGGTGTTTAACTGATGGAGATATAGGTTTCGATAATTCCTCAAGAATGGAACCAAAGC
GTGGTCATCAGTGGCTTATGGATAGCACTGGACCTGAATTGTTCAGTAACAAGAAACAAGCAGTTGAACCATCTAGCAACAACAGGCCAGTAATGGGAAT
GTCGCACATGAACATTTCTCCATGGAATAACACTTCCTGCTTCCAGTCGGTTTCAGGGCAATTTAATGACCGTCTGTTTGGGTTTGAACCATTGCGGATT
AACAGTGGTAGCAACGTGCCTTCTGCCAGCAATGGAAATATGAACATGGAAAGAAAGGATTTCAATGACCTATATGGCAGCAATTGCTCGATGGGTTTGT
CTATGTCTCATAACGTTGAAGATCCCCCTTCATCTTTTAGTTTTGGTGGCCTCAGAAAAGTGAGAGTCAATCAAGTTAGAGACTCAAGCAATGACATTTC
TTCATCTGTGGGGCACTCCTATAGTAGAGGAGATGACAATATAATTTCAATGGGTACTGCCTATAATAAGAGGGAGAGCAATGCCATCTCTTTGGGCTCT
ACTTATAACAATGGGGATGAGAATACCATCTCAATCAGCCCCACATTCAGCAAGGCAGATGGCAGTTTCATTTCAATGGGTCATGCATTTAACAAGGATG
ATGACAATTTCATATCAATGGGCCAGGGTTATAACAAGGGAGATGAGAGTATCTTATCAATGGGTCAGCCGTTTGACAAGAAGGATGCCAATTTCATAAC
AATGGGTCCATCTTATGACAAGGAAGATAACCATTTCATATCAATGGCTCTCTCCTACAATAAGGGACATGAGAGTTTCATTTCAATGGGACCCTCCTAT
GATAAAACGAGTGAAAATTTCATTTTGATGGGCTCCTCATTTAGCAAGGGAGGAGATAATGTCATATCAAACAGTCCAATATATGACAAGGCAGATATTG
ATATCGCATCCATGACTCCTGCCCAGGACAAGGGGAATTCTGGTATTCTATCCATTGGTCACAACTACAACAAGGGTGACAACAATTCCATTTCTTTTCA
AAGCTTTCATGATGAACCTGAGACAAATATGTCTGGCAATGTTATTAGAGGCTATGACCTGCTAGTGAGCAATCAGAACACTGCTCAAACTTCAGAAGTA
CCGGTTCAGAACAATTTGCCTCAGACAAATGTTGATCCACAATTGAATACCAATACTGCATCAAAGGTCATTGCAAATACACAATTAAATAGTGGTCTGG
AAGCTAATTCCAAAACTGATACTGACACCAAGGGTAAAGAGCCAAAGACATCAATCAATGCAGCCCCCAAGAATAAAGAGCCAAAGACAACAATTGATTC
TGCATCCAAGAATAAAGAGCTGAAGACATCTAAGAAGATCCCTCCTAACAACTTCCCCTCAAATGTCAAGAGCTTGCTATCAACTGGCTTGCTTGACGGG
GTTGCTGTGAAATATGTTTCCTGGTCGAGGGAGAAAACTCTTAGAGGAACTATAAAAGGGACTGGGTATTTATGTAGTTGCAAAGTATGCGGCAATAAGG
TTTTGAATGCATATGAATTTGAGCGACATGCTAACTGCAAGACCAAGCATCCCAATAACCATATTTACTTTGAGAACGGGAAGACCATCTATGCAGTTGT
TCAAGAACTGAAGAACACTCCTCAGGAGATGCTGTTCAATGCGATCGAAACTGTTACTGGATCTGCTATTAATCAGAAGAATTTCCTTAGCTGGAAAGCA
TCATATGAAGCTGCCACCCGTGAACTTCAGCGCATCTATGGAAAGGAGGAAGTCACTCTACCATCTTGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.010G209300.6 pacid=42799712 polypeptide=Potri.010G209300.6.p locus=Potri.010G209300 ID=Potri.010G209300.6.v4.1 annot-version=v4.1
MSFQQKSFWMTRDVGCLTDGDIGFDNSSRMEPKRGHQWLMDSTGPELFSNKKQAVEPSSNNRPVMGMSHMNISPWNNTSCFQSVSGQFNDRLFGFEPLRI
NSGSNVPSASNGNMNMERKDFNDLYGSNCSMGLSMSHNVEDPPSSFSFGGLRKVRVNQVRDSSNDISSSVGHSYSRGDDNIISMGTAYNKRESNAISLGS
TYNNGDENTISISPTFSKADGSFISMGHAFNKDDDNFISMGQGYNKGDESILSMGQPFDKKDANFITMGPSYDKEDNHFISMALSYNKGHESFISMGPSY
DKTSENFILMGSSFSKGGDNVISNSPIYDKADIDIASMTPAQDKGNSGILSIGHNYNKGDNNSISFQSFHDEPETNMSGNVIRGYDLLVSNQNTAQTSEV
PVQNNLPQTNVDPQLNTNTASKVIANTQLNSGLEANSKTDTDTKGKEPKTSINAAPKNKEPKTTIDSASKNKELKTSKKIPPNNFPSNVKSLLSTGLLDG
VAVKYVSWSREKTLRGTIKGTGYLCSCKVCGNKVLNAYEFERHANCKTKHPNNHIYFENGKTIYAVVQELKNTPQEMLFNAIETVTGSAINQKNFLSWKA
SYEAATRELQRIYGKEEVTLPS
|
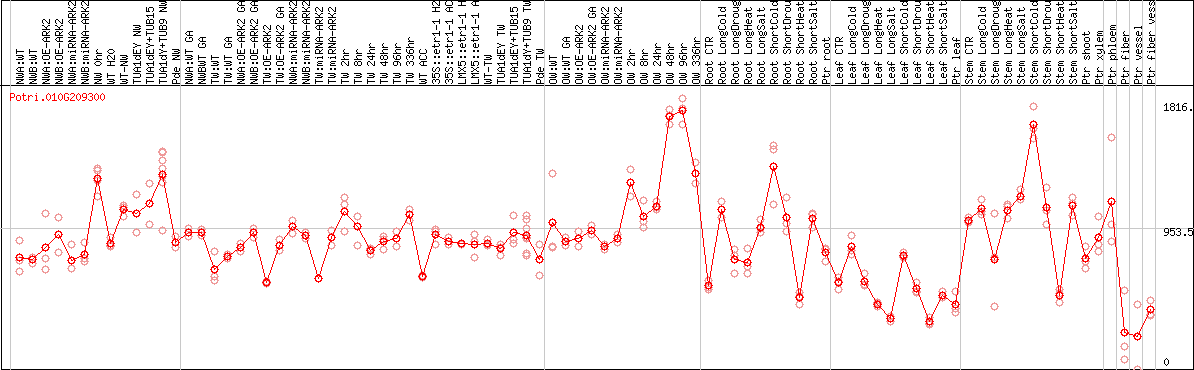
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.010G209300 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
