Potri.010G217900 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.010G217900.1 pacid=42797523 polypeptide=Potri.010G217900.1.p locus=Potri.010G217900 ID=Potri.010G217900.1.v4.1 annot-version=v4.1
ATGTTGTCTACAAATGATCCAATGGAGTCCTTTATGAATTCAATTCAAGTGGTCAAAGATGCACTTTCACCACTTGAACTCGGGATTCGGAAAGCTGCTA
AAGATCTTGAAACTTGTTGGGGGGTTTCAAAGAATGATGATAAGGCAACCCATGATCTAGTTACTGATAATAGCAGCAAAGTTTCTATCTTTACTGTGAA
AAATAAGAGTGTTAGTCTTGGAAATAGTGAGAATAGGCAGGGTGTGGTTAATGAAGAGAAAAAGAAAGGATTTTTATCTATTAAGTTTCCTATTAGGAGT
TTGTTGGGGATGTTTTCTATGAATTTGGAAGGTGGGCATAGAAATGGAGGTGATAATAAGGCTGGGCTTCCGAAGAAAGTCTTGAAAGAGAAGGAAATGA
GTAATGAAGACGGGTCTTGTGTAAATTGTTTGCGGTTTGCCATGACTTTGTCTTTGTTGGTTAATGGTTTGGTTCAGGCATTTCCTGGCCCATTTAAGAT
GAACAAGAAACGGTTTCAGAAAGTAGGTGACGAGGATAAAGATTATTTGCATTCGTCCAAAAATGGGTCGAAAGCTAAGGTTTCAGGTGAAATGAAGCTA
AGGAAATCAAAGGGTCAATCTGTTAAAGGGTACCAGAATGTGAGTGAGAAGGGTAAGGAGGAGAAACCTGTGTCCCTCGAATGCTTTATAGGGTTTCTAT
TTGACCAATTGGCTCAGAATCTACAGAAGTTTGATCTGGGTTTACAGGAAAGAGATATCAAGGGTTGTGAAAATGATTGTTCAACCTCACCTCCAGCATA
CTCCCAGTTTGATCATTTGAGGGCAATCATAAGTATTTGGGAAGGTCAAAAGGTGTATGTAGATGGGGTTTTGGGAAATTTGAGTTTTGCGAGAGTGGGT
GGTGTGCCATCAAGTATAGTTGGAGTATCCTCTTCAGTTAATGAAGAGGGTGATGATGGTGCAAGTAGTGCACCTACCAACAGCGCTGAGGACACAGGGA
GTAGTTCACCACAGAATCTAGCAAGTGGGTTACTGAGCATTCCGCTATCAAATGTAGAGCGTTTGAGATCCACATTATCTACTGTTTCATTGACGGAGCT
AATTGAGCTTGTGCCACAACTTGGTCGATCTTCTAAAGACTATCCTGACAAGAAAAAGCTATTCTCTGTCCAGGATTTCTTTAGATACACAGAGGCTGAA
GGAAGAAGATTCTTTGAGGAACTTGACAGAGATGGTGATGGTCAAGTGAATTTGGAAGATCTTGAAATCGCATTGAGAAAGAGAAAATTGCCCCAGAGAT
ATGCCCGGGAATTCATGCGGCGTGCTAGAAGTCACTTGTTCTCAAAATCATTTGGGTGGAAACAATTTTTGTCCTTGATGGAACAGAAGGAACCAACAAT
TCTGCGTGCTTACACTTCTCTGTGTTTAAGCAAGTCTGGGACCCTGCAAAAGAGTGAAATATTGGCATCACTAAAAAATTCTGGACTTCCAGTGAATGAA
GACAATGCTGTTGCTATGATGCGATTTCTGAATGCTGATACAGAAGAATCTATTTCTTATGGGCATTTTCGTAATTTTATGCTTTTGCTGCCTTCTGATC
GACTTCAAGATGATCCCAGGAATATATGGTTTGAAGCAGCAACTGTTGTTGCCGTTGCACCACCCGTGGAAATACCTGCTGGAAGCGTTTTACGATCTGC
ATTGGCCGGGGGTCTTTCATGTGCACTCTCTTGTTCTCTTATGCACCCTGTTGATACAATAAAGACTCGGGTGCAAGCTTCAACATTGGCCTTCCCAGAA
ATAATTTCAAAGCTTCCACAAGTTGGAGTTCGAGGATTGTATAGGGGTTCAATTCCTGCAATTTGGGGACAGTTTACAAGCCACGGCTTGCGAACTGGGA
TATTTGAAGCTACCAAACTTGTGCTAATAAATGTTGCCCCAACACTCCCAGATATCCAGGTTCAATCTGTAGCATCATTATGCAGCACAGTTTTGGGGAC
AGCAGTGCGGATCCCTTGTGAGGTATTGAAGCAGCGTTTGCAGGCTGGCCTTTTTGACAATGTAGGACAGGCTATTGTTGGTACATGGCAGCAAGATGGC
CTAAATGGGTTCTTTCGTGGGACTGGTGCCACTCTTCTCCGTGAGGTTCCATTTTATGTTGCTGGCATGTGTCTATATGGTGAATCCAAGAAGGTTGCTC
AACAACTTTTAAGGCGGGAGCTGGAACCCTGGGAAACAATTGCTGTTGGGGCTTTATCTGGAGGCCTAACTGCTGTTATCACTACCCCTTTCGATGTACT
GAAAACTAGAATGATGACGGCGCCACCAGGAAGAACTGTATCAATGTCATTGATAGCTTTCTCTATACTCCGACATGAGGGGCCCCTTGGCTTATTCAAA
GGAGCAGTGCCGAGGTTCTTCTGGATTGCTCCCTTAGGTGCCATGAACTTTGCAGGCTATGAGCTAGCAAGGAAGGCCATGGACAAAAACGAGGAGGCCA
CAAAAGTTGAGTAG
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.010G217900.1 pacid=42797523 polypeptide=Potri.010G217900.1.p locus=Potri.010G217900 ID=Potri.010G217900.1.v4.1 annot-version=v4.1
MLSTNDPMESFMNSIQVVKDALSPLELGIRKAAKDLETCWGVSKNDDKATHDLVTDNSSKVSIFTVKNKSVSLGNSENRQGVVNEEKKKGFLSIKFPIRS
LLGMFSMNLEGGHRNGGDNKAGLPKKVLKEKEMSNEDGSCVNCLRFAMTLSLLVNGLVQAFPGPFKMNKKRFQKVGDEDKDYLHSSKNGSKAKVSGEMKL
RKSKGQSVKGYQNVSEKGKEEKPVSLECFIGFLFDQLAQNLQKFDLGLQERDIKGCENDCSTSPPAYSQFDHLRAIISIWEGQKVYVDGVLGNLSFARVG
GVPSSIVGVSSSVNEEGDDGASSAPTNSAEDTGSSSPQNLASGLLSIPLSNVERLRSTLSTVSLTELIELVPQLGRSSKDYPDKKKLFSVQDFFRYTEAE
GRRFFEELDRDGDGQVNLEDLEIALRKRKLPQRYAREFMRRARSHLFSKSFGWKQFLSLMEQKEPTILRAYTSLCLSKSGTLQKSEILASLKNSGLPVNE
DNAVAMMRFLNADTEESISYGHFRNFMLLLPSDRLQDDPRNIWFEAATVVAVAPPVEIPAGSVLRSALAGGLSCALSCSLMHPVDTIKTRVQASTLAFPE
IISKLPQVGVRGLYRGSIPAIWGQFTSHGLRTGIFEATKLVLINVAPTLPDIQVQSVASLCSTVLGTAVRIPCEVLKQRLQAGLFDNVGQAIVGTWQQDG
LNGFFRGTGATLLREVPFYVAGMCLYGESKKVAQQLLRRELEPWETIAVGALSGGLTAVITTPFDVLKTRMMTAPPGRTVSMSLIAFSILRHEGPLGLFK
GAVPRFFWIAPLGAMNFAGYELARKAMDKNEEATKVE
|
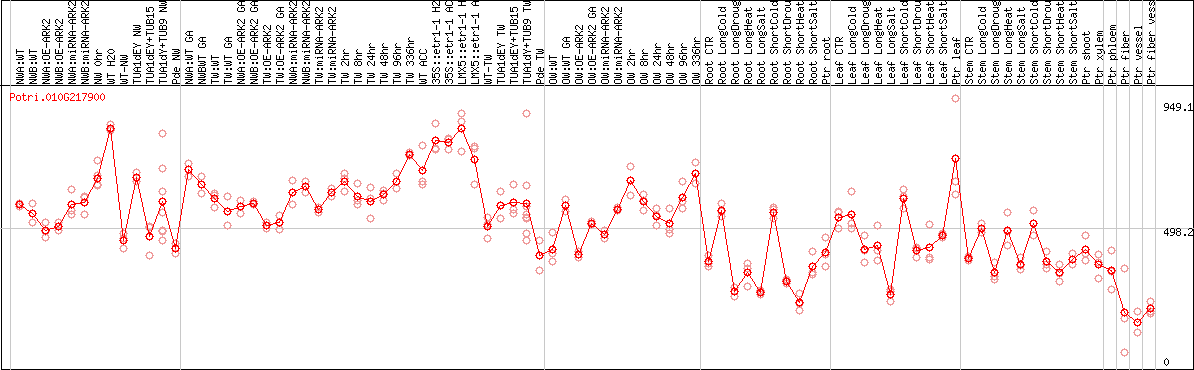
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.010G217900 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
