PABAS (Potri.010G221500) [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | PABAS | |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.010G221500.5 pacid=42797520 polypeptide=Potri.010G221500.5.p locus=Potri.010G221500 ID=Potri.010G221500.5.v4.1 annot-version=v4.1
ATGGCTCTGTGTTCGTCGACTTCGTTGGAACTGAAAAATCCTTTTATTGAGGGCACGAAAGGGAAAATAGCGACATCCAAGTCTTTTCTAAGAGTTGGTT
ATGTTGCCAAGAATGAGAAATCTTGTTGCTGTAATGGTAGAAAGGTGGCGGTATCCAGTCATTTGATGCCCGGACATTTAGAAGGGTCGTTCATGGGAAA
GAAACGGCTGGAGGAGCCTAGCCAGAAAATGGATTTTGTGAGGACATTGTTGATTGACAACTATGATAGTTATACCTATAATATTTACCAAGAGCTATCT
GTCGTTAATGGGGTGCCTCCTGTGGTCATCCAAAATGATGAGTGGACATGGGAGGATGCGTGCCATTACTTGTATGAAAAGAGAGCTTTTGATAATATTG
TTATCTCACCTGGACCTGGTTCTCCAACATGCGCAGCGGATATAGGAATATGTCTTCGACTGCTGCTCGAGTGCAGGGACATTCCTATTCTGGGTGTTTG
CCTTGGACACCAGGCCTTGGGTTATGTCAATGGTGCCAGAATTGTCCATGCATCTGAACCAGTCCATGGACGCTTAAGTGAAATCGAACATAATGGAAGC
AGACTGTTTGATAACATCCCTTCTGGCAGAAAATCTGGTTTTAAGGTGGTACGGTATCATTCTCTCATCATAGATTCAGAAGCACTGCCAAAGGAACTTA
TACCGACTGCTTGGACTTCTAGCAGCACTCATTCTTTCCTTGAGTCTCCGAATTCTGGTCTTAACCTTGATGCGTGCAAGAATCAGATTAGGCCAAGCAC
ATCTTCTGATACTTTTTCAACAGGATCACACAATGGAGCTTCTTGGTCTTTTAGCCACCCTGGAAGAATGCAAGGGGGAAAAGTTCTCATGGGCATTATG
CATTCTACTAGGCCGCACTATGGCTTACAGTTTCATCCAGAAAGTATTGCAACCTGTCACGGAAGGCAAATATTTGAGAATTTTAGAGAGATTACAGAGG
ATTATTGGCAGAGACTGAGGCCAAGGTCAACATTTATTAACGAGAGAAATGTTCATTATACTGCATGCATGCAGGTGCATGTTGCTAGTCAGCTGTTTAG
AGTTCCTAGAATTGGATCATTAGTACACAAGGAAGATGCTCAACCATTCAAAGAAGCTTTTAGAAGAAGCCAATTACTTGGAAATGCCAACGTGAATTGT
CTCAGCATTTCCAGTGCATTGAAGTTTCCAGAATCAAGCATCAACGTTAGGCATTTAAAGTTGAAATGGAGGAAATTTGATAAATTAGCAGCCCGAGTTG
GGGGAGCAAGAAACATATTTAACGAACTGTTTGGGGTTTGTAAGGCTGAAAATACGTTTTGGCTGGACAGTTCCTCAGTAGAAAAGAAAAGAGCACGCTT
TTCATTTATGGGGGGTAAAGACGGACCACTTTGGAGGCAAATGACATTTAGATTGTCAGATCAAAGTGATATGGATTTTAAAGGTGGAGGTTACTTATCA
ATCAAAGATACTCAAGGATCCACCGAAAGCATGTTTTTGGAAAAAGGTTTTCTTGATTTTTTGAACCAGGAGCTACTATCATTTACTTACGATGAAGAAG
ATTTTGAAGAACTGCCATTTGACTTTCATGGTGGTTATATTGGTTACTTCGGGTATAGCCTCAAAGTTGAATGTGGCATGTTATCCAACCGTCACAAATC
TACGACTCCAGATGCTTGCTTTTTCTTTGCAGATAATTTTGTAGTCATCGATCATCTTAATGACAATGTTTATATATTGTCTTTACATGAGGAAAGCACA
ACCAGCATTCCATGGTTGGATGACACGGAAAATAAGTTGCTTTGCTTAGAAGCTTCTACCACAAGGAAATTAGGGGAGCAGGCTTCCCCTACTGCAACAG
TTTCCCCATACAAGGCAGGTTTTCTTGGTGAGAAATCTAGAGAGCAGTATATTAAAGATGTTAGTAAGTGTCTAGAATACATTAAAGATGGGGAAAGCTA
CGAGTTATGTCTCACTTCTCAGATGAGAAAAACAGTTGGAGAGATAGATTCCTTGGGACTTTACCTTCACCTCAGAGAGAAAAATCCAGCACCATATGCT
GCGTGGCTTAATTTTTCAAATGAAGATTTGTGCATCTGCTGTTCTTCTCCTGAGAGATTCCTATGCTTGGACAGAAATGGTATATTAGAAGCAAAGCCGA
TTAAAGGTACCATAGCTCGTGGGGTGACCCTGGAGGAAGATGAAGAGCTTAAATTGAAACTGCAGTACAGTGAAAAGGATCAAGCTGAAAATCTTATGAT
TGTCGATCTTTTGAGGAACGATCTTGGTCGTGTCTGTGAGCCTGGTTCTGTTCATGTCCCACATCTTATGGAAGTAGAGTCGTATGCAACTGTGCATACT
ATGGTGAGTACAATTCGAGGAAAGAAGAGGTCAAATGTAAGTGCAGTTGACTGTGTCAGGGCTGCATTTCCTGGTGGCTCAATGACAGGTGCACCAAAAT
TGAGATCAATGGAGCTTCTCGATTCTCTTGAAAGCAGTTCTCGTGGGATCTACTCAGGTTCCATTGGATTTTTCTCATATAATCAGACATTTGATCTGAA
TATAGTGATAAGAACAATAGTTATTCATGACGGTGAAGCTTCTATAGGTGCAGGAGGAGCTATTGTTGCTCTATCAAATCCTGAAGACGAATACGATGAA
ATGTTACTGAAAACACGAGCACCAGCAAGCGCAGTGATAGAATTTCAGTAG
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.010G221500.5 pacid=42797520 polypeptide=Potri.010G221500.5.p locus=Potri.010G221500 ID=Potri.010G221500.5.v4.1 annot-version=v4.1
MALCSSTSLELKNPFIEGTKGKIATSKSFLRVGYVAKNEKSCCCNGRKVAVSSHLMPGHLEGSFMGKKRLEEPSQKMDFVRTLLIDNYDSYTYNIYQELS
VVNGVPPVVIQNDEWTWEDACHYLYEKRAFDNIVISPGPGSPTCAADIGICLRLLLECRDIPILGVCLGHQALGYVNGARIVHASEPVHGRLSEIEHNGS
RLFDNIPSGRKSGFKVVRYHSLIIDSEALPKELIPTAWTSSSTHSFLESPNSGLNLDACKNQIRPSTSSDTFSTGSHNGASWSFSHPGRMQGGKVLMGIM
HSTRPHYGLQFHPESIATCHGRQIFENFREITEDYWQRLRPRSTFINERNVHYTACMQVHVASQLFRVPRIGSLVHKEDAQPFKEAFRRSQLLGNANVNC
LSISSALKFPESSINVRHLKLKWRKFDKLAARVGGARNIFNELFGVCKAENTFWLDSSSVEKKRARFSFMGGKDGPLWRQMTFRLSDQSDMDFKGGGYLS
IKDTQGSTESMFLEKGFLDFLNQELLSFTYDEEDFEELPFDFHGGYIGYFGYSLKVECGMLSNRHKSTTPDACFFFADNFVVIDHLNDNVYILSLHEEST
TSIPWLDDTENKLLCLEASTTRKLGEQASPTATVSPYKAGFLGEKSREQYIKDVSKCLEYIKDGESYELCLTSQMRKTVGEIDSLGLYLHLREKNPAPYA
AWLNFSNEDLCICCSSPERFLCLDRNGILEAKPIKGTIARGVTLEEDEELKLKLQYSEKDQAENLMIVDLLRNDLGRVCEPGSVHVPHLMEVESYATVHT
MVSTIRGKKRSNVSAVDCVRAAFPGGSMTGAPKLRSMELLDSLESSSRGIYSGSIGFFSYNQTFDLNIVIRTIVIHDGEASIGAGGAIVALSNPEDEYDE
MLLKTRAPASAVIEFQ
|
DESeq2's median of ratios [POPLAR]
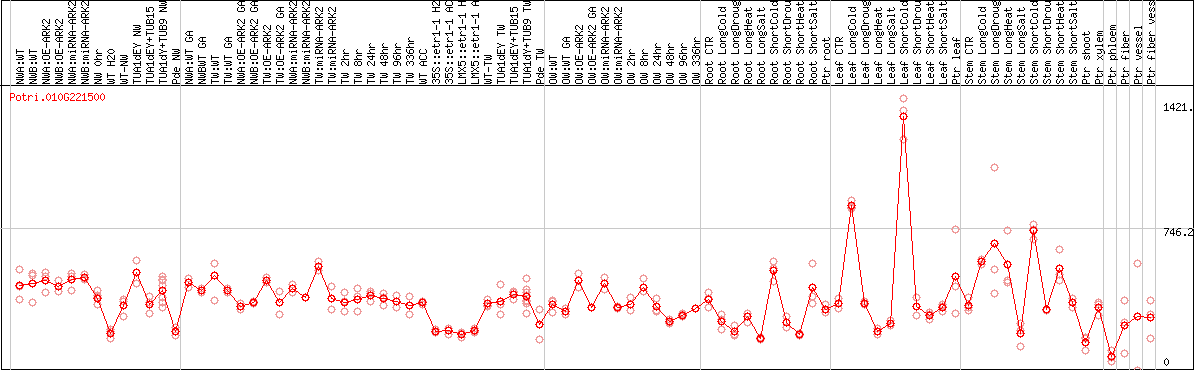
Coexpressed genes
Potri.010G221500 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
