Potri.010G234400 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.010G234400.4 pacid=42799829 polypeptide=Potri.010G234400.4.p locus=Potri.010G234400 ID=Potri.010G234400.4.v4.1 annot-version=v4.1
ATGGGTTCAAGGATTAATTATGGAGAAAGATTGATTCTGAGTTCGTTCTTACCAGCAACGCCAGAGAAGCCCATTCCAACAAGATCCAATTTGATGCCAG
CTGACGGGCAAGGGAACCTACAGGGCAGAACGAATTGGCAGCTGGCTGGATTTGCCGATGGATGTGTACAAGATGTGTCGAATTACAGCAGAGCGGCCCA
GCATACTGACCAGATTGATCAACTGTTTAGAAATGGCGGGGATAATATTAGGAATGCAGGTTTCGAAGAGGTGAAAAGAATGAGCAATACTACACGCAAT
TATATAGAGAGTCAAACTGGAGGTCTGGGCAGTGGTTTATTAGCAAATATTTTGGACCATTCAAATATATCTACAATTACCTCTTCAATGGCACCTCCTG
GTATAAGCACAAGCATGGTTTGGAGGCCTTCGTTTCCCAACTTGCATCCCCAAGTCAACAATTACAGAGAACCCAATTTGTTGCTCGGAAACCAAACTCA
CTGCTCAGGCTTACGGCATTTGGGTAGCAACTATATCTCATCCCAGGAACCCAATTATGAACCTATGATGCCTTGCCCTCATAACTATGATCTAAACTTT
CCACCAAGAATGGAAGCTGATGCAGCCTCATATTTTACCACCTCTTTTAAATTGGCCACGGTAGTACCGGATCAGTGCAAGAGATTGGAGAGTAGGCTTT
CTGCAACAGCAAGTCCCTCGCAAGAAAAAAACTCAAGTGGGGAAAAGGAGAAAACTGATTTAGTCATATTCAAAGAGTGCGAAGCCAATCAACATAATTC
AAAGGAGCTCTCATGCAACATTACAGACGCACCATCTGCTGTCATATCTACACCATTTGAGGAAGCAAAAGATCTAGCAACAGCAAATGCACAAGGCATC
GACCTTAATAGAACACCACAGCAGAAACCTCAAAAGAGAAGAAAGCATAGGCCCAAGGTGATTGTGGAAGGCAAGCCAAAAAGGACTCCCAAGGCTGCAA
CCACAAAAATTACTGATCCCAAGGAGAAGCCAATTGAAAAGAGGAAGTATGTTCGCAAGGCTTTGAAAGAGCCAGCAACTAAACCTACTGAATCTACCGT
AGATACTGCACCTCCCAGCTCTGCAAAGAGGAAGTATGTGCGCAAGAAGGCCCTGGATGAATCAGCAGTTCAACACACAGATTCTATTGGAGAGACCATA
AATACCCATGCTGTAAAGAGAAAGTATGTGCGCAAGAAAGACCTAAACAAATCAGCAAATCGACATGCAGACTCCACCGTGGAGATTACACAATCCAGTT
CAGCTGATGCAAAATCATGTAGAAGAGCATTGAGGTTTGATTTGGAAACTGCAACAGACAGGAGCTGCAGTAATGCAGCTGCTCAGCAGGACATGCTGAA
TCAAAAAAGAGGGACTTTTGATCTGAATGCAAGTCTTCAAGTGGCAGATTTAAGCACCACAACTTCACAGATGAGCCAACAGCATAGGTTACTGGTAGAA
AACCAGCAATCAGGTGCTCCAAGTAACCAGACACCTTTTATGAATCAGCCACGTGGTGACTATATATCAATTTCAGAAATACAAGTAGTGGCAGCGGAAC
TAACTCCAAGAAAAAACATGCACATGGAGAAATTAAACCTCAATGCGGGGGATGTGGAGAGAAGTATCCATGCACAAGGAATTGGTCAGGTTGTGTTTCC
AGAAAAAGGACCAGAGTGGACTAGACAAATAACTTCTCAGAATAACTCTCAATCAGCACAAAAGATAACACCTTATTTAATTGAGGGCAGAGGATTCAAG
AGAGAACACTTTCATATTAAGAAAACAAACCCGTGTACAGCATATCCAGTGGGCTCATTGACTGATGGATATGACCAAAATGGCAGCATTCCAGGTTCTG
GATGTTCAGAGACCCAGAAAAGAAAGAAAACAGAGGATGGAATTCAAACAAACACACACAGCATATCTTCTTTTGTTTCAAAAGTTAAATATCCTGGTGA
GTGGTATGTTCACTCTATGGCTCTGCAGAACCTTCCAAAGCAATGTATTTCACCTCAACCACATTTATGCCTAGAAATGCTGGGAGAGACTAATGGATCG
ACTCAGGTCCAAAACTCCCTCTGCCCCACCACTATAGAAACTTCTCACAGGCTTTCACAAACGTCTCTCAAAACAAGCCGTGCATCGGATAACCAACTGC
AACCCAAAACTTGCAACGCTGAAATGTCAAGAATTCAGCAAATGTCCGAAGCTACTGTGCCCATATCAATTCCATCTGAAAAGGGTAAGATTCCACAAGA
ACCAAAGGATGACTTGAAAGTTCATCAGCAGCCATATGCAAAGAGAAGAGGGCGACCAGCAAAGCAGACTTTCTCCTCTACAATAGAACAAATCATATAC
CAGATGGAGGGCCTCCGTCTCAATGCAGGGAGCAAGAAAATAGAAAACAAAGAGCAAAATGCACTTGTTCCTTACAAAGGAGATGGCAAACTTGTTCCCT
ATGATGGGTTTGAAGTTGTCAAGAAACACAAGCCAAGGCCTAAAGTAGATCTTGACCCAGAGTCAGACAGAGTATGGAAACTGCTGATGGGGAAAGAAGG
AAGTCAAGGCCTTGAAGGAACTGACAAGGGAAAGGAGCAATGGTGGGGGGAGGAGAGAAAAGTTTTTCATGGGCGAGTTGATTCATTCATTGCAAGAATG
CATCTCGTGCAAGGAGATAGGCGCTTTTCAAAATGGAAGGGATCAGTTGTTGATTCAGTAATAGGAGTTTTCCTAACTCAAAATGTTTCAGATCATCTGT
CAAGCTCTGCCTTCATGTCTCTAGCTTCATTGTTTCCTCTGAAGTTGAGGAGCAGTGGAGCCTGTGACAGAGAAAGGACAAGCATCGTCATTGAAGAACC
AGATACCTGCATTCTAAACCCAAATGATATCAAATGGAACAGCAATCCACTCTACAATCAAAGCTCCGTGACACACCATGGATCAGCAGAACCTCATAAA
GACTCTGAAACCTTGTTTATAGAAAGAGCAAGCATGGTAGAGACACAAAGTCACAGCCTGGAGGAAGAATTTGTATTATCACAAGATTCTTTCGATTCCT
CAACTGTCCAAGCAAATGGAGTCAGATCCTACTCGGGCTCCAACTCTGAAGCAGAAGATCCTGCTACTGGGTGCAAACCTAGCATGAATGATGATTTATC
TTTCATGGATCTTTTACAGATGGAGAGTCCCACCTTGCTAGGAGAATTTTATGGTTGTGAGGGTGGGAGCTCACTGTTTCATAAGGAATCCAGGCATGAA
AAGGAACAAGCAGAAGATTTACAAAACAGGCAGCCAGGACCTGGATTGGAAAGATTAGGTAATCTGAACTGCTTTTCTACATATAATCAGCATTTCGATT
ATTGCAATCCACAGATGCTAGGGAAAGTGGTTCCTTGCAGTGACTACGGGTTGCTACACATGACTTCACAATCCAATGTACAGCAAGCAGAAGGCTTCGA
ATTGTACAGTGAAGAAAACATATCTTCTTGGCTGTCATATTCTTCCAGATTCGACAAAGAAAAGGCTGCAACCTGCACAAGCAAAGCTGTTGGGCAAGAG
GCAGAAAGTGTAGGTAAAACTGCTGCCAAGCAATATGAGTTGCCAAGGTATGGACAAAGCAGCTCTCAGTCTTGCCATGAAAGACAAGTTGATGAGAGGA
ACAAGACCTTGCAATGGCAAAGCATGTCAGTTGGCGGACCTGTAAACCTTGCTGAAGAACTGCCTAAGAAGCAAAACAGTTACAGACAGCAAGTCTCAAG
CCTCACAGGAAACATTTTTGACGTTGAGAGAATCACTTCAGTGAATAAGCAAACCCCTCTAGAAAACAATGTGGTTGACCCAAATACAAAGGAGAAAGTA
CATCATAATAACAGGGAAAACCTCAAGGAAAATGCAAGTACCTCAAAAGCAAGAAAAGGAAAAGTGGAGGGTGAGAAAAAGGATGCATTTGACTGGGATA
GTTTAAGGAAGCAAGTGCAGGCAAATGGAAGAAAAGAGAGAGCCAAAGACACAATGGATTCCTTGGACTATGAAGCAGTGCGAAGTGCTAGGGTCAAGGA
GATTTCTGATGCGATAAAAGAAAGAGGGATGAATAACATGCTAGCAGAACGAATCCAGGAATTCCTCAACCGACTGGTAAGGGAACACGGAAGCATCGAT
TTGGAATGGTTAAGAGATGTGCCCCCTGACAAAGCAAAGGATTATCTGCTAAGTATAAGGGGGTTGGGGTTGAAAAGCGTGGAGTGTGTGAGGCTCCTGA
CACTTCATCATCTTGCCTTTCCAGTTGACACAAATGTCGGGAGAATCGCAGTTCGACTAGGATGGGTCCCTCTCCAGCCATTGCCAGAGTCACTTCAGTT
GCACCTCCTAGAACTGTACCCAATACTCGAGTCAATTCAGAAATATCTCTGGCCAAGATTGTGCAAACTAGATCAAAGAACATTGTATGAATTGCATTAC
CAGATGATCACATTTGGAAAGGTTTTTTGCACAAAGAGTAGACCCAACTGTAATGCATGTCCCATGAGAGCAGAATGTAGACATTTTGCAAGTGCCTTTG
CAAGTGCTAGGCTTGCATTACCTGGACCAGAAGAGAAGGGTATAACAACTTCAACTGTTCCCTTTATGCCTGAAAGAAGCCCCGGCATAGGTATCAATCC
CATGCCATTACCTCCACCCGAGGACAACCCACATAAAAGACACGGGTCTGATATTGGCAGTTGCGTGCCCATTATTGAAGAGCCAGCAACACCAGATCAA
GAGAACACTGAATTAACAGAGACTGATATTGAGGATTTTGGAGAAGATCCTGATGAAATACCCACAATAAAACTGAACATGGAAGAGTTCACTGAGAATT
TACAGAATTACATGCATACAAACTTGGAGCTCCAAGAAGGCGACATGTCCAAGGCCTTAGTTGCTTTGAATCCAAATGCTTCTATACCCACACCCAAATT
GAAGAATGTTAGTCGTCTACGGACAGAACACCAAGTGTATGAACTTCCAGATTCACATCCTCTCCTTGAGGGGATGGATAGAAGAGAACCTGATGATCCT
AGTCCATACCTTCTAGCAATATGGACACCAGGTGAAACTGCAAATTCTATTGAGCCACCAGACCAACAATGCCAGTCCAGAGAACCAAACAAATTGTGCG
ATGAGAAGACATGTTTTTCATGCAATAGTATCAGAGAAGCTAATTCACAAACAGTAAGAGGAACCCTTCTGATACCTTGTAGAACAGCAATGAGAGGAAG
CTTCCCACTGAATGGAACGTACTTTCAAGTGAACGAGATGTTTGCAGATCACGAATCTAGCTTAAACCCAATTGATGTTCCAAGATCATTGATATGGAAT
TTGCCGAGACGAATTGTCTACTTTGGAACCTCTGTATCATCAATATTTAAAGGACTTTCAACAGAAGGAATTCAGTTCTGCTTTTGGAGAGGGTTTGTTT
GTGTGAGGGGGTTCGACCAGAAAACAAGAGCACCCCGTCCTCTTAAGGCCAGATTACACTTTCCTGCAAGTCGGTTGGTAAAGACAAAGAACGAGAAAAA
ATGA
|
|||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.010G234400.4 pacid=42799829 polypeptide=Potri.010G234400.4.p locus=Potri.010G234400 ID=Potri.010G234400.4.v4.1 annot-version=v4.1
MGSRINYGERLILSSFLPATPEKPIPTRSNLMPADGQGNLQGRTNWQLAGFADGCVQDVSNYSRAAQHTDQIDQLFRNGGDNIRNAGFEEVKRMSNTTRN
YIESQTGGLGSGLLANILDHSNISTITSSMAPPGISTSMVWRPSFPNLHPQVNNYREPNLLLGNQTHCSGLRHLGSNYISSQEPNYEPMMPCPHNYDLNF
PPRMEADAASYFTTSFKLATVVPDQCKRLESRLSATASPSQEKNSSGEKEKTDLVIFKECEANQHNSKELSCNITDAPSAVISTPFEEAKDLATANAQGI
DLNRTPQQKPQKRRKHRPKVIVEGKPKRTPKAATTKITDPKEKPIEKRKYVRKALKEPATKPTESTVDTAPPSSAKRKYVRKKALDESAVQHTDSIGETI
NTHAVKRKYVRKKDLNKSANRHADSTVEITQSSSADAKSCRRALRFDLETATDRSCSNAAAQQDMLNQKRGTFDLNASLQVADLSTTTSQMSQQHRLLVE
NQQSGAPSNQTPFMNQPRGDYISISEIQVVAAELTPRKNMHMEKLNLNAGDVERSIHAQGIGQVVFPEKGPEWTRQITSQNNSQSAQKITPYLIEGRGFK
REHFHIKKTNPCTAYPVGSLTDGYDQNGSIPGSGCSETQKRKKTEDGIQTNTHSISSFVSKVKYPGEWYVHSMALQNLPKQCISPQPHLCLEMLGETNGS
TQVQNSLCPTTIETSHRLSQTSLKTSRASDNQLQPKTCNAEMSRIQQMSEATVPISIPSEKGKIPQEPKDDLKVHQQPYAKRRGRPAKQTFSSTIEQIIY
QMEGLRLNAGSKKIENKEQNALVPYKGDGKLVPYDGFEVVKKHKPRPKVDLDPESDRVWKLLMGKEGSQGLEGTDKGKEQWWGEERKVFHGRVDSFIARM
HLVQGDRRFSKWKGSVVDSVIGVFLTQNVSDHLSSSAFMSLASLFPLKLRSSGACDRERTSIVIEEPDTCILNPNDIKWNSNPLYNQSSVTHHGSAEPHK
DSETLFIERASMVETQSHSLEEEFVLSQDSFDSSTVQANGVRSYSGSNSEAEDPATGCKPSMNDDLSFMDLLQMESPTLLGEFYGCEGGSSLFHKESRHE
KEQAEDLQNRQPGPGLERLGNLNCFSTYNQHFDYCNPQMLGKVVPCSDYGLLHMTSQSNVQQAEGFELYSEENISSWLSYSSRFDKEKAATCTSKAVGQE
AESVGKTAAKQYELPRYGQSSSQSCHERQVDERNKTLQWQSMSVGGPVNLAEELPKKQNSYRQQVSSLTGNIFDVERITSVNKQTPLENNVVDPNTKEKV
HHNNRENLKENASTSKARKGKVEGEKKDAFDWDSLRKQVQANGRKERAKDTMDSLDYEAVRSARVKEISDAIKERGMNNMLAERIQEFLNRLVREHGSID
LEWLRDVPPDKAKDYLLSIRGLGLKSVECVRLLTLHHLAFPVDTNVGRIAVRLGWVPLQPLPESLQLHLLELYPILESIQKYLWPRLCKLDQRTLYELHY
QMITFGKVFCTKSRPNCNACPMRAECRHFASAFASARLALPGPEEKGITTSTVPFMPERSPGIGINPMPLPPPEDNPHKRHGSDIGSCVPIIEEPATPDQ
ENTELTETDIEDFGEDPDEIPTIKLNMEEFTENLQNYMHTNLELQEGDMSKALVALNPNASIPTPKLKNVSRLRTEHQVYELPDSHPLLEGMDRREPDDP
SPYLLAIWTPGETANSIEPPDQQCQSREPNKLCDEKTCFSCNSIREANSQTVRGTLLIPCRTAMRGSFPLNGTYFQVNEMFADHESSLNPIDVPRSLIWN
LPRRIVYFGTSVSSIFKGLSTEGIQFCFWRGFVCVRGFDQKTRAPRPLKARLHFPASRLVKTKNEKK
|
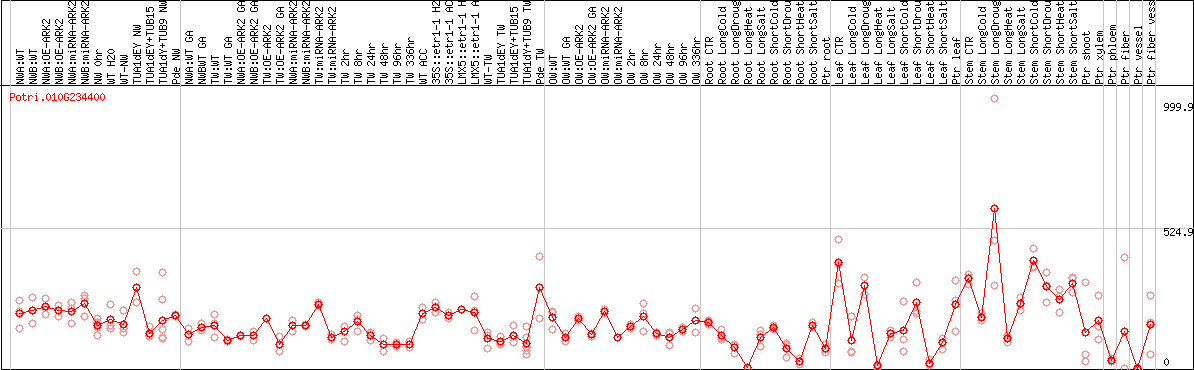
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.010G234400 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
