Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G66950 236 / 3e-71
ABCG39, PDR11, ATPDR11
ATP-binding cassette G39, pleiotropic drug resistance 11 (.1)
AT2G36380 224 / 3e-67
ABCG34, PDR6, ATPDR6
ATP-binding cassette G34, pleiotropic drug resistance 6 (.1)
AT2G26910 169 / 4e-48
PEC1, ABCG32, PDR4, ATPDR4
PERMEABLE CUTICLE 1, ATP-binding cassette G32, pleiotropic drug resistance 4 (.1)
AT2G29940 169 / 8e-48
ABCG31, PDR3, ATPDR3
ATP-binding cassette G31, pleiotropic drug resistance 3 (.1)
AT1G15520 164 / 2e-46
ATABCG40, ABCG40, PDR12, ATPDR12
Arabidopsis thaliana ATP-binding cassette G40, ATP-binding cassette G40, pleiotropic drug resistance 12 (.1)
AT3G16340 152 / 4e-42
ABCG29, ATPDR1, PDR1, AtABCG29
ATP-binding cassette G29, pleiotropic drug resistance 1 (.1.2)
AT1G15210 148 / 1e-40
ABCG35, PDR7, ATPDR7
ATP-binding cassette G35, pleiotropic drug resistance 7 (.1)
AT3G53480 143 / 5e-39
PIS1, ABCG37, PDR9, ATPDR9
polar auxin transport inhibitor sensitive 1, ATP-binding cassette G37, pleiotropic drug resistance 9 (.1)
AT1G59870 142 / 1e-38
ATABCG36, ABCG36, PEN3, PDR8, ATPDR8
PENETRATION 3, ARABIDOPSIS PLEIOTROPIC DRUG RESISTANCE 8, Arabidopsis thaliana ATP-binding cassette G36, ATP-binding cassette G36, ABC-2 and Plant PDR ABC-type transporter family protein (.1)
AT2G37280 135 / 3e-36
ABCG33, PDR5, ATPDR5
ATP-binding cassette G33, pleiotropic drug resistance 5 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.008G024400
355 / 6e-114
AT1G66950 2011 / 0.0
ATP-binding cassette G39, pleiotropic drug resistance 11 (.1)
Potri.006G115000
262 / 2e-80
AT1G66950 2148 / 0.0
ATP-binding cassette G39, pleiotropic drug resistance 11 (.1)
Potri.009G147100
202 / 1e-59
AT2G36380 1759 / 0.0
ATP-binding cassette G34, pleiotropic drug resistance 6 (.1)
Potri.018G032900
178 / 4e-51
AT1G15520 2041 / 0.0
Arabidopsis thaliana ATP-binding cassette G40, ATP-binding cassette G40, pleiotropic drug resistance 12 (.1)
Potri.003G057301
168 / 2e-50
AT1G15520 382 / 1e-121
Arabidopsis thaliana ATP-binding cassette G40, ATP-binding cassette G40, pleiotropic drug resistance 12 (.1)
Potri.003G183200
174 / 6e-50
AT1G15520 2031 / 0.0
Arabidopsis thaliana ATP-binding cassette G40, ATP-binding cassette G40, pleiotropic drug resistance 12 (.1)
Potri.006G248500
173 / 2e-49
AT1G15520 2050 / 0.0
Arabidopsis thaliana ATP-binding cassette G40, ATP-binding cassette G40, pleiotropic drug resistance 12 (.1)
Potri.001G048900
172 / 4e-49
AT1G15520 2055 / 0.0
Arabidopsis thaliana ATP-binding cassette G40, ATP-binding cassette G40, pleiotropic drug resistance 12 (.1)
Potri.001G048700
171 / 1e-48
AT1G15520 2000 / 0.0
Arabidopsis thaliana ATP-binding cassette G40, ATP-binding cassette G40, pleiotropic drug resistance 12 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10008984
254 / 2e-77
AT2G36380 2007 / 0.0
ATP-binding cassette G34, pleiotropic drug resistance 6 (.1)
Lus10028841
228 / 2e-68
AT1G66950 1374 / 0.0
ATP-binding cassette G39, pleiotropic drug resistance 11 (.1)
Lus10014384
223 / 1e-66
AT1G66950 1917 / 0.0
ATP-binding cassette G39, pleiotropic drug resistance 11 (.1)
Lus10027725
211 / 1e-62
AT1G66950 1077 / 0.0
ATP-binding cassette G39, pleiotropic drug resistance 11 (.1)
Lus10006277
196 / 2e-57
AT1G66950 1791 / 0.0
ATP-binding cassette G39, pleiotropic drug resistance 11 (.1)
Lus10020568
196 / 3e-57
AT1G66950 1761 / 0.0
ATP-binding cassette G39, pleiotropic drug resistance 11 (.1)
Lus10023879
191 / 8e-56
AT2G36380 1369 / 0.0
ATP-binding cassette G34, pleiotropic drug resistance 6 (.1)
Lus10019453
179 / 3e-51
AT2G26910 2354 / 0.0
PERMEABLE CUTICLE 1, ATP-binding cassette G32, pleiotropic drug resistance 4 (.1)
Lus10041668
168 / 1e-47
AT1G15520 2128 / 0.0
Arabidopsis thaliana ATP-binding cassette G40, ATP-binding cassette G40, pleiotropic drug resistance 12 (.1)
Lus10041685
167 / 2e-47
AT1G15520 1271 / 0.0
Arabidopsis thaliana ATP-binding cassette G40, ATP-binding cassette G40, pleiotropic drug resistance 12 (.1)
Representative CDS sequence
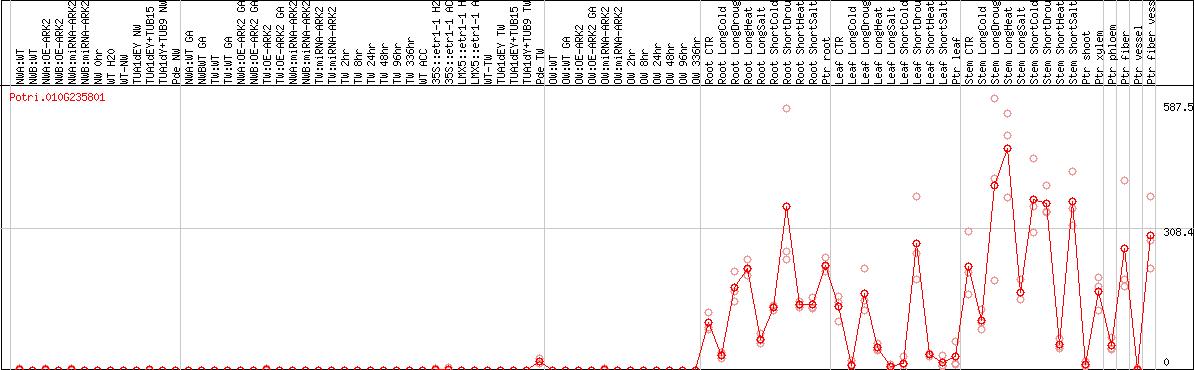
>Potri.010G235801.1 pacid=42798009 polypeptide=Potri.010G235801.1.p locus=Potri.010G235801 ID=Potri.010G235801.1.v4.1 annot-version=v4.1
ATGACATACATGAAAAAAAAATTACACCATTTTGCAGATAATATTGGACCCTGGATGATCTGGGGCTACCATATTTCTCCCATGATGCATGGACAGAAAG
CCATAGTCCTCAAAGAGTTTCTCGACGAAAGATGGAGTGTGCCAAATCATGACAAAGCATTTTCTGAGCCCACTGTTGGAAAGGTTCTTCTGAAAATGAG
AGGCATGTTCATGGAAGGATATTGGTATTGGATTTCGATTGGGGTACTTCGAGGATTTGCTCTGGTTTTTAATGTCCTTTTTGTAGCCGCACTGACTTAT
TTGGATCCTCTGGGAGAATCTAAATCCATTATTCTGGAAGAGGATGAGACTACGAAGTCATCCTCAACTGGAAAGCAAGCAGCAAAGGCCTTAGAAATGA
CTCCAGCATCTGCTGCCCAATTCTCTGAAGTTGCAGAGCAAGCGCCTAAGCAACGAGGAATGGTGTTGCCCTTCCAGCCCTTGTCACTTGCATTTAATCA
TGTGAATTACTATGTTGATGTGCCTATGGAAATGAAAATGCAAGGAATTGAAGAGGATAGCCTCCAGCTGTTACGTGATGTTAGTGGTGTTTTCAGGCCC
GGTGTTCTAACAGCATTGGTTGGTGTAACTTGTAAGTGGTGCTGGAAAGACAACTTTGATGGATGTTCTTGCTGGTCGGAAAACTGGAGGGTACATTGA
AA sequence
>Potri.010G235801.1 pacid=42798009 polypeptide=Potri.010G235801.1.p locus=Potri.010G235801 ID=Potri.010G235801.1.v4.1 annot-version=v4.1
MTYMKKKLHHFADNIGPWMIWGYHISPMMHGQKAIVLKEFLDERWSVPNHDKAFSEPTVGKVLLKMRGMFMEGYWYWISIGVLRGFALVFNVLFVAALTY
LDPLGESKSIILEEDETTKSSSTGKQAAKALEMTPASAAQFSEVAEQAPKQRGMVLPFQPLSLAFNHVNYYVDVPMEMKMQGIEEDSLQLLRDVSGVFRP
GVLTALVGVTCKWCWKDNFDGCSCWSENWRVH