External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G66240 129 / 1e-40
ATX1, ATATX1
homolog of anti-oxidant 1 (.1.2.3)
AT3G56240 122 / 1e-37
ATX1, CCH
copper chaperone (.1)
AT5G02600 70 / 9e-16
NPCC6, NAKR1
nuclear-enriched phloem companion cell gene 6, SODIUM POTASSIUM ROOT DEFECTIVE 1, Heavy metal transport/detoxification superfamily protein (.1.2)
AT5G27690 69 / 3e-15
Heavy metal transport/detoxification superfamily protein (.1)
AT1G71050 62 / 9e-14
HIPP20
heavy metal associated isoprenylated plant protein 20, Heavy metal transport/detoxification superfamily protein (.1)
AT2G37390 64 / 1e-13
NAKR2
SODIUM POTASSIUM ROOT DEFECTIVE 2, Chloroplast-targeted copper chaperone protein (.1.2)
AT5G17450 61 / 2e-13
HIPP21
heavy metal associated isoprenylated plant protein 21, Heavy metal transport/detoxification superfamily protein (.1.2)
AT2G28660 62 / 4e-13
Chloroplast-targeted copper chaperone protein (.1)
AT1G06330 60 / 6e-13
Heavy metal transport/detoxification superfamily protein (.1)
AT2G18196 61 / 8e-13
Heavy metal transport/detoxification superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.008G023800
129 / 5e-41
AT1G66240 124 / 7e-39
homolog of anti-oxidant 1 (.1.2.3)
Potri.002G092200
69 / 3e-16
AT4G08570 193 / 3e-64
Heavy metal transport/detoxification superfamily protein (.1)
Potri.006G024800
68 / 4e-16
AT3G56891 167 / 5e-54
Heavy metal transport/detoxification superfamily protein (.1)
Potri.010G114600
68 / 4e-16
AT1G22990 194 / 9e-65
heavy metal associated isoprenylated plant protein 22, Heavy metal transport/detoxification superfamily protein (.1)
Potri.005G167000
67 / 1e-15
AT4G08570 229 / 1e-78
Heavy metal transport/detoxification superfamily protein (.1)
Potri.006G213900
67 / 8e-15
AT2G37390 127 / 2e-35
SODIUM POTASSIUM ROOT DEFECTIVE 2, Chloroplast-targeted copper chaperone protein (.1.2)
Potri.016G080400
66 / 2e-14
AT2G37390 133 / 2e-37
SODIUM POTASSIUM ROOT DEFECTIVE 2, Chloroplast-targeted copper chaperone protein (.1.2)
Potri.005G003700
64 / 2e-14
AT4G08570 207 / 5e-70
Heavy metal transport/detoxification superfamily protein (.1)
Potri.001G234700
65 / 4e-14
AT2G37390 114 / 2e-30
SODIUM POTASSIUM ROOT DEFECTIVE 2, Chloroplast-targeted copper chaperone protein (.1.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10043444
118 / 6e-36
AT1G66240 127 / 4e-39
homolog of anti-oxidant 1 (.1.2.3)
Lus10028859
117 / 2e-35
AT1G66240 125 / 8e-38
homolog of anti-oxidant 1 (.1.2.3)
Lus10034138
103 / 1e-28
AT1G66240 115 / 1e-32
homolog of anti-oxidant 1 (.1.2.3)
Lus10008963
79 / 2e-19
AT1G66240 81 / 5e-20
homolog of anti-oxidant 1 (.1.2.3)
Lus10024435
71 / 3e-16
AT2G37390 144 / 2e-41
SODIUM POTASSIUM ROOT DEFECTIVE 2, Chloroplast-targeted copper chaperone protein (.1.2)
Lus10025294
71 / 4e-16
AT2G37390 141 / 3e-40
SODIUM POTASSIUM ROOT DEFECTIVE 2, Chloroplast-targeted copper chaperone protein (.1.2)
Lus10042946
63 / 5e-14
AT1G71050 196 / 3e-65
heavy metal associated isoprenylated plant protein 20, Heavy metal transport/detoxification superfamily protein (.1)
Lus10036004
63 / 6e-14
AT1G22990 187 / 4e-62
heavy metal associated isoprenylated plant protein 22, Heavy metal transport/detoxification superfamily protein (.1)
Lus10016436
63 / 8e-14
AT4G39700 209 / 1e-70
Heavy metal transport/detoxification superfamily protein (.1)
Lus10019676
63 / 8e-14
AT4G39700 211 / 3e-71
Heavy metal transport/detoxification superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF00403
HMA
Heavy-metal-associated domain
Representative CDS sequence
>Potri.010G236500.1 pacid=42798515 polypeptide=Potri.010G236500.1.p locus=Potri.010G236500 ID=Potri.010G236500.1.v4.1 annot-version=v4.1
ATGTCTCAGACTGTTGTCCTCAAGGTTGGTATGTCATGCGAAGGCTGTGTTGGGGCTGTGAAAAGGGTTTTGGGAAAAATGGAAGGTGTGGAATCATATG
ACATTGATTTGAAGGAGCAAAAAGTCACAGTGAAAGGAAATGTGCAGCCAGATGCTGTTCTTCAGACCGTCTCTAAGACCGGGAAGAAGACTGCCTTCTG
GGAAGCAGAGGCACCAGCTGAACCCGCAAAGCCTGCAGAAACCGTGGCTGCTGCATAA
AA sequence
>Potri.010G236500.1 pacid=42798515 polypeptide=Potri.010G236500.1.p locus=Potri.010G236500 ID=Potri.010G236500.1.v4.1 annot-version=v4.1
MSQTVVLKVGMSCEGCVGAVKRVLGKMEGVESYDIDLKEQKVTVKGNVQPDAVLQTVSKTGKKTAFWEAEAPAEPAKPAETVAAA
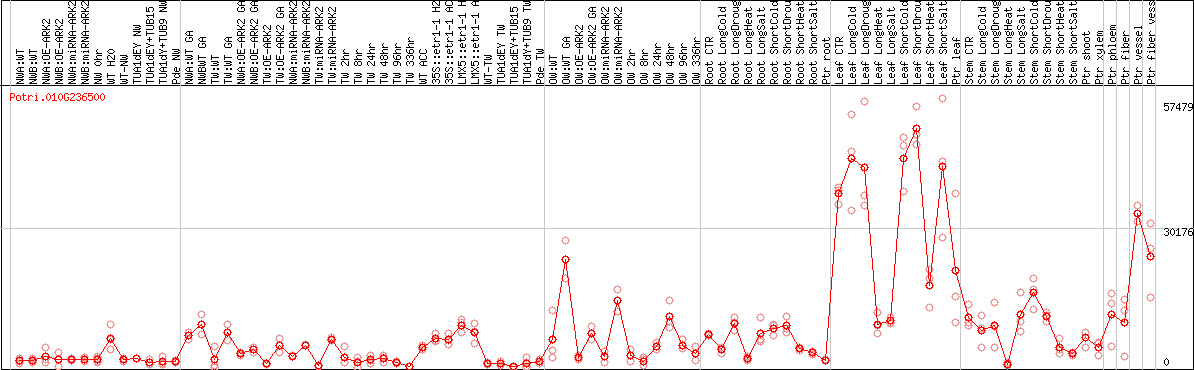
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.010G236500 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.