External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G04840 228 / 2e-73
bZIP
bZIP protein (.1)
AT3G58120 88 / 8e-20
bZIP
ATBZIP61
Basic-leucine zipper (bZIP) transcription factor family protein (.1)
AT2G42380 84 / 1e-18
bZIP
ATBZIP34
Basic-leucine zipper (bZIP) transcription factor family protein (.1), Basic-leucine zipper (bZIP) transcription factor family protein (.2)
AT1G58110 71 / 1e-13
bZIP
Basic-leucine zipper (bZIP) transcription factor family protein (.1), Basic-leucine zipper (bZIP) transcription factor family protein (.2)
AT1G06070 64 / 3e-11
bZIP
AtbZIP69
Basic-leucine zipper (bZIP) transcription factor family protein (.1)
AT2G40620 59 / 7e-10
bZIP
AtbZIP18
Basic-leucine zipper (bZIP) transcription factor family protein (.1)
AT4G38900 58 / 3e-09
bZIP
AtbZIP29
Basic-leucine zipper (bZIP) transcription factor family protein (.1), Basic-leucine zipper (bZIP) transcription factor family protein (.2), Basic-leucine zipper (bZIP) transcription factor family protein (.3)
AT1G35490 57 / 4e-09
bZIP
bZIP family transcription factor (.1)
AT4G06598 57 / 4e-09
unknown protein
AT2G31370 56 / 1e-08
bZIP
AtbZIP59, PosF21
Basic-leucine zipper (bZIP) transcription factor family protein
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.008G018400
495 / 6e-179
AT5G04840 222 / 3e-71
bZIP protein (.1)
Potri.001G374200
87 / 1e-19
AT3G58120 282 / 6e-94
Basic-leucine zipper (bZIP) transcription factor family protein (.1)
Potri.013G124400
84 / 3e-18
AT3G58120 293 / 2e-98
Basic-leucine zipper (bZIP) transcription factor family protein (.1)
Potri.003G014800
82 / 2e-17
AT1G58110 304 / 3e-101
Basic-leucine zipper (bZIP) transcription factor family protein (.1), Basic-leucine zipper (bZIP) transcription factor family protein (.2)
Potri.019G091900
81 / 2e-17
AT3G58120 285 / 4e-95
Basic-leucine zipper (bZIP) transcription factor family protein (.1)
Potri.004G219100
74 / 8e-15
AT1G58110 303 / 8e-101
Basic-leucine zipper (bZIP) transcription factor family protein (.1), Basic-leucine zipper (bZIP) transcription factor family protein (.2)
Potri.015G147800
66 / 3e-12
AT1G58110 153 / 4e-43
Basic-leucine zipper (bZIP) transcription factor family protein (.1), Basic-leucine zipper (bZIP) transcription factor family protein (.2)
Potri.013G091400
64 / 3e-11
AT2G40620 300 / 2e-100
Basic-leucine zipper (bZIP) transcription factor family protein (.1)
Potri.005G190700
62 / 8e-11
AT1G43700 219 / 2e-69
sulphate utilization efficiency 3, VIRE2-interacting protein 1 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10017656
84 / 2e-18
AT2G42380 305 / 1e-103
Basic-leucine zipper (bZIP) transcription factor family protein (.1), Basic-leucine zipper (bZIP) transcription factor family protein (.2)
Lus10033630
84 / 2e-18
AT2G42380 294 / 7e-99
Basic-leucine zipper (bZIP) transcription factor family protein (.1), Basic-leucine zipper (bZIP) transcription factor family protein (.2)
Lus10039902
83 / 2e-18
AT3G58120 181 / 8e-56
Basic-leucine zipper (bZIP) transcription factor family protein (.1)
Lus10002185
83 / 3e-18
AT3G58120 219 / 1e-71
Basic-leucine zipper (bZIP) transcription factor family protein (.1)
Lus10015457
81 / 4e-17
AT1G58110 315 / 2e-105
Basic-leucine zipper (bZIP) transcription factor family protein (.1), Basic-leucine zipper (bZIP) transcription factor family protein (.2)
Lus10028595
60 / 4e-10
AT1G43700 289 / 1e-96
sulphate utilization efficiency 3, VIRE2-interacting protein 1 (.1)
Lus10008605
60 / 5e-10
AT2G40620 276 / 1e-90
Basic-leucine zipper (bZIP) transcription factor family protein (.1)
Lus10024204
60 / 5e-10
AT1G06850 271 / 5e-90
basic leucine-zipper 52 (.1.2)
Lus10018899
60 / 5e-10
AT1G43700 287 / 7e-96
sulphate utilization efficiency 3, VIRE2-interacting protein 1 (.1)
Lus10009907
60 / 6e-10
AT1G06850 270 / 2e-89
basic leucine-zipper 52 (.1.2)
PFAM info
Representative CDS sequence
>Potri.010G242800.3 pacid=42798971 polypeptide=Potri.010G242800.3.p locus=Potri.010G242800 ID=Potri.010G242800.3.v4.1 annot-version=v4.1
ATGTCAAGGCAATCTCTCCTTCCTCCTCGCTGCCCATTTCAGAAGCCCGTGGTTTCTCGACCAATTCATGACTCCTACCCACAACACCATAGATCTCCTT
CCCAAGGTTCAATTTTGGGGGAGAAGCCGGCATGGCTTGATGATTTGTTAAGTGATGAAGATGCAGATTCCAGGGGGACTTGTCTTCGCCGGTCAGCCAG
TGATTCAGTCACCCTCTTGGAAGGTATTGTGGATTCGTTTTCGGGTTCGAGTCCTTATAACAATGAAGCTGCCTCAGGTGGGGGTGAAACTTGTAGTGGG
CTGGAGTCAGCTTCTATGTATGGTCCTAATTCTCCTAGGCGAAGAGGGAATGTGACCTTTTCTGAGAATGCCATAGCTTCAGCACTATCGGAGTATGCGT
TTCAGAATCCTCTGCAGTATGTAGACGGCAGCTTGTGCATATGGGGGAATACTCCCTTGGATTCAATGGGGAATGCATGTGGTTCGGCTGGGGAGCTCAA
TGGCGAGACGAGCACTGTGAAACGGCAATCTGGGCAGCGCTCAAGAGTTCGCAAACTCCAGTATATTGCTGAACTCGAAAGAACTGTTAATGTTCTGCAG
ACTTTGGAGTCAGAGTTGGCGTTCAAGGTTGCTTCAATGCTTCAGAAACGTGCTGCCTTGTCCCTGGAAAACAACACATTAAAGCAGCAGGTGGCCAGAC
TGCGGCAGGAAAAATTAATCGTGGATGCTCAACATAAGACTTTGAAGAAGGAAGCTGAGAGATTGAAAAACAAGTTAGGTAGTTCCACAAATAATAAGTT
CAGAACATATTCCAGATCAAGTCTTTCTCCAGAGGCAGCTAGATCAGAAGTTACATGGCAGATGGCGAGGCTTAATCTTAACTAA
AA sequence
>Potri.010G242800.3 pacid=42798971 polypeptide=Potri.010G242800.3.p locus=Potri.010G242800 ID=Potri.010G242800.3.v4.1 annot-version=v4.1
MSRQSLLPPRCPFQKPVVSRPIHDSYPQHHRSPSQGSILGEKPAWLDDLLSDEDADSRGTCLRRSASDSVTLLEGIVDSFSGSSPYNNEAASGGGETCSG
LESASMYGPNSPRRRGNVTFSENAIASALSEYAFQNPLQYVDGSLCIWGNTPLDSMGNACGSAGELNGETSTVKRQSGQRSRVRKLQYIAELERTVNVLQ
TLESELAFKVASMLQKRAALSLENNTLKQQVARLRQEKLIVDAQHKTLKKEAERLKNKLGSSTNNKFRTYSRSSLSPEAARSEVTWQMARLNLN
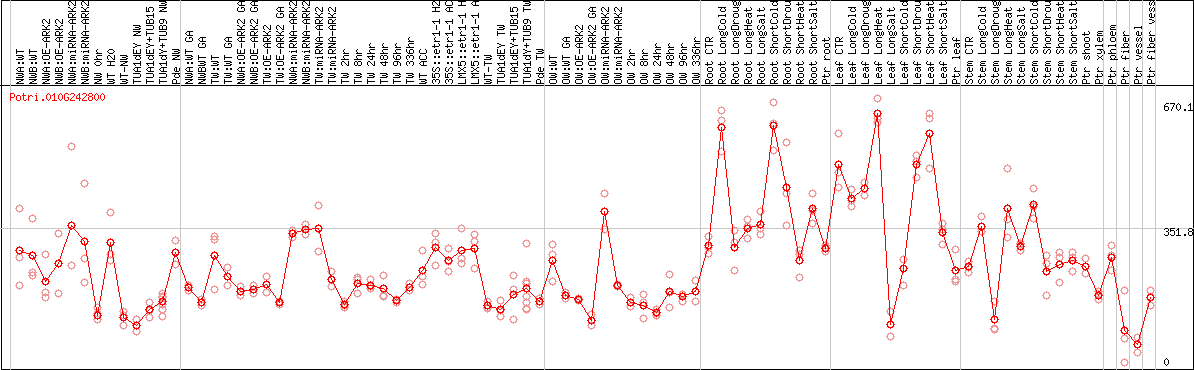
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.010G242800 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.