External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G16010 295 / 7e-101
3-oxo-5-alpha-steroid 4-dehydrogenase family protein (.1)
AT2G38050 77 / 2e-16
DWF6, DET2, ATDET2
DWARF 6, DE-ETIOLATED 2, 3-oxo-5-alpha-steroid 4-dehydrogenase family protein (.1)
AT1G72590 72 / 2e-14
3-oxo-5-alpha-steroid 4-dehydrogenase family protein (.1)
AT2G16530 69 / 2e-13
3-oxo-5-alpha-steroid 4-dehydrogenase family protein (.1.2)
AT3G55360 58 / 2e-09
ECR, CER10, ATTSC13, TSC13
ENOYL-COA REDUCTASE, ECERIFERUM 10, 3-oxo-5-alpha-steroid 4-dehydrogenase family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.008G012700
428 / 2e-153
AT5G16010 278 / 2e-94
3-oxo-5-alpha-steroid 4-dehydrogenase family protein (.1)
Potri.008G013100
332 / 2e-115
AT5G16010 298 / 4e-102
3-oxo-5-alpha-steroid 4-dehydrogenase family protein (.1)
Potri.010G245266
332 / 3e-115
AT5G16010 291 / 2e-99
3-oxo-5-alpha-steroid 4-dehydrogenase family protein (.1)
Potri.010G245332
326 / 6e-113
AT5G16010 289 / 1e-98
3-oxo-5-alpha-steroid 4-dehydrogenase family protein (.1)
Potri.008G012800
317 / 2e-109
AT5G16010 266 / 3e-89
3-oxo-5-alpha-steroid 4-dehydrogenase family protein (.1)
Potri.013G134433
79 / 4e-17
AT2G38050 303 / 3e-104
DWARF 6, DE-ETIOLATED 2, 3-oxo-5-alpha-steroid 4-dehydrogenase family protein (.1)
Potri.016G110600
79 / 5e-17
AT2G38050 350 / 1e-122
DWARF 6, DE-ETIOLATED 2, 3-oxo-5-alpha-steroid 4-dehydrogenase family protein (.1)
Potri.005G047800
77 / 2e-16
AT2G38050 295 / 8e-101
DWARF 6, DE-ETIOLATED 2, 3-oxo-5-alpha-steroid 4-dehydrogenase family protein (.1)
Potri.004G163600
67 / 1e-12
AT2G16530 418 / 6e-147
3-oxo-5-alpha-steroid 4-dehydrogenase family protein (.1.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10008951
248 / 1e-81
AT5G16010 272 / 4e-91
3-oxo-5-alpha-steroid 4-dehydrogenase family protein (.1)
Lus10028869
127 / 2e-36
AT5G16010 123 / 2e-35
3-oxo-5-alpha-steroid 4-dehydrogenase family protein (.1)
Lus10021127
80 / 6e-18
AT2G38050 256 / 3e-87
DWARF 6, DE-ETIOLATED 2, 3-oxo-5-alpha-steroid 4-dehydrogenase family protein (.1)
Lus10017183
74 / 9e-16
AT2G38050 138 / 2e-64
DWARF 6, DE-ETIOLATED 2, 3-oxo-5-alpha-steroid 4-dehydrogenase family protein (.1)
Lus10028371
69 / 2e-13
AT2G16530 336 / 4e-115
3-oxo-5-alpha-steroid 4-dehydrogenase family protein (.1.2)
Lus10001695
59 / 5e-10
AT3G55360 540 / 0.0
ENOYL-COA REDUCTASE, ECERIFERUM 10, 3-oxo-5-alpha-steroid 4-dehydrogenase family protein (.1)
Lus10042055
59 / 9e-10
AT2G16530 347 / 8e-120
3-oxo-5-alpha-steroid 4-dehydrogenase family protein (.1.2)
Lus10005162
58 / 2e-09
AT3G55360 541 / 0.0
ENOYL-COA REDUCTASE, ECERIFERUM 10, 3-oxo-5-alpha-steroid 4-dehydrogenase family protein (.1)
Lus10041820
0 / 1
AT2G16530 289 / 2e-96
3-oxo-5-alpha-steroid 4-dehydrogenase family protein (.1.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0115
Steroid_dh
PF02544
Steroid_dh
3-oxo-5-alpha-steroid 4-dehydrogenase
Representative CDS sequence
>Potri.010G245200.1 pacid=42799236 polypeptide=Potri.010G245200.1.p locus=Potri.010G245200 ID=Potri.010G245200.1.v4.1 annot-version=v4.1
ATGGCATCCATGTTGTCTAGATTAATTATCTTTCCTACACCACCTTCTATGATCGTTCCAGTCATGTCTGTGGTTAGTCTGGTAGCAATAGCAGGTCTTG
GTGTTTCTGAAATCTTAGGAAAGCATTTACAGTACTCAAAATTCTGGAACTTGAATTCTGCAAAATCTTCAAGAAAGCAGATACAACTCTCCAGCAGAAC
TGGCATGCTTGTACTTTATGTGCCAGCTTTCCTTTCTGGTGTTGCTTCTTTTGTGCTCTATCCAAATCATGATCTCAGGTTGTTCCTGGTTAAAGCAACC
CTTACCATACATTTCTTCAAGAGGGTCATTGAGGTGCTGTGTGTTCACAAATATAGTACTAGTGGGGTGGTTCTTGATTCTGCAATTCTTATAAGTTCAA
GTTATTTCTCAGCTACTTCAACCATGATATATGGTCAGTATCTAACACAAGGGTTTCCTGAGCCACAGCTGGACTTGAAGTATCCAGGGATTTTGCTGTT
CATGTTGGGAACCTTCGGCAACTTCTACCACCACCAAGTTCTTGCCTCTTTGAGAACAAATGATGACAAAGAATACAAGATTCCAAAGGGTGGTTTATTT
GACCTGGTGATCTGCCCTCACTATCTCTTTGAGGTGTTGGGGTTTATAGGGATATTTTTCATTTCTCAAACACTGTATTCCTTTTGCTTCACTTTAGGCA
CTATTGTTTACTTGATTGGAAGGAGTTACGCTACTAGGAGATGGTACCTTTCCAAGTTCGAAGATTTTCCCAAGGATGTCAAGGCTCTTATTCCTTTTGT
ATTTTGA
AA sequence
>Potri.010G245200.1 pacid=42799236 polypeptide=Potri.010G245200.1.p locus=Potri.010G245200 ID=Potri.010G245200.1.v4.1 annot-version=v4.1
MASMLSRLIIFPTPPSMIVPVMSVVSLVAIAGLGVSEILGKHLQYSKFWNLNSAKSSRKQIQLSSRTGMLVLYVPAFLSGVASFVLYPNHDLRLFLVKAT
LTIHFFKRVIEVLCVHKYSTSGVVLDSAILISSSYFSATSTMIYGQYLTQGFPEPQLDLKYPGILLFMLGTFGNFYHHQVLASLRTNDDKEYKIPKGGLF
DLVICPHYLFEVLGFIGIFFISQTLYSFCFTLGTIVYLIGRSYATRRWYLSKFEDFPKDVKALIPFVF
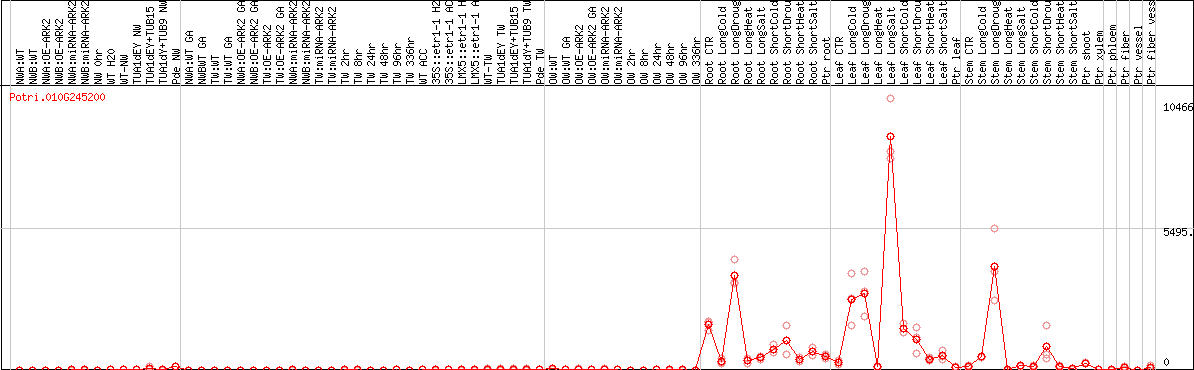
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.010G245200 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.