External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G22120 275 / 2e-88
ERD (early-responsive to dehydration stress) family protein
AT1G11960 265 / 2e-84
ERD (early-responsive to dehydration stress) family protein (.1)
AT4G04340 265 / 2e-84
ERD (early-responsive to dehydration stress) family protein (.1), ERD (early-responsive to dehydration stress) family protein (.2), ERD (early-responsive to dehydration stress) family protein (.3)
AT4G15430 263 / 5e-84
ERD (early-responsive to dehydration stress) family protein (.1), ERD (early-responsive to dehydration stress) family protein (.2)
AT4G02900 263 / 2e-83
ERD (early-responsive to dehydration stress) family protein (.1)
AT1G62320 258 / 8e-82
ERD (early-responsive to dehydration stress) family protein (.1)
AT3G21620 253 / 6e-80
ERD (early-responsive to dehydration stress) family protein (.1)
AT1G32090 247 / 2e-77
early-responsive to dehydration stress protein (ERD4) (.1)
AT1G58520 124 / 5e-33
RXW8
lipases;hydrolases, acting on ester bonds (.1.2.3)
AT1G10090 121 / 6e-32
Early-responsive to dehydration stress protein (ERD4) (.1)
Paralogs
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10020032
295 / 8e-96
AT4G22120 1186 / 0.0
ERD (early-responsive to dehydration stress) family protein
Lus10015551
295 / 2e-95
AT4G22120 1183 / 0.0
ERD (early-responsive to dehydration stress) family protein
Lus10024477
282 / 7e-91
AT1G11960 1072 / 0.0
ERD (early-responsive to dehydration stress) family protein (.1)
Lus10002996
282 / 8e-91
AT4G22120 1087 / 0.0
ERD (early-responsive to dehydration stress) family protein
Lus10030024
276 / 1e-88
AT3G21620 1147 / 0.0
ERD (early-responsive to dehydration stress) family protein (.1)
Lus10035300
275 / 4e-88
AT3G21620 1146 / 0.0
ERD (early-responsive to dehydration stress) family protein (.1)
Lus10002133
263 / 3e-83
AT1G32090 1136 / 0.0
early-responsive to dehydration stress protein (ERD4) (.1)
Lus10028000
251 / 2e-79
AT4G02900 1116 / 0.0
ERD (early-responsive to dehydration stress) family protein (.1)
Lus10000660
249 / 2e-78
AT4G02900 1113 / 0.0
ERD (early-responsive to dehydration stress) family protein (.1)
Lus10004597
127 / 7e-34
AT1G58520 845 / 0.0
lipases;hydrolases, acting on ester bonds (.1.2.3)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0416
Anoctamin-like
PF02714
RSN1_7TM
Calcium-dependent channel, 7TM region, putative phosphate
Representative CDS sequence
>Potri.011G009801.1 pacid=42781923 polypeptide=Potri.011G009801.1.p locus=Potri.011G009801 ID=Potri.011G009801.1.v4.1 annot-version=v4.1
ATGATGGATGGTTCTGTTATTATTCAGATAGCAGAAGATAGGGAAAAGATTTTAAATGATCCGAATTCTATCATGCCAGCTGCATTCGTTTCGTTCAAGA
CTCGATGGGGTGCAGCGTTTTGTGCACAAACTCAACAATCAAGAAATCCAACTTTGTGGTTAACAGAGTGGGCTCCTGAGCCTCGCGATGTGTATTGGGA
AAACTTAGCCATTCCATATGTGTCACTTTCTGTTAGGAGGCTGATAGTTGGAGTTTCATTCTTTTTTCTTGCCTTCTTATTCTTGATCCCTATTGCATTT
GTACAATCTCTAGCAAGCATTGAGGGAATTGAGAAAAACCTTCCCTTTTTGAAGCCTGTTATTGAAATAGAATTTATCAAATCAGTTGCCCAAGGATTTC
TACCTGGGATTGCACTGAAACTCTTTCTCACCTTCCTGCCGACAGTATTGATGATGATGTCTAAATTGGAAGGCTTCATGTCTCTATCATCTTTGGAAAG
GATATCAGCAATGAGATATTACATTTTCATCATTATTGATGTGTTCCTTGGAAGCATACTCACCGGGGCTGTGTTCGAACAGCTAAATTCTTTTATCAAT
CAGTCATCAGTCTGCTAG
AA sequence
>Potri.011G009801.1 pacid=42781923 polypeptide=Potri.011G009801.1.p locus=Potri.011G009801 ID=Potri.011G009801.1.v4.1 annot-version=v4.1
MMDGSVIIQIAEDREKILNDPNSIMPAAFVSFKTRWGAAFCAQTQQSRNPTLWLTEWAPEPRDVYWENLAIPYVSLSVRRLIVGVSFFFLAFLFLIPIAF
VQSLASIEGIEKNLPFLKPVIEIEFIKSVAQGFLPGIALKLFLTFLPTVLMMMSKLEGFMSLSSLERISAMRYYIFIIIDVFLGSILTGAVFEQLNSFIN
QSSVC
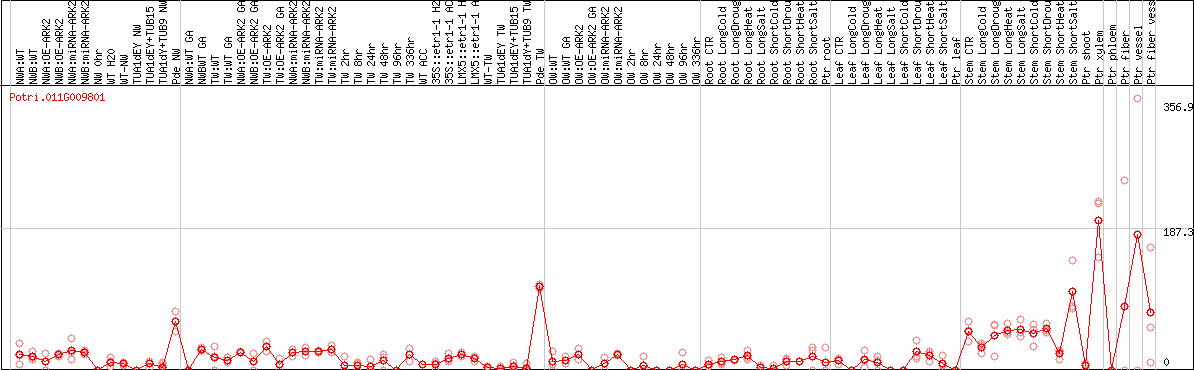
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.011G009801 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.