External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G36930 134 / 2e-35
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
AT1G27170 105 / 2e-25
transmembrane receptors;ATP binding (.1.2)
AT1G27180 99 / 2e-23
disease resistance protein (TIR-NBS-LRR class), putative (.1)
AT4G19510 86 / 6e-19
Disease resistance protein (TIR-NBS-LRR class) (.1), Disease resistance protein (TIR-NBS-LRR class) (.2)
AT5G46260 82 / 1e-17
disease resistance protein (TIR-NBS-LRR class) family (.1)
AT5G46270 81 / 3e-17
Disease resistance protein (TIR-NBS-LRR class) family (.1)
AT5G45260 81 / 4e-17
WRKY
ATWRKY52, SLH1, RRS1
SENSITIVE TO LOW HUMIDITY 1, RESISTANT TO RALSTONIA SOLANACEARUM 1, ARABIDOPSIS THALIANA WRKY DOMAIN PROTEIN 52, Disease resistance protein (TIR-NBS-LRR class) (.1), Disease resistance protein (TIR-NBS-LRR class) (.2)
AT5G22690 80 / 1e-16
Disease resistance protein (TIR-NBS-LRR class) family (.1)
AT5G46490 79 / 1e-16
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
AT1G72840 79 / 2e-16
Disease resistance protein (TIR-NBS-LRR class) (.1), Disease resistance protein (TIR-NBS-LRR class) (.2)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.011G013700
523 / 0
AT5G36930 378 / 3e-117
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Potri.006G269900
497 / 1e-172
AT5G36930 284 / 2e-81
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Potri.011G014101
494 / 1e-172
AT5G36930 435 / 1e-137
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Potri.006G282200
503 / 1e-171
AT5G36930 454 / 2e-140
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Potri.011G013900
503 / 2e-171
AT5G36930 441 / 1e-135
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Potri.006G283000
501 / 5e-171
AT5G36930 460 / 8e-143
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Potri.011G013450
498 / 7e-171
AT5G36930 422 / 2e-129
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Potri.011G011600
499 / 3e-170
AT5G36930 492 / 1e-154
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Potri.011G015400
498 / 1e-169
AT5G36930 438 / 1e-134
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10005171
175 / 2e-49
AT1G27170 576 / 0.0
transmembrane receptors;ATP binding (.1.2)
Lus10033825
170 / 6e-48
AT5G36930 400 / 3e-123
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Lus10011741
162 / 4e-45
AT5G36930 540 / 4e-172
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Lus10039850
129 / 8e-34
AT5G36930 573 / 0.0
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Lus10018616
122 / 3e-31
AT5G36930 578 / 0.0
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Lus10030345
98 / 7e-23
AT5G17680 528 / 5e-168
disease resistance protein (TIR-NBS-LRR class), putative (.1)
Lus10041078
97 / 2e-22
AT5G17680 399 / 6e-126
disease resistance protein (TIR-NBS-LRR class), putative (.1)
Lus10003749
96 / 3e-22
AT5G17680 517 / 5e-161
disease resistance protein (TIR-NBS-LRR class), putative (.1)
Lus10024886
94 / 3e-21
AT5G36930 370 / 1e-108
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Lus10018972
92 / 5e-21
AT5G36930 192 / 3e-51
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
PFAM info
Representative CDS sequence
>Potri.011G014301.1 pacid=42781477 polypeptide=Potri.011G014301.1.p locus=Potri.011G014301 ID=Potri.011G014301.1.v4.1 annot-version=v4.1
ATGGAAGTGATTCCTAATTTTGAAGTTCAAAAGGTTCTTCGAATAAGTTACGACTTTCTTGATGGTGATTATCCGAAGATCTTATTCCTTGATATCACAT
GTTTCTTCAATGGAATGGATGTGGATGATGCAGTTAGGATACTGGATGGGCTCGATAAAGGTGCAAGATTTGGGATTGACAATCTCATCGATAGATGTCT
TGTTGAAATCAACAATGATCAAAGGTTGTGGATGCATCAACTAGTAATAGATATGGGAAGGGAAATTGCTCGTCAAGAATCACCCAAATGTCAAAGAATA
TGGCATCAAGGGGATGCTTTTACAGTTTTGAAAGGAACTACTGATGCTGAAAAATTGCGTGGCCTTACCATTGATATGCATGCATTAATGGAATATCATT
ATGCAGAAGTTGTCTGTACTGATTCAATGGTTTGTCGCAAGCGCCGCAGGCTTAACTTCTTTCAACAATGGCTTTCCGATTTTTTCGATGGGGGAAAATT
ACAAACTGGCCAAACAAGTTTGTTTCCCATCCTCAACACGGATGCTTTTAGAAAGATGCCAGATGTAAAATTTCTCCAACTAAACTACACTAATTTTCAT
GGAAGTTTTGAGCACTTTCCCAAGAATTTGATATGGTTATGTTGGCATGGATTGTCTTGGAGCTCCATACCAAATCACGTATGCTTGGAGAAGCTGGTGG
TTCTTGATCTATCCAGAAGTTGTCTAGTTGATGCTTGGAAGGGCAAACCGAACCCCAGACTTCTCGGGTCTCCCAGCCCTTGA
AA sequence
>Potri.011G014301.1 pacid=42781477 polypeptide=Potri.011G014301.1.p locus=Potri.011G014301 ID=Potri.011G014301.1.v4.1 annot-version=v4.1
MEVIPNFEVQKVLRISYDFLDGDYPKILFLDITCFFNGMDVDDAVRILDGLDKGARFGIDNLIDRCLVEINNDQRLWMHQLVIDMGREIARQESPKCQRI
WHQGDAFTVLKGTTDAEKLRGLTIDMHALMEYHYAEVVCTDSMVCRKRRRLNFFQQWLSDFFDGGKLQTGQTSLFPILNTDAFRKMPDVKFLQLNYTNFH
GSFEHFPKNLIWLCWHGLSWSSIPNHVCLEKLVVLDLSRSCLVDAWKGKPNPRLLGSPSP
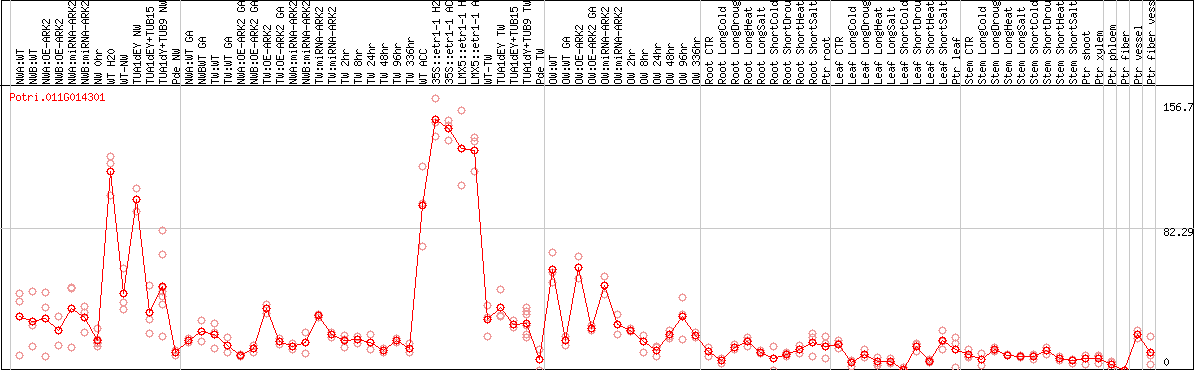
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.011G014301 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.