External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G22310 205 / 3e-70
Uncharacterised protein family (UPF0041) (.1)
AT4G14695 196 / 1e-66
Uncharacterised protein family (UPF0041) (.1), Uncharacterised protein family (UPF0041) (.2)
AT4G05590 192 / 3e-65
unknown protein
AT5G20090 77 / 6e-20
Uncharacterised protein family (UPF0041) (.1), Uncharacterised protein family (UPF0041) (.2), Uncharacterised protein family (UPF0041) (.3)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.001G333902
77 / 2e-19
AT5G20090 144 / 5e-46
Uncharacterised protein family (UPF0041) (.1), Uncharacterised protein family (UPF0041) (.2), Uncharacterised protein family (UPF0041) (.3)
Potri.012G095133
70 / 8e-17
AT5G20090 199 / 5e-68
Uncharacterised protein family (UPF0041) (.1), Uncharacterised protein family (UPF0041) (.2), Uncharacterised protein family (UPF0041) (.3)
Potri.015G092500
67 / 1e-15
AT5G20090 189 / 7e-64
Uncharacterised protein family (UPF0041) (.1), Uncharacterised protein family (UPF0041) (.2), Uncharacterised protein family (UPF0041) (.3)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10023745
211 / 2e-72
AT4G22310 206 / 1e-70
Uncharacterised protein family (UPF0041) (.1)
Lus10011790
207 / 4e-71
AT4G22310 202 / 3e-69
Uncharacterised protein family (UPF0041) (.1)
Lus10032564
162 / 2e-53
AT4G22310 163 / 5e-54
Uncharacterised protein family (UPF0041) (.1)
Lus10025851
75 / 9e-19
AT5G20090 190 / 2e-64
Uncharacterised protein family (UPF0041) (.1), Uncharacterised protein family (UPF0041) (.2), Uncharacterised protein family (UPF0041) (.3)
Lus10038250
75 / 9e-19
AT5G20090 190 / 2e-64
Uncharacterised protein family (UPF0041) (.1), Uncharacterised protein family (UPF0041) (.2), Uncharacterised protein family (UPF0041) (.3)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0141
MtN3-like
PF03650
MPC
Mitochondrial pyruvate carriers
Representative CDS sequence
>Potri.011G023100.2 pacid=42781410 polypeptide=Potri.011G023100.2.p locus=Potri.011G023100 ID=Potri.011G023100.2.v4.1 annot-version=v4.1
ATGGCGACATCGAAGCTACAAGCTCTGTGGAATCACCCAGCCGGTCCCAAAACCATTCATTTTTGGGCACCTACTTTTAAATGGGGTATTAGCATTGCCA
ATATTGCTGACTTCGCTAAACCTCCGGAAAAACTGTCCTACCCTCAGCAAATAGCGGTCACTTGTACTGGAGTTATATGGTCACGCTACAGTACTGTAAT
TACACCAAAAAATTGGAATCTATTTAGTGTAAATGTTGCGATGGCTGCAACAGGCATCTACCAACTCAGTCGCAAAATACAGCACGATTACTTTTCTGAG
GAAGAAGCTGCTGTTGCAAAGGAATAA
AA sequence
>Potri.011G023100.2 pacid=42781410 polypeptide=Potri.011G023100.2.p locus=Potri.011G023100 ID=Potri.011G023100.2.v4.1 annot-version=v4.1
MATSKLQALWNHPAGPKTIHFWAPTFKWGISIANIADFAKPPEKLSYPQQIAVTCTGVIWSRYSTVITPKNWNLFSVNVAMAATGIYQLSRKIQHDYFSE
EEAAVAKE
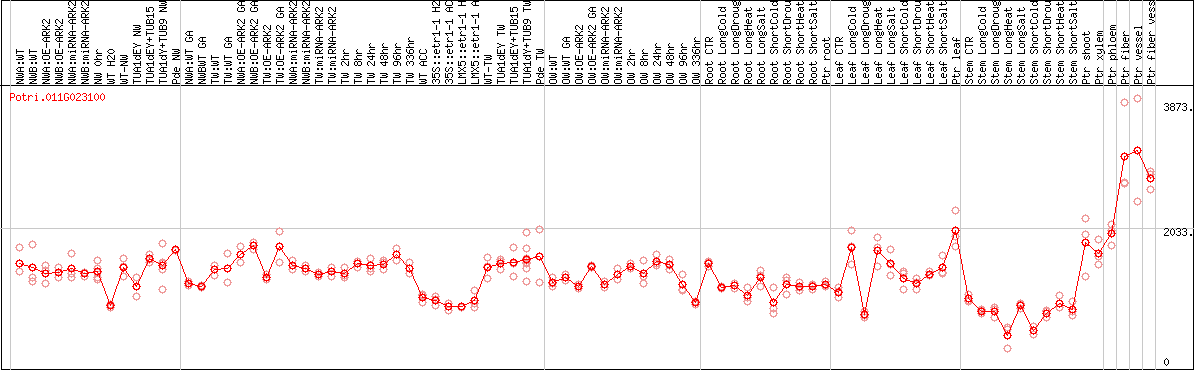
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.011G023100 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.