Pt-SPIKE1.3 (Potri.011G024000) [POPLAR]

| External link |
|
|||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | Pt-SPIKE1.3 | |||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||
|
Paralogs
|
|
|||||||||||||||
|
Flax homologues |
|
|||||||||||||||
| PFAM info |
| |||||||||||||||
|
Representative CDS sequence |
>Potri.011G024000.1 pacid=42780392 polypeptide=Potri.011G024000.1.p locus=Potri.011G024000 ID=Potri.011G024000.1.v4.1 annot-version=v4.1
ATGGATAATAGTAATAGTAACGGCAGCAGCGGGGGGCAGCGGTTCCGGAAAATACCTCGGCATTCTCAGTCCCTTTCTCATCTCAAGTTGGACCCGCTGG
TAGATGAAAATTTGGAGCAGTGGCCACATCTTAATGAACTTGTTCAATGTTATAGAACTGATTGGGTGAAGGATGAAAACAAGTATGGTCATTATGAAAG
TATCTCTCCAGTATCTTTTCAAAACCAGATTTTTGAAGGTCCGGATACTGATCTCGAGACAGAAATGCATCTTGCCAATTCAAGACGAAACAAGGCTGAA
GAGACCACCGATGATGACATCCCAAGTACTTCAGGAAGGCAATTTGTGGAGGCCGCCTTCCCCGACTCATCAAATTCTGTTGTTTCAAAGCATTTTGGTG
AATCTCCTCTCCCTGCATATGAACCAGCTTTTGATTGGGATAATGAGAGGTCGATGATATTTGGCCAAAGGATTCCTGAAACTCCATTGCCACAGTATGA
CAGTGGATTGAAGATCTCTGTGAAAGTACTGTCCCTATCATTTCAAGCAGGATTAGCTGAGCCATTTTATGGTACAATTTGCATATATAATAAGGAGAGA
AGAGAAAAGTTGTCCGAGGATTTTTATTTTTCTGTGGTGCCAACAGACACACAAGATGCAAAAATTTCTCATGATCCTCGTGGAATTTTCTATTTAGATG
CTCCTTCATCATCAATATGTTTGCTAATCCAACTAGAGAAGCCTGCCACAGAAGAAGGCGGGGTCACTGCCTCTGTTTATTCACGTAAAGAACCTGTGCA
CTTGAGCGAGAGAGAGAAGCAGAAACTGCAGGTCTGGTCTCGTATTATGCCTTACAAAGAATCCTTTGCCTGGACTATTGTTCCATTATTTGATAACAGT
ATTGCTGCGACTTCCGGGGGGGCTGCTTCTCCAAGCAGCCCTCTTGCTCCAAGTGTATCTGGCTCAAGCTCCCATGACGGTGTTTTCGAGCCTGTTGCAA
AGATCACTCTGGATGGGAAGTTGGGTTATTCAAGTGGAAGCTCTGTTGTGGTTGAAATATCAAATTTAAATAAAGTTAAGGAAAGCTACACTGAGGACTC
ACTTCAGGATCCTAAACGCAAAGTTCATAAACCTGTCAAGGGTGTTCTCCGACTAGAAATTGAAAAGCATCAGACAGCCCATGCTGAGTTGGAAAATTTA
TCAGAAACTGGCAGTATAACCAATGATTCCATTGATCTGGGGGATCGTGTTGCTGATTCTGCATTTACAAAATCCCCTAGCAATGGCTTTGATGATCCTC
AAACTAGTGGTTCGAAGTGGAATATCTTTGATGGGAAAGAAACATCTGGAAATATATCAAATGCTCGTGAAAATCCTGATTTCACTGCTGATGATTTTCA
AGCTTTTGATTTCCGCACGACCACGAGGAACGAGCCTTTCTTGCAGCTTTTCCATTGCCTGTATGTTTACCCATTAACTGTCAGTTTGAGTCGGAAGAGG
AATTTGTTCATTCGGGTTGAACTAAGGAAGGATGACGTGGATGTTCGTAGACAGCCTTTAGAGGCAATGCATCCCAGGGAGCCAGGAACATCGCTCCAAA
AGTGGGCTCACACACAAGTAGCTGCTGGGACTAGAGTGGCTTGCTATCATGATGAAATCAAACTTTCTCTGCCAGCGATTTGGACACCATCCCATCACCT
CTTGTTCACTTTCTTCCATGTTGATCTTCAAACCAAACTGGAAGCTCCAAAACCAGTGGTTATCGGATATGCAGTGCTTCCATTATCCACACATGCACAG
TTGAGATCTGAAATTTCTTTGCCAATAATGAGAGAGCTGGTTCCACATTATCTCCAGGAAATGGGAAAGGAGAGGCTGGATTATTTGGAAGATGGAAAAA
ATGTCTTCAGATTGCGCTTGAGACTATGCTCATCTCTGTATCCCATTAATGAGCGCATTAGAGATTTTTTTATTGAATATGATAGGCACACTCTTCGTAC
TAGTCCACCTTGGGGATCTGAACTTCTTGAGGCCATTAATAGTCTGAAGAACGTTGATTCCACTGCTTTGCTTCAGTTTCTCCACCCAATTTTGAATATG
CTTCTCCATCTTATTGGCAGTGGTGGAGAAACTCTCCAGGTTGCAGCCTTCAGGGCCATGGTTAATATCCTAACTAGGGTGCAGCAAGAGTCAGTTGATG
ATACTGAACGGAATCGTTTTCTGGTTAACTATGTTGATTATGCTTTTGATGACTTTGGTGGTCGTCAACCACCTGTTTATCCTGGTTTGTCCACTGTGTG
GGGGAGTCTGGCTCGTAGCAAGGCTAAAGGCTATCGTGTTGGACCAGTATATGATGATGTATTGGCAATGGCATGGTTTTTCCTTGAGCTTATTGTCAAG
TCAATGGCATTGGAGCAAGCTCGTCTTTTCTATCATAGTCTTCCATTGGGTGAAGATGTTCCACCGATGCAATTGAAAGAAGGTGTGTTCAGATGCATCA
TGCAGTTATATGATTGCCTTCTTACAGAGGTGCATGAGCGTTGTAAGAAGGGTTTAAGCTTGGCAAAACGTTTGAATAGCAGTTTGGCCTTCTTCTGCTA
TGATCTTTTGTCAATCATTGAACCTCGTCAAGTTTTTGAATTGGTGTCCTTGTACCTGGACAAGTTTTCTGGGGTATGCCAATCAGTTCTTCATGACTGC
AAGCTTACCTTTTTGCAAATCATATGTGATCATGATCTTTTTGTGGAGATGCCGGGGAGGGACCCTTCAGATAGAAACTACCTTGCATCTGTTTTAATAC
AAGAGCTTTTCCTTACTTGGGATCATGACGAGCTATCTCAACGGTCAAAGGCAGCCAGAATCTTAGTAGTACTGCTTTGCAAGCATGAATTTGATGCTCG
ATACCAAAAGCCTGAAGATAAACTATATATCGCACAGCTATATTTTCCACTTGTTGGCCAGATCCTGGATGAGATGCCTGTCTTCTACAACCTGAATGCT
GTTGAAAAGCGTGAAGTTTTAATTGTTATTTTGCAAATCATGCGCAATTTGGATGATACATCGCTTGTCAAGGCGTGGCAGCAAAGCATTGCTCGGACTA
GATTATTTTTCAAACTCATGGAGGAATGCCTTGTTCTTTTTGAGCACAGAAAACCTGCTGATGGCATTCTTATGGGATCTAGTTCTCGCAGTCCTGTTGG
GGATGGACCTGCTTCTCCCAAGTACTCCGATAGGCTCTCTCCAGCAATCAACAATTATCTGTCTGAGGCATCACGGCAAGAAGTCAGGCCTCAGGGTAAA
ACTGATAATGGATATTTGTGGCAGAGAGTGAATTCCCAATTAAGCTCCCCCAGCCAGCCATATTCCTTGAGAGAAGCACTTGCTCAGGCACAATCTTCCA
GGATTGGAGCTTCAGCCCAAGCACTTAGAGAATCCTTGCATCCAATATTGAGACAAAAACTGGAGCTTTGGGAAGAAAACTTGAGTGCTGCTGTGAGTCT
TCAGGTTCTGGAAATAACTGAGAAGTTTTCTATGATGGCAGCTTCCCATAGCATTGCTACTGATTATGGGAAACTTGATTGCCTCACAGCCATATTTACG
AGCTTCTTCTCTCGAAATCAACCATTGTCTTTCTGGAAAGCTCTATTTCCTGTCTTTAACAATGTGTTTGATCTTCATGGGGCAACATTAATGGCCAGGG
AGAACGATCGTTTCTTAAAGCAGGTTGCTTTCCATCTTCTTCGGCTTGCAGTTTTTAGAAACGAAAGTGTCAAGAAAAGAGCTGTTATTGGCCTCCAGAT
TCTTGTCAGGAGCGCTTTCTATTACTTCATGCAAACAGCAAGGTTGAGGGTCATGCTGACTATCACACTGTCAGAGCTGATGTCTGATGTGCAAGTGACT
CAGATGAAGTCTGATGGAATGCTAGAGGAGAGTGGTGAAGCAAAGCGTCTTCGCAAGTCTTTGGAGGAGGTGGCAGATGAATTGAAGACTCCTGACCTAC
TACGGGAATGTGGAGTTCCTGAAAGTGCTCTGGTGGCAGTTCCAAAAAAGTTGGCAGACAATAGGTGGTCCTGGAGTGAGGTGAAATATCTCTCAGACTG
CCTTATCCTGGCCCTTGATGCTAGCCTTGAACATGCACTCTTGGGTTCTGTAATGACTGTGGATAGATATGCGGCTGCAGAGAGCTTCTATAAACTAGCA
ATGGCATTTGCTCCTGTTCCTGATCTTCACATAATGTGGTTACTGCATTTATGCGATGCGCATCAGGAAATGCAGTCTTGGGCTGAAGCTGCACAGTGTG
CTGTGGCTGTTGCTGGTGTTGTAATGCAGGCTCTTGTGGCTAGGAATGATGGTGTGTGGAGCAAAGATCACGTGATCTCTCTACGTAAAATTTGCCCAAT
GGTCAGCAGTGAGATCACTGCTGAGGCATCTGCAGCTGAGGTAGAGGGGTATGGTTCTTCAAAGCTCACAGTGGACTCTGCTGTAAAATATCTACAGCTT
GCTAATAGGCTTTTCTCTCAAGCTGAGCTTTTTCATTTCTGTGCAAACATTCTGGAGCTTGTAATACCAGTTCATAAAAGTAGGAGGGCATATGGGCAGC
TGGCTAAATGTCACACCATGCTTACTGATATCTATGAATCAATTCTGGAGCAGGAATCAAGCCCAATTCCGTTCACTGATGCCACATACTATAGGGTGGG
ATTCTATGGTGAAAGGTTTGGGAAACTAGATAGGAAGGAATATGTATACCGAGAGCCCCGTGATGTACGGCTAGGTGACATAATGGAGAAACTTAGTCAT
ATTTATGAGTCTAGGATGGACGACAATCACACTTTGCATATTATTCCAGATTCTAGGCAAGTAAAGGCTGACGAGCTGCAGCCCGGAGTCTGCTACCTGC
AGATAACTGCTGTTGATCCGGTCATGGAAGATGAAGATTTAGGAAGCAGAAGGGAAAGGATCTTTTCTCTTTCCACTGGAACTGTCCGGGCACGTGTCTT
TGACCGCTTCTTGTTTGACACTCCATTTACAAAAAATGGTAAAACCCAAGGTGGATTGGAAGACCAATGGAAGAGGAGGACAGTTCTTCAGACTGAAGGC
TCATTCCCTGCTCTGGTGAACAGGCTTTTGGTAATGAAATCTGAATCTCTTGAGTTCTCACCAGTCGAGAATGCAATTGGAATGATTGAAACTCGAACAG
CTGCCTTACGAAACGAGCTTGAGGAGCCCCGCAGCTCTGAAGGCGATCAGCTTCCACGTCTCCAAAGTTTGCAGAGAATTCTCCAAGGCAGTGTTGCCGT
TCAAGTAAACAGCGGCGTTTTGAGTGTCTGCACGGCCTTCCTGTCGGGTGAGCCTGCTACAAGGTTGCGTTCTCAAGAACTGCAGCAACTCATTGCTGCC
TTGCTTGAATTCATGGCTGTTTGCAAGCGCGCCATTCGTGTACATTTTAGATTGATTGGAGAGGAAGATCAAGATTTCCACACGCAGCTTGTGAACGGAT
TTCAATCGCTCACTGCAGAGTTATCTCATTACATCCCAGCCATTCTTGCAGAACTATGA
|
|||||||||||||||
|
AA sequence
|
>Potri.011G024000.1 pacid=42780392 polypeptide=Potri.011G024000.1.p locus=Potri.011G024000 ID=Potri.011G024000.1.v4.1 annot-version=v4.1
MDNSNSNGSSGGQRFRKIPRHSQSLSHLKLDPLVDENLEQWPHLNELVQCYRTDWVKDENKYGHYESISPVSFQNQIFEGPDTDLETEMHLANSRRNKAE
ETTDDDIPSTSGRQFVEAAFPDSSNSVVSKHFGESPLPAYEPAFDWDNERSMIFGQRIPETPLPQYDSGLKISVKVLSLSFQAGLAEPFYGTICIYNKER
REKLSEDFYFSVVPTDTQDAKISHDPRGIFYLDAPSSSICLLIQLEKPATEEGGVTASVYSRKEPVHLSEREKQKLQVWSRIMPYKESFAWTIVPLFDNS
IAATSGGAASPSSPLAPSVSGSSSHDGVFEPVAKITLDGKLGYSSGSSVVVEISNLNKVKESYTEDSLQDPKRKVHKPVKGVLRLEIEKHQTAHAELENL
SETGSITNDSIDLGDRVADSAFTKSPSNGFDDPQTSGSKWNIFDGKETSGNISNARENPDFTADDFQAFDFRTTTRNEPFLQLFHCLYVYPLTVSLSRKR
NLFIRVELRKDDVDVRRQPLEAMHPREPGTSLQKWAHTQVAAGTRVACYHDEIKLSLPAIWTPSHHLLFTFFHVDLQTKLEAPKPVVIGYAVLPLSTHAQ
LRSEISLPIMRELVPHYLQEMGKERLDYLEDGKNVFRLRLRLCSSLYPINERIRDFFIEYDRHTLRTSPPWGSELLEAINSLKNVDSTALLQFLHPILNM
LLHLIGSGGETLQVAAFRAMVNILTRVQQESVDDTERNRFLVNYVDYAFDDFGGRQPPVYPGLSTVWGSLARSKAKGYRVGPVYDDVLAMAWFFLELIVK
SMALEQARLFYHSLPLGEDVPPMQLKEGVFRCIMQLYDCLLTEVHERCKKGLSLAKRLNSSLAFFCYDLLSIIEPRQVFELVSLYLDKFSGVCQSVLHDC
KLTFLQIICDHDLFVEMPGRDPSDRNYLASVLIQELFLTWDHDELSQRSKAARILVVLLCKHEFDARYQKPEDKLYIAQLYFPLVGQILDEMPVFYNLNA
VEKREVLIVILQIMRNLDDTSLVKAWQQSIARTRLFFKLMEECLVLFEHRKPADGILMGSSSRSPVGDGPASPKYSDRLSPAINNYLSEASRQEVRPQGK
TDNGYLWQRVNSQLSSPSQPYSLREALAQAQSSRIGASAQALRESLHPILRQKLELWEENLSAAVSLQVLEITEKFSMMAASHSIATDYGKLDCLTAIFT
SFFSRNQPLSFWKALFPVFNNVFDLHGATLMARENDRFLKQVAFHLLRLAVFRNESVKKRAVIGLQILVRSAFYYFMQTARLRVMLTITLSELMSDVQVT
QMKSDGMLEESGEAKRLRKSLEEVADELKTPDLLRECGVPESALVAVPKKLADNRWSWSEVKYLSDCLILALDASLEHALLGSVMTVDRYAAAESFYKLA
MAFAPVPDLHIMWLLHLCDAHQEMQSWAEAAQCAVAVAGVVMQALVARNDGVWSKDHVISLRKICPMVSSEITAEASAAEVEGYGSSKLTVDSAVKYLQL
ANRLFSQAELFHFCANILELVIPVHKSRRAYGQLAKCHTMLTDIYESILEQESSPIPFTDATYYRVGFYGERFGKLDRKEYVYREPRDVRLGDIMEKLSH
IYESRMDDNHTLHIIPDSRQVKADELQPGVCYLQITAVDPVMEDEDLGSRRERIFSLSTGTVRARVFDRFLFDTPFTKNGKTQGGLEDQWKRRTVLQTEG
SFPALVNRLLVMKSESLEFSPVENAIGMIETRTAALRNELEEPRSSEGDQLPRLQSLQRILQGSVAVQVNSGVLSVCTAFLSGEPATRLRSQELQQLIAA
LLEFMAVCKRAIRVHFRLIGEEDQDFHTQLVNGFQSLTAELSHYIPAILAEL
|
DESeq2's median of ratios [POPLAR]
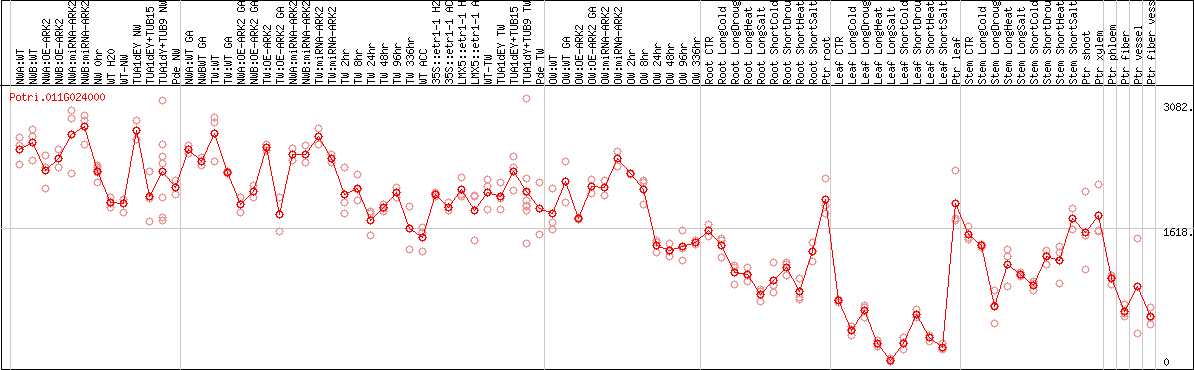
Coexpressed genes
Potri.011G024000 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
