External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G22060 67 / 3e-13
Receptor-like protein kinase-related family protein (.1)
AT5G48540 60 / 8e-11
receptor-like protein kinase-related family protein (.1)
AT3G60720 56 / 2e-09
PDLP8
plasmodesmata-located protein 8 (.1)
AT4G23220 56 / 3e-09
CRK14
cysteine-rich RLK (RECEPTOR-like protein kinase) 14 (.1)
AT4G20670 54 / 1e-08
Domain of unknown function (DUF26) (.1)
AT4G23280 54 / 2e-08
CRK20
cysteine-rich RLK (RECEPTOR-like protein kinase) 20 (.1)
AT1G61750 53 / 3e-08
Receptor-like protein kinase-related family protein (.1)
AT4G23180 50 / 2e-07
RLK4, CRK10
cysteine-rich RLK (RECEPTOR-like protein kinase) 10 (.1)
AT4G23250 50 / 4e-07
RKC1, CRK17, DUF26-21, EMB1290
RECEPTOR-LIKE KINASE THALIANACOL-0GENOMICLIBRARY\(CLONTECH\).THEPOSITIVEPHAGECLONES C-X8-C-X2-C CLASS 1, EMBRYO DEFECTIVE 1290, CYSTEINE-RICH RLK \(RECEPTOR-LIKE PROTEIN KINASE\) 17, kinases;protein kinases (.1)
AT4G38830 49 / 6e-07
CRK26
cysteine-rich RLK (RECEPTOR-like protein kinase) 26 (.1)
Paralogs
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10006752
166 / 8e-51
AT5G48540 100 / 1e-24
receptor-like protein kinase-related family protein (.1)
Lus10007631
68 / 3e-13
AT4G23180 257 / 1e-76
cysteine-rich RLK (RECEPTOR-like protein kinase) 10 (.1)
Lus10027144
63 / 2e-12
ND 121 / 1e-34
Lus10027145
62 / 1e-11
AT3G22060 253 / 8e-85
Receptor-like protein kinase-related family protein (.1)
Lus10007632
63 / 2e-11
AT4G05200 624 / 0.0
cysteine-rich RLK (RECEPTOR-like protein kinase) 25 (.1)
Lus10038227
62 / 2e-11
AT5G48540 261 / 1e-87
receptor-like protein kinase-related family protein (.1)
Lus10025875
61 / 6e-11
AT5G48540 261 / 2e-87
receptor-like protein kinase-related family protein (.1)
Lus10018380
61 / 1e-10
AT4G05200 324 / 2e-104
cysteine-rich RLK (RECEPTOR-like protein kinase) 25 (.1)
Lus10031582
60 / 2e-10
AT4G05200 436 / 6e-145
cysteine-rich RLK (RECEPTOR-like protein kinase) 25 (.1)
Lus10039699
57 / 1e-09
AT3G22060 171 / 5e-52
Receptor-like protein kinase-related family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF01657
Stress-antifung
Salt stress response/antifungal
Representative CDS sequence
>Potri.011G029500.2 pacid=42780590 polypeptide=Potri.011G029500.2.p locus=Potri.011G029500 ID=Potri.011G029500.2.v4.1 annot-version=v4.1
ATGCGCCTATCCAAAATGGCTTCTACACGACCGCAGCAGGTAAAGGTGCTAACAAGATTTATGGCCTTACACAATGCAGAGGTGATATTTCTGCCACAGA
CTGTGCAGCCTGCATCAAGAATGTTACTTGGACCGGAGCTTTGGTTTAAGTGGTGTTTCCTAAGGTATTCAGACCGGAGCTTTTTCGGTGAATTGGATCA
ATCAGGAATGGTGGCAACTTACAATGACACCAATTTTGAAGATGCAAAAGTGGTTTCTGAAGGACTTAACTTCACAAAAACGCTTGCTTCTACAACTCCC
AACCAGCCCTCGATGTTCTATACCGCAGTATTGGATGTTGGACAGAGTGGGAAGAGGTATGGCATGGCTCAATGCACTAGAGATCTCAGCAAGAGTGACT
GTGGCAAGTGCTTGGATTTCCAATTGGCGACATATTTAAATATTATTGGAAATAAGAGAAGTTGGGATATTTATGGATCTAGTTGTAGAATGTGGTACTA
TGACTACCAGTTCTACTTCAACTTTTCAACCCCTGCAGCAAAGGGAGGTTCCACAAGATCCTCACCGCATCGAGTTGCAATTGGCATGGCCTTTCCAGTG
CTAGTGTTTCTGTTAGTTCTTTAA
AA sequence
>Potri.011G029500.2 pacid=42780590 polypeptide=Potri.011G029500.2.p locus=Potri.011G029500 ID=Potri.011G029500.2.v4.1 annot-version=v4.1
MRLSKMASTRPQQVKVLTRFMALHNAEVIFLPQTVQPASRMLLGPELWFKWCFLRYSDRSFFGELDQSGMVATYNDTNFEDAKVVSEGLNFTKTLASTTP
NQPSMFYTAVLDVGQSGKRYGMAQCTRDLSKSDCGKCLDFQLATYLNIIGNKRSWDIYGSSCRMWYYDYQFYFNFSTPAAKGGSTRSSPHRVAIGMAFPV
LVFLLVL
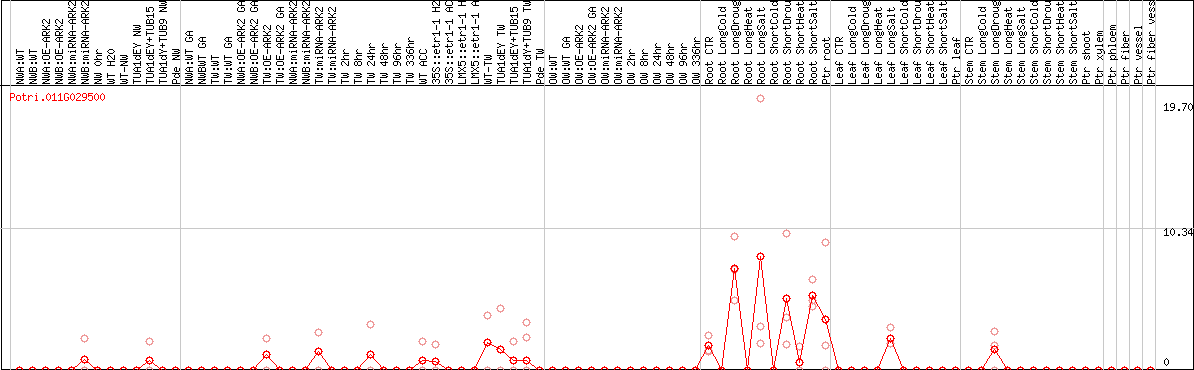
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.011G029500 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.