Pt-ATMRP12.3 (Potri.011G043100) [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | Pt-ATMRP12.3 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.011G043100.2 pacid=42781746 polypeptide=Potri.011G043100.2.p locus=Potri.011G043100 ID=Potri.011G043100.2.v4.1 annot-version=v4.1
ATGGGTTTGGAGGCATTGGTTTGGTATTGTCGACCTGTGCCAAATGGGGTTTGGGCAACAAAAGTGGATAATGCATTTGGTGCTTACACACCCTGTGTTG
TTGACTCCCTAGTCATTTGCATTTCTCATCTGGTTCTTCTTGGCCTTTGTTTGTATCGAATATGGTTGATTACTGACAAAAATTCCAAAGCTCAACACTA
TTGTCTCAGGACAAATTACTATAATTACTCGTTGGGATTGCTGGCTGCTTATTGCACAGTCCAGCCTTTATTCAGGTTGTTTATGAATGTTTCAATCTTC
AATCTGGATGGGCAGACTGCTCTCGCTCCCTTTGAGATGGTCTCCTTGATTGTTGAGGCCCTTTCCTGGTGCTCCACTCTTATTATGATTGGTTTGGAAA
CAAGAATATACATTCAACAGTTTCGATGGTATGTGCGGTTTGGAGTCATTTATGTGTTGGTAGGGGAAGCGGCAATGCTCAATCTCATTCTGTCAGTGAG
TGATAATTACGACAGTAGATTTATATTCTATATGTATTTGAGCACAGTTTTCTGCCAGGTTTTGTTTGGGATACATCTACTTGTTTATATTCCCAATTTG
GATCCTTGTTCAGATTATGTTATGATGGAACCTGAGTCTCCTGATAATAGTGCATATGAAGCACTTCCTGGAAGAGAGCAAATATGTCCCGAGAGGAATG
CCACCTTATTTTCTAGAATATTTTATTGGTGGTTGACTCCACTCATGAAGCAAGCCCATAAAAGACCTATTAGTGAAAAGGATGTTTGGAAGCTAGACAC
ATGGGACCAGACTGAGACATCGATGAACAAGTTCCAGACTTGTTGGGTTGAAGAATCTCAAAGGCCAAAGCCATGTCTTTTAAGAGCACTAAACAATAGT
CTAGGGGGAAGGTTTTGGTTAGGAGGATTTTTTAAGATTGGCTATGATCTTTCTGAATTTGTGGGACCTGTTGTATTGAGCCACCTCCTGCAGTCGATGC
AACGAGGAGATCCAGCCTGGATCGGTTATGTCTATGCCTTCGTAATTTTCCTTGGGATGTTGTTTAGTGCGCTATGCGAATCTCGGTATTACCAGAATGT
TCTGCGTGTTGGTTTTCGGTTGCGGTCAACTTTGGTGGCTGGTATATTTCGTAAATCCTTGAAACTAACTCATGAGGGTCAAAAGAACTTTCCATCAGGG
AAGATAACAAATATGATCACAACAGATGCTGATGTACTTCAGCAAATATGCCTACTACTTCATGGATTGTGGTCCGCGCCATTTTGCATCACCATGTCCA
TGGTTCTCCTTTACCAGCAATTAGGTGTTGCTTCACTTTTTGGTTCGCTAGTGCTTGTTATCATGGTCCCTACACAAGCAATCTTGTTGAACAGAATGAC
ACGACTAACTAAGGAAGGATTGCATAGGACAGACAAAAGAGTCAGCCTCATGAATGAAATCTTGGCTGCCATGGACACTGTGAAATGTTATGCATGGGAG
AAGAGCTTTCAATTTAGGGTTCAAAGTGTGCGGAATGATGAGCTGTCCTTGTTTCGCAGTGCCCAATTACTATTTGCTTTCAACAGTTTTATGGTGAATA
GCATTCCAGTTGTTGTGACACTTGTTTCATTCGGCACATTCACCTTGCTTGGTGGAGACTTGACCCCTGCAAAGGCTTTTACATCACTTTCTTTGTTTCA
AGTGTTAAGATACCCTCTAAACATGCTTCCTAACTTACTGAGCCAGGTTGTAAATGCAAATATATCATTGCAACGCTTGGAGGAACTGTTTCTAGCTGAA
GAGAGAATTTTAGCACCAAATCCACCTCTTGAACCAGGAATTCCAGCTATCTCAATTGAAAATGGGAACTTTTCATGGGATTTAAAGCTAGAAAATCCAA
CGTTGACAAATATCAAATTAAACATACAAGTTGGCAGCTTAGTCGCAATCGTTGGTGGGACTGGAGAAGGAAAAACATCACTTATATCAGCAATGCTTGG
AGAGCTGCCTCCTATGGAAGATGCATGTGTTGTTATAAGAGGGACTGTTGCATATGCTCCTCAAGTTCCATGGATCTTCAATGCTACTGTTCGTGACAAC
ATATTATTTGGGTCAAAATATGAACCTTCACGATATGGGAAAGCTATTGATGTAACTGCATTACAACATGATCTTGACTTATTTGCAGGCCATGACCTCA
CTGAGATAGGTGAAAGAGGAGTCAATATTAGTGGTGGGCAGAAGCAAAGAATTTCCATGGCCAGGGCTTTCTACTCCAATTCAGATATATACATATTCGA
TGACCCTTTAAGTGCTTTAGATGCTCACGTTGCCCGACAGGTCTTTAACAGTTGTATTAAAGAAGGGTTGCAAGGAAAAACCAGGGTTCTTGTGACAAAC
CAGCTACATTTCCTTCCACAAGTGGAAAAAATTATTCTTCTCAGTGAAGGTATGATTAAAGAGGAGGGAACCTTCGAGGAACTCTTTAAAAACAGCGAAC
TATTCCAGAAGCTGATGGAAAATGCAGGGAAAATGGAAGAGCAAGTGAAAGAAAAGGAGAAAAGTGACAATTTGGACCACAAAAGCTCAAAAGCAGAAGC
TAATTGGGAAAATGAGTTGCCACAAAAGGCAGCCTCTACAATGAAAGGGAAAGAAGGGAAGTCTATTCTCATCAAGCAAGAAGAACGAGAAAGAGGTGTT
GTCAGTTGGAATGTTTTGATAAGGTACAACAACGCATTAGGAGGTGTATGGGTGGTTTCAATACTTTTTCTGTGCTACTTATTAACTGAAGTTTTTCGAG
TTTCAAGAAGTACATGGTTGAGTTTTTGGACAAATCAAAGCACTTTGGAGAGTTATAGGCCAGGATACTTCATTTTCGTGTATGGACTTCTATCATTTGG
TCAGGTGACTGTGACACTAGCAAACTCTTATTGGTTAATCAGTTCAAGTCTTCATGCGTCCAAAAGACTTCATGATGCCATGCTAGATTCCATCCTACGA
ACTCCAATGCTATTCTTCCACACCAACCCAACTGGGAGGATAATCAATAGGTTTGCGAAGGATGTAGGTGAAATAGATCGTAACGTTGCCAATTCTGCCA
ACAATTTTCTAAACCTAGCTTGGCAGTTGCTTTCAACTTTTGTGCTGATAGGCACTGTGAGCACAATATCATTGTGGGCCATAATGCCACTTCTGATCTT
ATTTTATTCAGCCTATCTATATTATCAGAATACATCCCGTGAAGTGAAACGTCTGGATTCCATCACCAGGTCTCCAGTTTATGCACAGTTTGGAGAAGCA
TTAAATGGTTTGTCAAGTATTCGTGCTTATAAAGCATATGATTGGATGTCCATCATCAATGGGAAGTATATGGACAATAATATTAGATTTAGTCTCGTAA
CTATTAGTTCAGATGGTTGGCTTGCCATAAGGTTGGTAACTTTAGGAGGGATGATGATTTGGTTGATAGCGTCATTTTCAGTTTTGGGGAATGGAAGAAC
TGAAAATCATGTGGGATTTGCATCTATAATGGGTCTACTTCTTAGTTACACTTCGAACATCACTGATTTGTTGAGTAATGTTCTAAGACAAGCTAGTAAA
GCTGAAAATAGCTTAAACTCTGTTGAGCGTGTTTCCACATATATAGATTTACCCTCCGAAGCTCCAGCCATAGACAAGAACAATCGTCCTCCTTCTTCTT
GGCCTTTGTCAGGATTAATTAAATTTACGGATGTTGTTCTACGTTACAGGCCCGAGCTTCCTCCAGTCTTACATGGTTTGTCCTTTGCAGTCTCACCAAG
TGAAAAGCTAGGAATAGTCGGAAGAACTGGAGCTGGGAAATCTAGCATGTTAAATGCATTGTTCCGTATCGTAGAACTGGAAAGAGGAGAAATCACCATT
GATGGTTGTGACATTACTAAGTTCGGACTGACAGATTTGAGGAGAGCTCTCAGTATCATACCACAATCACCAGTTCTCTTCTCAGGGACTGTGCGATTTA
ATCTTGATCCATTCAGTGAACACAATGATGCTGACCTATGGAAGGCTTTAGAGAGGGCACATTTGAAGGATGCTGTTAGAAATAGTTCTTTTGGTCTGGA
TGCTCAGGTTTTTGGAGGGGAGAGTTTTAGTGTTGGACAGAGGCAACTACTAAGTCTCGCTCGAGCACTGCTGCGGAGATCTAAGATTCTTGTTCTTGAT
GAAGCTACTTCTTCTGTTGATGTTAGAATTGATGCTCTTATCCAGAAAACCATCCGAGAAGAATTCAGATCCTGCACAATGCTAATTATCGCTCATAGAC
TAAATACCATTATTGACTGTGACAGAATTCTTGTGCTTGAGGCTGGTCAGGTTTTAGAACATTCTACCCCAGAGGAACTGCTATCGAACGAAGGAAGCGC
ATTCTCTAGGATGGTTCAAAGTACAGGACCAGCAAATGCACAGTACTTGCATAGCCTGGTGTTTGAAAGCAAAGAGAACAAGTTAATGTTTATACATTCC
TGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.011G043100.2 pacid=42781746 polypeptide=Potri.011G043100.2.p locus=Potri.011G043100 ID=Potri.011G043100.2.v4.1 annot-version=v4.1
MGLEALVWYCRPVPNGVWATKVDNAFGAYTPCVVDSLVICISHLVLLGLCLYRIWLITDKNSKAQHYCLRTNYYNYSLGLLAAYCTVQPLFRLFMNVSIF
NLDGQTALAPFEMVSLIVEALSWCSTLIMIGLETRIYIQQFRWYVRFGVIYVLVGEAAMLNLILSVSDNYDSRFIFYMYLSTVFCQVLFGIHLLVYIPNL
DPCSDYVMMEPESPDNSAYEALPGREQICPERNATLFSRIFYWWLTPLMKQAHKRPISEKDVWKLDTWDQTETSMNKFQTCWVEESQRPKPCLLRALNNS
LGGRFWLGGFFKIGYDLSEFVGPVVLSHLLQSMQRGDPAWIGYVYAFVIFLGMLFSALCESRYYQNVLRVGFRLRSTLVAGIFRKSLKLTHEGQKNFPSG
KITNMITTDADVLQQICLLLHGLWSAPFCITMSMVLLYQQLGVASLFGSLVLVIMVPTQAILLNRMTRLTKEGLHRTDKRVSLMNEILAAMDTVKCYAWE
KSFQFRVQSVRNDELSLFRSAQLLFAFNSFMVNSIPVVVTLVSFGTFTLLGGDLTPAKAFTSLSLFQVLRYPLNMLPNLLSQVVNANISLQRLEELFLAE
ERILAPNPPLEPGIPAISIENGNFSWDLKLENPTLTNIKLNIQVGSLVAIVGGTGEGKTSLISAMLGELPPMEDACVVIRGTVAYAPQVPWIFNATVRDN
ILFGSKYEPSRYGKAIDVTALQHDLDLFAGHDLTEIGERGVNISGGQKQRISMARAFYSNSDIYIFDDPLSALDAHVARQVFNSCIKEGLQGKTRVLVTN
QLHFLPQVEKIILLSEGMIKEEGTFEELFKNSELFQKLMENAGKMEEQVKEKEKSDNLDHKSSKAEANWENELPQKAASTMKGKEGKSILIKQEERERGV
VSWNVLIRYNNALGGVWVVSILFLCYLLTEVFRVSRSTWLSFWTNQSTLESYRPGYFIFVYGLLSFGQVTVTLANSYWLISSSLHASKRLHDAMLDSILR
TPMLFFHTNPTGRIINRFAKDVGEIDRNVANSANNFLNLAWQLLSTFVLIGTVSTISLWAIMPLLILFYSAYLYYQNTSREVKRLDSITRSPVYAQFGEA
LNGLSSIRAYKAYDWMSIINGKYMDNNIRFSLVTISSDGWLAIRLVTLGGMMIWLIASFSVLGNGRTENHVGFASIMGLLLSYTSNITDLLSNVLRQASK
AENSLNSVERVSTYIDLPSEAPAIDKNNRPPSSWPLSGLIKFTDVVLRYRPELPPVLHGLSFAVSPSEKLGIVGRTGAGKSSMLNALFRIVELERGEITI
DGCDITKFGLTDLRRALSIIPQSPVLFSGTVRFNLDPFSEHNDADLWKALERAHLKDAVRNSSFGLDAQVFGGESFSVGQRQLLSLARALLRRSKILVLD
EATSSVDVRIDALIQKTIREEFRSCTMLIIAHRLNTIIDCDRILVLEAGQVLEHSTPEELLSNEGSAFSRMVQSTGPANAQYLHSLVFESKENKLMFIHS
|
DESeq2's median of ratios [POPLAR]
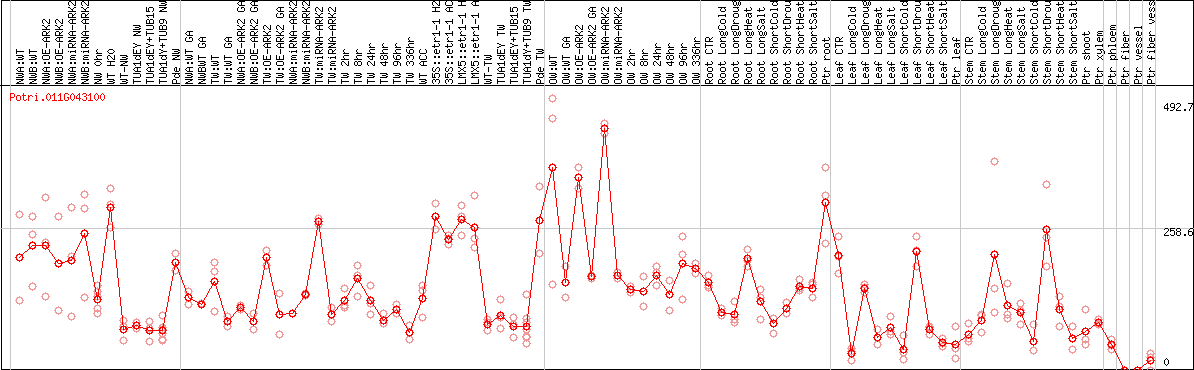
Coexpressed genes
Potri.011G043100 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
